The Transcriptome Difference Between Colorectal Tumor and Normal Tissues Revealed by Single-Cell Sequencing
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Genomics of Inherited Bone Marrow Failure and Myelodysplasia Michael
Genomics of inherited bone marrow failure and myelodysplasia Michael Yu Zhang A dissertation submitted in partial fulfillment of the requirements for the degree of Doctor of Philosophy University of Washington 2015 Reading Committee: Mary-Claire King, Chair Akiko Shimamura Marshall Horwitz Program Authorized to Offer Degree: Molecular and Cellular Biology 1 ©Copyright 2015 Michael Yu Zhang 2 University of Washington ABSTRACT Genomics of inherited bone marrow failure and myelodysplasia Michael Yu Zhang Chair of the Supervisory Committee: Professor Mary-Claire King Department of Medicine (Medical Genetics) and Genome Sciences Bone marrow failure and myelodysplastic syndromes (BMF/MDS) are disorders of impaired blood cell production with increased leukemia risk. BMF/MDS may be acquired or inherited, a distinction critical for treatment selection. Currently, diagnosis of these inherited syndromes is based on clinical history, family history, and laboratory studies, which directs the ordering of genetic tests on a gene-by-gene basis. However, despite extensive clinical workup and serial genetic testing, many cases remain unexplained. We sought to define the genetic etiology and pathophysiology of unclassified bone marrow failure and myelodysplastic syndromes. First, to determine the extent to which patients remained undiagnosed due to atypical or cryptic presentations of known inherited BMF/MDS, we developed a massively-parallel, next- generation DNA sequencing assay to simultaneously screen for mutations in 85 BMF/MDS genes. Querying 71 pediatric and adult patients with unclassified BMF/MDS using this assay revealed 8 (11%) patients with constitutional, pathogenic mutations in GATA2 , RUNX1 , DKC1 , or LIG4 . All eight patients lacked classic features or laboratory findings for their syndromes. -

Somatic Mutations and Progressive Monosomy Modify SAMD9-Related Phenotypes in Humans
Downloaded from http://www.jci.org on June 26, 2017. https://doi.org/10.1172/JCI91913 RESEARCH ARTICLE The Journal of Clinical Investigation Somatic mutations and progressive monosomy modify SAMD9-related phenotypes in humans Federica Buonocore,1 Peter Kühnen,2 Jenifer P. Suntharalingham,1 Ignacio Del Valle,1 Martin Digweed,3 Harald Stachelscheid,4 Noushafarin Khajavi,2 Mohammed Didi,5 Angela F. Brady,6 Oliver Blankenstein,2 Annie M. Procter,7 Paul Dimitri,8 Jerry K.H. Wales,9 Paolo Ghirri,10 Dieter Knöbl,11 Brigitte Strahm,12 Miriam Erlacher,12,13 Marcin W. Wlodarski,12,13 Wei Chen,14 George K. Kokai,15 Glenn Anderson,16 Deborah Morrogh,17 Dale A. Moulding,18 Shane A. McKee,19 Charlotte M. Niemeyer,12,13 Annette Grüters,2 and John C. Achermann1 1Genetics and Genomic Medicine, University College London (UCL) Great Ormond Street Institute of Child Health, London, United Kingdom. 2Institute of Experimental Pediatric Endocrinology and Department of Pediatric Endocrinology, Charité, Berlin, Germany. 3Department of Human and Medical Genetics, Charité, Berlin, Germany. 4Berlin Institute of Health, Berlin, Germany, and Berlin-Brandenburg Centrum for Regenerative Therapies, Charité, Berlin, Germany. 5Department of Paediatric Endocrinology, Alder Hey Children’s NHS Foundation Trust, Liverpool, United Kingdom. 6North West Thames Regional Genetics Service, Northwick Park Hospital, Harrow, United Kingdom. 7Institute of Medical Genetics, University Hospital of Wales, Cardiff, United Kingdom. 8Academic Unit of Child Health, University of Sheffield, Sheffield, United Kingdom. 9Department of Endocrinology, Children’s Health Queensland Clinical Unit, University of Queensland, Brisbane, Australia. 10Department of Neonatology, University of Pisa, Pisa, Italy. 11Pediatric Endocrinology, Karlsruhe, Germany. -

