This Is the Published Version of a Paper
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Dynamic Molecular Linkers of the Genome: the First Decade of SMC Proteins
Downloaded from genesdev.cshlp.org on October 8, 2021 - Published by Cold Spring Harbor Laboratory Press REVIEW Dynamic molecular linkers of the genome: the first decade of SMC proteins Ana Losada1 and Tatsuya Hirano2,3 1Spanish National Cancer Center (CNIO), Madrid E-28029, Spain; 2Cold Spring Harbor Laboratory, Cold Spring Harbor, New York 11724, USA Structural maintenance of chromosomes (SMC) proteins in eukaryotes. The proposed actions of cohesin and con- are chromosomal ATPases, highly conserved from bac- densins offer a plausible, if not complete, explanation for teria to humans, that play fundamental roles in many the sudden appearance of thread-like “substances” (the aspects of higher-order chromosome organization and chromosomes) and their longitudinal splitting during dynamics. In eukaryotes, SMC1 and SMC3 act as the mitosis, first described by Walther Flemming (1882). core of the cohesin complexes that mediate sister chro- Remarkably, SMC proteins are conserved among the matid cohesion, whereas SMC2 and SMC4 function as three phyla of life, indicating that the basic strategy of the core of the condensin complexes that are essential chromosome organization may be evolutionarily con- for chromosome assembly and segregation. Another served from bacteria to humans. An emerging theme is complex containing SMC5 and SMC6 is implicated in that SMC proteins are dynamic molecular linkers of the DNA repair and checkpoint responses. The SMC com- genome that actively fold, tether, and manipulate DNA plexes form unique ring- or V-shaped structures with strands. Their diverse functions range far beyond chro- long coiled-coil arms, and function as ATP-modulated, mosome segregation, and involve nearly all aspects of dynamic molecular linkers of the genome. -

2020 Program Book
PROGRAM BOOK Note that TAGC was cancelled and held online with a different schedule and program. This document serves as a record of the original program designed for the in-person meeting. April 22–26, 2020 Gaylord National Resort & Convention Center Metro Washington, DC TABLE OF CONTENTS About the GSA ........................................................................................................................................................ 3 Conference Organizers ...........................................................................................................................................4 General Information ...............................................................................................................................................7 Mobile App ....................................................................................................................................................7 Registration, Badges, and Pre-ordered T-shirts .............................................................................................7 Oral Presenters: Speaker Ready Room - Camellia 4.......................................................................................7 Poster Sessions and Exhibits - Prince George’s Exhibition Hall ......................................................................7 GSA Central - Booth 520 ................................................................................................................................8 Internet Access ..............................................................................................................................................8 -

Familial Cortical Myoclonus Caused by Mutation in NOL3 by Jonathan Foster Rnsseil DISSERTATION Submitted in Partial Satisfaction
Familial Cortical Myoclonus Caused by Mutation in NOL3 by Jonathan Foster Rnsseil DISSERTATION Submitted in partial satisfaction of the requirements for the degree of DOCTOR OF PHILOSOPHY in Biomedical Sciences in the Copyright 2013 by Jonathan Foster Russell ii I dedicate this dissertation to Mom and Dad, for their adamantine love and support iii No man has earned the right to intellectual ambition until he has learned to lay his course by a star which he has never seen—to dig by the divining rod for springs which he may never reach. In saying this, I point to that which will make your study heroic. For I say to you in all sadness of conviction, that to think great thoughts you must be heroes as well as idealists. Only when you have worked alone – when you have felt around you a black gulf of solitude more isolating than that which surrounds the dying man, and in hope and in despair have trusted to your own unshaken will – then only will you have achieved. Thus only can you gain the secret isolated joy of the thinker, who knows that, a hundred years after he is dead and forgotten, men who never heard of him will be moving to the measure of his thought—the subtile rapture of a postponed power, which the world knows not because it has no external trappings, but which to his prophetic vision is more real than that which commands an army. -Oliver Wendell Holmes, Jr. iv ACKNOWLEDGMENTS I am humbled by the efforts of many, many others who were essential for this work. -

Bioinformatic Identification of Genes Suppressing Genome Instability
Bioinformatic identification of genes suppressing PNAS PLUS genome instability Christopher D. Putnama,b, Stephanie R. Allen-Solteroa,c, Sandra L. Martineza, Jason E. Chana,b, Tikvah K. Hayesa,1, and Richard D. Kolodnera,b,c,d,e,2 aLudwig Institute for Cancer Research, Departments of bMedicine and cCellular and Molecular Medicine, dMoores-University of California at San Diego Cancer Center, and eInstitute of Genomic Medicine, University of California School of Medicine at San Diego, La Jolla, CA 92093 Contributed by Richard D. Kolodner, September 28, 2012 (sent for review August 25, 2011) Unbiased forward genetic screens for mutations causing increased in concert to prevent genome rearrangements (reviewed in 12). gross chromosomal rearrangement (GCR) rates in Saccharomyces Modifications of the original GCR assay demonstrated that sup- cerevisiae are hampered by the difficulty in reliably using qualitative pression of GCRs mediated by segmental duplications and Ty GCR assays to detect mutants with small but significantly increased elements involves additional genes and pathways that do not GCR rates. We therefore developed a bioinformatic procedure using suppress single-copy sequence-mediated GCRs (13–15). Inter- genome-wide functional genomics screens to identify and prioritize estingly, homologs of some GCR-suppressing genes and pathways candidate GCR-suppressing genes on the basis of the shared drug suppress the development of cancer in mammals (16). Most of the sensitivity suppression and similar genetic interactions as known genes that suppress GCRs have been identified through a candi- GCR suppressors. The number of known suppressors was increased date gene approach. Some studies have screened collections of from 75 to 110 by testing 87 predicted genes, which identified un- arrayed S. -

Identification of Novel Dna Damage Response Genes Using Functional Genomics