Of Small Intestine Harboring Driver Gene Mutations: a Case Report and a Literature Review
1161 Case Report A rare multiple primary sarcomatoid carcinoma (SCA) of small intestine harboring driver gene mutations: a case report and a literature review Zhu Zhu1#, Xinyi Liu2#, Wenliang Li1, Zhengqi Wen1, Xiang Ji1, Ruize Zhou1, Xiaoyu Tuo3, Yaru Chen2, Xian Gong2, Guifeng Liu2, Yanqing Zhou2, Shifu Chen2, Lele Song2#^, Jian Huang1 1Department of Oncology, First Affiliated Hospital of Kunming Medical University, Kunming, China; 2HaploX Biotechnology, Shenzhen, China; 3Department of Pathology, First Affiliated Hospital of Kunming Medical University, Kunming, China #These authors contributed equally to this work. Correspondence to: Jian Huang. Department of Oncology, First Affiliated Hospital of Kunming Medical University, No. 295, Xichang Road, Kunming 560032, Yunnan Province, China. Email: [email protected]; Lele Song. HaploX Biotechnology, 8th floor, Auto Electric Power Building, Songpingshan Road, Nanshan District, Shenzhen 518057, Guangdong Province, China. Email: [email protected]. Abstract: Primary sarcomatoid carcinoma (SCA) is a type of rare tumor consisting of both malignant epithelial and mesenchymal components. Only 32 cases of SCA of the small bowel have been reported in the literature to date. Due to its rarity and complexity, this cancer has not been genetically studied and its diagnosis and treatment remain difficult. Here we report a 54-year-old male underwent emergency surgical resection in the small intestine due to severe obstruction and was diagnosed with multiple SCA based on postoperative pathological examination. Over 100 polypoid tumors scattered along his whole jejunum and proximal ileum. Chemotherapy (IFO+Epirubicin) was performed after surgery while the patient died two months after the surgery due to severe malnutrition. Whole-exome sequencing was performed for the tumor tissue with normal tissue as the control. -

Somatic Mutations and Progressive Monosomy Modify SAMD9-Related Phenotypes in Humans
Downloaded from http://www.jci.org on June 5, 2017. https://doi.org/10.1172/JCI91913 RESEARCH ARTICLE The Journal of Clinical Investigation Somatic mutations and progressive monosomy modify SAMD9-related phenotypes in humans Federica Buonocore,1 Peter Kühnen,2 Jenifer P. Suntharalingham,1 Ignacio Del Valle,1 Martin Digweed,3 Harald Stachelscheid,4 Noushafarin Khajavi,2 Mohammed Didi,5 Angela F. Brady,6 Oliver Blankenstein,2 Annie M. Procter,7 Paul Dimitri,8 Jerry K.H. Wales,9 Paolo Ghirri,10 Dieter Knöbl,11 Brigitte Strahm,12 Miriam Erlacher,12,13 Marcin W. Wlodarski,12,13 Wei Chen,14 George K. Kokai,15 Glenn Anderson,16 Deborah Morrogh,17 Dale A. Moulding,18 Shane A. McKee,19 Charlotte M. Niemeyer,12,13 Annette Grüters,2 and John C. Achermann1 1Genetics and Genomic Medicine, University College London (UCL) Great Ormond Street Institute of Child Health, London, United Kingdom. 2Institute of Experimental Pediatric Endocrinology and Department of Pediatric Endocrinology, Charité, Berlin, Germany. 3Department of Human and Medical Genetics, Charité, Berlin, Germany. 4Berlin Institute of Health, Berlin, Germany, and Berlin-Brandenburg Centrum for Regenerative Therapies, Charité, Berlin, Germany. 5Department of Paediatric Endocrinology, Alder Hey Children’s NHS Foundation Trust, Liverpool, United Kingdom. 6North West Thames Regional Genetics Service, Northwick Park Hospital, Harrow, United Kingdom. 7Institute of Medical Genetics, University Hospital of Wales, Cardiff, United Kingdom. 8Academic Unit of Child Health, University of Sheffield, Sheffield, United Kingdom. 9Department of Endocrinology, Children’s Health Queensland Clinical Unit, University of Queensland, Brisbane, Australia. 10Department of Neonatology, University of Pisa, Pisa, Italy. 11Pediatric Endocrinology, Karlsruhe, Germany. -

Genome-Wide Association Study for Feed Efficiency and Growth Traits in U.S. Beef Cattle Christopher M
Seabury et al. BMC Genomics (2017) 18:386 DOI 10.1186/s12864-017-3754-y RESEARCH ARTICLE Open Access Genome-wide association study for feed efficiency and growth traits in U.S. beef cattle Christopher M. Seabury1*, David L. Oldeschulte1, Mahdi Saatchi2, Jonathan E. Beever3, Jared E. Decker4,5, Yvette A. Halley1, Eric K. Bhattarai1, Maral Molaei1, Harvey C. Freetly6, Stephanie L. Hansen2, Helen Yampara-Iquise4, Kristen A. Johnson7, Monty S. Kerley4, JaeWoo Kim4, Daniel D. Loy2, Elisa Marques8, Holly L. Neibergs7, Robert D. Schnabel4,5, Daniel W. Shike3, Matthew L. Spangler9, Robert L. Weaber10, Dorian J. Garrick2,11 and Jeremy F. Taylor4* Abstract Background: Single nucleotide polymorphism (SNP) arrays for domestic cattle have catalyzed the identification of genetic markers associated with complex traits for inclusion in modern breeding and selection programs. Using actual and imputed Illumina 778K genotypes for 3887 U.S. beef cattle from 3 populations (Angus, Hereford, SimAngus), we performed genome-wide association analyses for feed efficiency and growth traits including average daily gain (ADG), dry matter intake (DMI), mid-test metabolic weight (MMWT), and residual feed intake (RFI), with marker-based heritability estimates produced for all traits and populations. Results: Moderate and/or large-effect QTL were detected for all traits in all populations, as jointly defined by the estimated proportion of variance explained (PVE) by marker effects (PVE ≥ 1.0%) and a nominal P-value threshold (P ≤ 5e-05). Lead SNPs with PVE ≥ 2.0% were considered putative evidence of large-effect QTL (n = 52), whereas those with PVE ≥ 1.0% but < 2.0% were considered putative evidence for moderate-effect QTL (n = 35). -

Early Mechanisms of Retinal Degeneration in the Harlequin Mouse
Western University Scholarship@Western Electronic Thesis and Dissertation Repository 7-17-2014 12:00 AM Early Mechanisms of Retinal Degeneration in the harlequin Mouse Eric Dolinar The University of Western Ontario Supervisor Dr. Kathleen Hill The University of Western Ontario Graduate Program in Biology A thesis submitted in partial fulfillment of the equirr ements for the degree in Master of Science © Eric Dolinar 2014 Follow this and additional works at: https://ir.lib.uwo.ca/etd Part of the Biology Commons, Genetics Commons, and the Molecular Genetics Commons Recommended Citation Dolinar, Eric, "Early Mechanisms of Retinal Degeneration in the harlequin Mouse" (2014). Electronic Thesis and Dissertation Repository. 