IDENTIFICATION OF NOVEL DNA DAMAGE RESPONSE GENES USING FUNCTIONAL GENOMICS by Michael Chang A thesis submitted in conformity with the requirements for the degree of Doctor of Philosophy Graduate Department of Biochemistry University of Toronto © Copyright by Michael Chang (2005) Identification of novel DNA damage response genes using functional genomics Doctor of Philosophy, 2005; Michael Chang; Department of Biochemistry, University of Toronto ABSTRACT The genetic information required for life is stored within molecules of DNA. This DNA is under constant attack as a result of normal cellular metabolic processes, as well as exposure to genotoxic agents. DNA damage left unrepaired can result in mutations that alter the genetic information encoded within DNA. Cells have consequently evolved complex pathways to combat damage to their DNA. Defects in the cellular response to DNA damage can result in genomic instability, a hallmark of cancer cells. Identifying all the components required for this response remains an important step in fully elucidating the molecular mechanisms involved. I used functional genomic approaches to identify genes required for the DNA damage response in Saccharomyces cerevisiae. I conducted a screen to identify genes required for resistance to a DNA damaging agent, methyl methanesulfonate, and identified several poorly characterized genes that are necessary for proper S phase progression in the presence of DNA damage. Among the genes identified, ESC4/RTT107 has since been shown to be essential for the resumption of DNA replication after DNA damage. Using genome-wide genetic interaction screens to identify genes that are required for viability in the absence of MUS81 and MMS4, two genes required for resistance to DNA damage, I helped identify ELG1, deletion of which causes DNA replication defects, genomic instability, and an inability to properly recover from DNA damage during S phase. -

Control of Genome Integrity by RFC Complexes; Conductors of PCNA Loading Onto and Unloading from Chromatin During DNA Replication
Review Control of Genome Integrity by RFC Complexes; Conductors of PCNA Loading onto and Unloading from Chromatin during DNA Replication Yasushi Shiomi *and Hideo Nishitani * Graduate School of Life Science, University of Hyogo, Kamigori, Ako‐gun, Hyogo 678‐1297, Japan Correspondence: [email protected]‐hyogo.ac.jp (Y.S.); [email protected]‐hyogo.ac.jp (H.N.) Academic Editor: Eishi Noguchi Received: 28 November 2016; Accepted: 21 January 2017; Published: 26 January 2017 Abstract: During cell division, genome integrity is maintained by faithful DNA replication during S phase, followed by accurate segregation in mitosis. Many DNA metabolic events linked with DNA replication are also regulated throughout the cell cycle. In eukaryotes, the DNA sliding clamp, proliferating cell nuclear antigen (PCNA), acts on chromatin as a processivity factor for DNA polymerases. Since its discovery, many other PCNA binding partners have been identified that function during DNA replication, repair, recombination, chromatin remodeling, cohesion, and proteolysis in cell‐cycle progression. PCNA not only recruits the proteins involved in such events, but it also actively controls their function as chromatin assembles. Therefore, control of PCNA‐loading onto chromatin is fundamental for various replication‐coupled reactions. PCNA is loaded onto chromatin by PCNA‐loading replication factor C (RFC) complexes. Both RFC1‐RFC and Ctf18‐RFC fundamentally function as PCNA loaders. On the other hand, after DNA synthesis, PCNA must be removed from chromatin by Elg1‐RFC. Functional defects in RFC complexes lead to chromosomal abnormalities. In this review, we summarize the structural and functional relationships among RFC complexes, and describe how the regulation of PCNA loading/unloading by RFC complexes contributes to maintaining genome integrity. -