2156. https://ir.lib.uwo.ca/etd/2156 This Dissertation/Thesis is brought to you for free and open access by Scholarship@Western. It has been accepted for inclusion in Electronic Thesis and Dissertation Repository by an authorized administrator of Scholarship@Western. For more information, please contact [email protected]. i Early Mechanisms of Retinal Degeneration in the harlequin Mouse (Thesis Format: Monograph) by Eric A Dolinar Graduate Program in Biology A thesis submitted in partial fulfillment of the requirements for the degree of Master of Science The School of Graduate and Postdoctoral Studies The University of Western Ontario, London, Ontario, Canada © Eric A Dolinar 2014 i ii Abstract Retinal diseases are personally debilitating and expensive, yet many early disease mechanisms leading to their onset and progression remain poorly understood. The harlequin mouse is a model of human mitochondrial dysfunction and parainflammation leading to subsequent cerebellar and retinal degeneration. Diagnosis of retinal degeneration can be tracked in vivo and is associated with AIF dysfunction. -
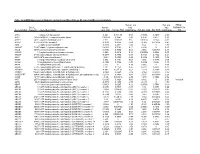
Mrna Expression in Human Leiomyoma and Eker Rats As Measured by Microarray Analysis
Table 3S: mRNA Expression in Human Leiomyoma and Eker Rats as Measured by Microarray Analysis Human_avg Rat_avg_ PENG_ Entrez. Human_ log2_ log2_ RAPAMYCIN Gene.Symbol Gene.ID Gene Description avg_tstat Human_FDR foldChange Rat_avg_tstat Rat_FDR foldChange _DN A1BG 1 alpha-1-B glycoprotein 4.982 9.52E-05 0.68 -0.8346 0.4639 -0.38 A1CF 29974 APOBEC1 complementation factor -0.08024 0.9541 -0.02 0.9141 0.421 0.10 A2BP1 54715 ataxin 2-binding protein 1 2.811 0.01093 0.65 0.07114 0.954 -0.01 A2LD1 87769 AIG2-like domain 1 -0.3033 0.8056 -0.09 -3.365 0.005704 -0.42 A2M 2 alpha-2-macroglobulin -0.8113 0.4691 -0.03 6.02 0 1.75 A4GALT 53947 alpha 1,4-galactosyltransferase 0.4383 0.7128 0.11 6.304 0 2.30 AACS 65985 acetoacetyl-CoA synthetase 0.3595 0.7664 0.03 3.534 0.00388 0.38 AADAC 13 arylacetamide deacetylase (esterase) 0.569 0.6216 0.16 0.005588 0.9968 0.00 AADAT 51166 aminoadipate aminotransferase -0.9577 0.3876 -0.11 0.8123 0.4752 0.24 AAK1 22848 AP2 associated kinase 1 -1.261 0.2505 -0.25 0.8232 0.4689 0.12 AAMP 14 angio-associated, migratory cell protein 0.873 0.4351 0.07 1.656 0.1476 0.06 AANAT 15 arylalkylamine N-acetyltransferase -0.3998 0.7394 -0.08 0.8486 0.456 0.18 AARS 16 alanyl-tRNA synthetase 5.517 0 0.34 8.616 0 0.69 AARS2 57505 alanyl-tRNA synthetase 2, mitochondrial (putative) 1.701 0.1158 0.35 0.5011 0.6622 0.07 AARSD1 80755 alanyl-tRNA synthetase domain containing 1 4.403 9.52E-05 0.52 1.279 0.2609 0.13 AASDH 132949 aminoadipate-semialdehyde dehydrogenase -0.8921 0.4247 -0.12 -2.564 0.02993 -0.32 AASDHPPT 60496 aminoadipate-semialdehyde -

Constitutional SAMD9L Mutations Cause Familial Myelodysplastic Syndrome and Transient Monosomy 7
Constitutional SAMD9L mutations cause familial myelodysplastic syndrome and transient monosomy 7 The Harvard community has made this article openly available. Please share how this access benefits you. Your story matters Citation Pastor, V. B., S. S. Sahoo, J. Boklan, G. C. Schwabe, E. Saribeyoglu, B. Strahm, D. Lebrecht, et al. 2018. “Constitutional SAMD9L mutations cause familial myelodysplastic syndrome and transient monosomy 7.” Haematologica 103 (3): 427-437. doi:10.3324/haematol.2017.180778. http://dx.doi.org/10.3324/ haematol.2017.180778. Published Version doi:10.3324/haematol.2017.180778 Citable link http://nrs.harvard.edu/urn-3:HUL.InstRepos:35982102 Terms of Use This article was downloaded from Harvard University’s DASH repository, and is made available under the terms and conditions applicable to Other Posted Material, as set forth at http:// nrs.harvard.edu/urn-3:HUL.InstRepos:dash.current.terms-of- use#LAA Myelodysplastic Syndromes ARTICLE Constitutional SAMD9L mutations cause familial myelodysplastic syndrome and transient Ferrata Storti monosomy 7 Foundation Victor B. Pastor,1,2* Sushree S. Sahoo,1,2,3* Jessica Boklan,4 Georg C. Schwabe,5 Ebru Saribeyoglu,5 Brigitte Strahm,1 Dirk Lebrecht,1 Matthias Voss,6 Yenan T. Bryceson,6 Miriam Erlacher,1,7 Gerhard Ehninger,8 Marena Niewisch,1 Brigitte Schlegelberger,9 Irith Baumann,10 John C. Achermann,11 Akiko Shimamura,12 Jochen Hochrein,13 Ulf Tedgård,14 Lars Nilsson,15 Henrik Hasle,16 Melanie Boerries,7,13 Hauke Busch,13,17 Charlotte M. Niemeyer1,7 Haematologica 2018 and Marcin -