DNA Replication and Sister Chromatid Cohesion 1 (DSCC1)
Journal of Cancer 2019, Vol. 10 6142 Ivyspring International Publisher Journal of Cancer 2019; 10(24): 6142-6153. doi: 10.7150/jca.32339 Research Paper DNA Replication and Sister Chromatid Cohesion 1 (DSCC1) of the Replication Factor Complex CTF18- RFC is Critical for Colon Cancer Cell Growth Jong-Tae Kim1*, Hee Jun Cho1*, Sang Yoon Park1, Byung Moo Oh1,2, Yo Sep Hwang1,2, Kyoung Eun Baek1, Young-Ha Lee3, Hee Cheol Kim4 and Hee Gu Lee1,2 1. Immunotherapy Research Center, Korea Research Institute of Bioscience and Biotechnology (KRIBB), Daejeon, Republic of Korea. 2. Department of Biomolecular Science, University of Science and Technology (UST), Daejeon, Republic of Korea. 3. Department of Infection Biology, Chungnam National University School of Medicine, Daejeon, Republic of Korea. 4. Department of Surgery, Samsung Medical Center, Sungkyunkwan University School of Medicine, Seoul, Republic of Korea. *These authors contributed equally to this work. Corresponding authors: Hee Cheol Kim, M.D., Ph.D., Department of Surgery, Samsung Medical Center, Sungkyunkwan University School of Medicine, Seoul, Republic of Korea. E-mail: [email protected] or Hee Gu Lee, Ph.D., Immunotherapy Research Center, Korea Research Institute of Bioscience and Biotechnology, Daejeon 34141, Republic of Korea. Tel: +82-42-860-4182; Fax: +82-42-860-4593; E-mail: [email protected] © The author(s). This is an open access article distributed under the terms of the Creative Commons Attribution License (https://creativecommons.org/licenses/by/4.0/). See http://ivyspring.com/terms for full terms and conditions. Received: 2018.12.17; Accepted: 2019.08.26; Published: 2019.10.15 Abstract DNA replication and sister chromatid cohesion 1 (DSCC1) combines with chromosome transmission-fidelity protein 18 (CTF18) to form a CTF18-DSCC1-CTF8 (CTF18-1-8) module, which in combination with CTF18-replication factor C (RFC) acts as a proliferating cell nuclear antigen (PCNA) loader during DNA replication-associated processes. -
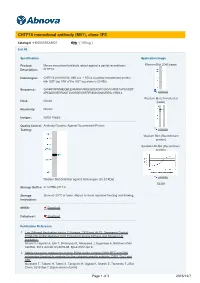
CHTF18 Monoclonal Antibody (M01), Clone 1F5
CHTF18 monoclonal antibody (M01), clone 1F5 Catalog # : H00063922-M01 規格 : [ 100 ug ] List All Specification Application Image Product Mouse monoclonal antibody raised against a partial recombinant Western Blot (Cell lysate) Description: CHTF18. Immunogen: CHTF18 (AAH18184, 886 a.a. ~ 975 a.a) partial recombinant protein with GST tag. MW of the GST tag alone is 26 KDa. Sequence: GVHRPAPRNHEQRLEHIMRRAAREEQPEKDFFGRVVVRSTAVPSAGDT APEQDSVERRMGTAVGRSEVWFRFNEGVSNAVRRSLYIRDLL enlarge Western Blot (Transfected Host: Mouse lysate) Reactivity: Human Isotype: IgG2a Kappa Quality Control Antibody Reactive Against Recombinant Protein. Testing: enlarge Western Blot (Recombinant protein) Sandwich ELISA (Recombinant protein) enlarge Western Blot detection against Immunogen (35.53 KDa) . ELISA Storage Buffer: In 1x PBS, pH 7.4 Storage Store at -20°C or lower. Aliquot to avoid repeated freezing and thawing. Instruction: MSDS: Download Datasheet: Download Publication Reference 1. Two Different Replication Factor C Proteins, Ctf18 and RFC1, Separately Control PCNA-CRL4Cdt2-Mediated Cdt1 Proteolysis during S Phase and following UV Irradiation. Shiomi Y, Hayashi A, Ishii T, Shinmyozu K, Nakayama J, Sugasawa K, Nishitani H.Mol Cell Biol. 2012 Jun;32(12):2279-88. Epub 2012 Apr 9. 2. Stable interaction between the human PCNA loader complex Ctf18-RFC and DNA polymerase {epsilon} is mediated by the cohesion specific subunits, Ctf18, Dcc1 and Ctf8. Murakami T, Takano R, Takeo S, Taniguchi R, Ogawa K, Ohashi E, Tsurimoto T.J Biol Chem. 2010 Sep 7. [Epub ahead of print] Page 1 of 3 2016/12/7 Applications Western Blot (Cell lysate) CHTF18 monoclonal antibody (M01), clone 1F5 Western Blot analysis of CHTF18 expression in HeLa ( Cat # L013V1 ). Protocol Download Western Blot (Transfected lysate) Western Blot analysis of CHTF18 expression in transfected 293T cell line by CHTF18 monoclonal antibody (M01), clone 1F5. -

S-Phase Checkpoint Genes Safeguard High-Fidelity Sister Chromatid Cohesion□D Cheryl D
Molecular Biology of the Cell Vol. 15, 1724–1735, April 2004 S-Phase Checkpoint Genes Safeguard High-Fidelity Sister Chromatid Cohesion□D Cheryl D. Warren,* D. Mark Eckley,* Marina S. Lee,* Joseph S. Hanna,* Adam Hughes,* Brian Peyser,* Chunfa Jie,* Rafael Irizarry,† and Forrest A. Spencer*‡ *McKusick-Nathans Institute of Genetic Medicine, Ross 850, Johns Hopkins University School of Medicine, Baltimore, Maryland 21205; and †Department of Biostatistics, Johns Hopkins University Bloomberg School of Public Health, Baltimore, Maryland 21205 Submitted September 2, 2003; Revised December 10, 2003; Accepted December 23, 2003 Monitoring Editor: Douglas Koshland Cohesion establishment and maintenance are carried out by proteins that modify the activity of Cohesin, an essential complex that holds sister chromatids together. Constituents of the replication fork, such as the DNA polymerase ␣-binding protein Ctf4, contribute to cohesion in ways that are poorly understood. To identify additional cohesion components, we analyzed a ctf4⌬ synthetic lethal screen performed on microarrays. We focused on a subset of ctf4⌬- interacting genes with genetic instability of their own. Our analyses revealed that 17 previously studied genes are also necessary for the maintenance of robust association of sisters in metaphase. Among these were subunits of the MRX complex, which forms a molecular structure similar to Cohesin. Further investigation indicated that the MRX complex did not contribute to metaphase cohesion independent of Cohesin, although an additional role may be contributed by XRS2. In general, results from the screen indicated a sister chromatid cohesion role for a specific subset of genes that function in DNA replication and repair. This subset is particularly enriched for genes that support the S-phase checkpoint. -

Cshperspect-REP-A015727 Table3 1..10
Table 3. Nomenclature for proteins and protein complexes in different organisms Mammals Budding yeast Fission yeast Flies Plants Archaea Bacteria Prereplication complex assembly H. sapiens S. cerevisiae S. pombe D. melanogaster A. thaliana S. solfataricus E. coli Hs Sc Sp Dm At Sso Eco ORC ORC ORC ORC ORC [Orc1/Cdc6]-1, 2, 3 DnaA Orc1/p97 Orc1/p104 Orc1/Orp1/p81 Orc1/p103 Orc1a, Orc1b Orc2/p82 Orc2/p71 Orc2/Orp2/p61 Orc2/p69 Orc2 Orc3/p66 Orc3/p72 Orc3/Orp3/p80 Orc3/Lat/p82 Orc3 Orc4/p50 Orc4/p61 Orc4/Orp4/p109 Orc4/p52 Orc4 Orc5L/p50 Orc5/p55 Orc5/Orp5/p52 Orc5/p52 Orc5 Orc6/p28 Orc6/p50 Orc6/Orp6/p31 Orc6/p29 Orc6 Cdc6 Cdc6 Cdc18 Cdc6 Cdc6a, Cdc6b [Orc1/Cdc6]-1, 2, 3 DnaC Cdt1/Rlf-B Tah11/Sid2/Cdt1 Cdt1 Dup/Cdt1 Cdt1a, Cdt1b Whip g MCM helicase MCM helicase MCM helicase MCM helicase MCM helicase Mcm DnaB Mcm2 Mcm2 Mcm2/Nda1/Cdc19 Mcm2 Mcm2 Mcm3 Mcm3 Mcm3 Mcm3 Mcm3 Mcm4 Mcm4/Cdc54 Mcm4/Cdc21 Mcm4/Dpa Mcm4 Mcm5 Mcm5/Cdc46/Bob1 Mcm5/Nda4 Mcm5 Mcm5 Mcm6 Mcm6 Mcm6/Mis5 Mcm6 Mcm6 Mcm7 Mcm7/Cdc47 Mcm7 Mcm7 Mcm7/Prolifera Gmnn/Geminin Geminin Mcm9 Mcm9 Hbo1 Chm/Hat1 Ham1 Ham2 DiaA Ihfa Ihfb Fis SeqA Replication fork assembly Hs Sc Sp Dm At Sso Eco Mcm8 Rec/Mcm8 Mcm8 Mcm10 Mcm10/Dna43 Mcm10/Cdc23 Mcm10 Mcm10 DDK complex DDK complex DDK complex DDK complex Cdc7 Cdc7 Hsk1 l(1)G0148 Hsk1-like 1 Dbf4/Ask Dbf4 Dfp1/Him1/Rad35 Chif/chiffon Drf1 Continued 2 Replication fork assembly (Continued ) Hs Sc Sp Dm At Sso Eco CDK complex CDK complex CDK complex CDK complex CDK complex Cdk1 Cdc28/Cdk1 Cdc2/Cdk1 Cdc2 CdkA Cdk2 Cdc2c CcnA1, A2 CycA CycA1, A2, -

Identification of Genomic Targets of Krüppel-Like Factor 9 in Mouse Hippocampal
Identification of Genomic Targets of Krüppel-like Factor 9 in Mouse Hippocampal Neurons: Evidence for a role in modulating peripheral circadian clocks by Joseph R. Knoedler A dissertation submitted in partial fulfillment of the requirements for the degree of Doctor of Philosophy (Neuroscience) in the University of Michigan 2016 Doctoral Committee: Professor Robert J. Denver, Chair Professor Daniel Goldman Professor Diane Robins Professor Audrey Seasholtz Associate Professor Bing Ye ©Joseph R. Knoedler All Rights Reserved 2016 To my parents, who never once questioned my decision to become the other kind of doctor, And to Lucy, who has pushed me to be a better person from day one. ii Acknowledgements I have a huge number of people to thank for having made it to this point, so in no particular order: -I would like to thank my adviser, Dr. Robert J. Denver, for his guidance, encouragement, and patience over the last seven years; his mentorship has been indispensable for my growth as a scientist -I would also like to thank my committee members, Drs. Audrey Seasholtz, Dan Goldman, Diane Robins and Bing Ye, for their constructive feedback and their willingness to meet in a frequently cold, windowless room across campus from where they work -I am hugely indebted to Pia Bagamasbad and Yasuhiro Kyono for teaching me almost everything I know about molecular biology and bioinformatics, and to Arasakumar Subramani for his tireless work during the home stretch to my dissertation -I am grateful for the Neuroscience Program leadership and staff, in particular -

CHTF18 Polyclonal Antibody Purified Rabbit Polyclonal Antibody (Pab) Catalog # AP55350
10320 Camino Santa Fe, Suite G San Diego, CA 92121 Tel: 858.875.1900 Fax: 858.622.0609 CHTF18 Polyclonal Antibody Purified Rabbit Polyclonal Antibody (Pab) Catalog # AP55350 Specification CHTF18 Polyclonal Antibody - Product Information Application WB, IHC-P, IHC-F, IF, ICC Primary Accession Q8WVB6 Reactivity Rat Host Rabbit Clonality Polyclonal Calculated MW 107383 CHTF18 Polyclonal Antibody - Additional Information Gene ID 63922 Other Names Chromosome transmission fidelity protein 18 homolog, hCTF18, CHL12, CHTF18, C16orf41, CTF18 Format 0.01M TBS(pH7.4) with 1% BSA, 0.09% (W/V) sodium azide and 50% Glyce Storage Store at -20 ℃ for one year. Avoid repeated freeze/thaw cycles. When reconstituted in sterile pH 7.4 0.01M PBS or diluent of antibody the antibody is stable for at least two weeks at 2-4 ℃. CHTF18 Polyclonal Antibody - Protein Information Name CHTF18 Synonyms C16orf41, CTF18 Function Chromosome cohesion factor involved in sister chromatid cohesion and fidelity of chromosome transmission. Component of one of the cell nuclear antigen loader complexes, CTF18-replication factor C (CTF18-RFC), which consists of CTF18, CTF8, DCC1, RFC2, RFC3, RFC4 and RFC5. Page 1/2 10320 Camino Santa Fe, Suite G San Diego, CA 92121 Tel: 858.875.1900 Fax: 858.622.0609 The CTF18-RFC complex binds to single-stranded and primed DNAs and has weak ATPase activity that is stimulated by the presence of primed DNA, replication protein A (RPA) and by proliferating cell nuclear antigen (PCNA). The CTF18-RFC complex catalyzes the ATP- dependent loading of PCNA onto primed and gapped DNA. Interacts with and stimulates DNA polymerase POLH.