Genome-Wide Association Study for Feed Efficiency and Growth Traits in U.S
University of Nebraska - Lincoln DigitalCommons@University of Nebraska - Lincoln Faculty Papers and Publications in Animal Science Animal Science Department 2017 Genome-wide association study for feed efficiency and owthgr traits in U.S. beef cattle Christopher M. Seabury Texas A&M University, [email protected] David L. Oldeschulte Texas A&M University, College Station, [email protected] Mahdi Saatchi Iowa State University, [email protected] Jonathan E. Beever University of Illinois at Urbana-Champaign, [email protected] Jared E. Decker University of Missouri, Columbia, [email protected] See next page for additional authors Follow this and additional works at: https://digitalcommons.unl.edu/animalscifacpub Part of the Genetics and Genomics Commons, and the Meat Science Commons Seabury, Christopher M.; Oldeschulte, David L.; Saatchi, Mahdi; Beever, Jonathan E.; Decker, Jared E.; Halley, Yvette A.; Bhattarai, Eric K.; Molael, Maral; Freetly, Harvey; Hansen, Stephanie L.; Yampara-Iquise, Helen; Johnson, Kristen A.; Kerley, Monty S.; Kim, JaeWoo; Loy, Daniel D.; Marques, Elisa; Neibergs, Holly L.; Schnabel, Robert D.; Shike, Daniel W.; Spangler, Matthew L.; Weaber, Bob; Garrick, D. J.; and Taylor, Jeremy F., "Genome-wide association study for feed efficiency and owthgr traits in U.S. beef cattle" (2017). Faculty Papers and Publications in Animal Science. 993. https://digitalcommons.unl.edu/animalscifacpub/993 This Article is brought to you for free and open access by the Animal Science Department at DigitalCommons@University of Nebraska - Lincoln. It has been accepted for inclusion in Faculty Papers and Publications in Animal Science by an authorized administrator of DigitalCommons@University of Nebraska - Lincoln. Authors Christopher M. -

Systematic Expression Analysis of the Diagnostic and Prognostic Value of HEPACAM Family Member 2 in Colon Adenocarcinoma
Systematic Expression Analysis of the Diagnostic and Prognostic Value of HEPACAM Family Member 2 in Colon Adenocarcinoma Shuai Wang Guangxi Medical University First Aliated Hospital Guo-Tian Ruan Guangxi Medical University First Aliated Hospital Yi-Zhen Gong Guilin Medical University Aliated Hospital Cun Liao Guangxi Medical University First Aliated Hospital Lei Zhang Guilin Medical University Aliated Hospital Feng Gao ( [email protected] ) Guangxi Medical University First Aliated Hospital https://orcid.org/0000-0002-1000-385X Research article Keywords: HEPACAM2, Guangxi cohort, COAD, prognosis, diagnosis Posted Date: January 29th, 2021 DOI: https://doi.org/10.21203/rs.3.rs-154487/v1 License: This work is licensed under a Creative Commons Attribution 4.0 International License. Read Full License Page 1/19 Abstract Background: The diagnostic and prognostic value of HEPACAM family member 2 (HEPACAM2) gene in patients with colon adenocarcinoma (COAD) is rarely reported. Therefore, the purpose of this study is to explore the diagnostic and prognostic value of HEPACAM2 gene in patients with COAD. Methods: Firstly, we analyzed the differential expression levels of HEPACAM2 gene and diagnostic value analysis from different databases. Secondly, univariate and multivariate survival analysis of the prognostic value of HEPACAM2 gene in patients with COAD was performed. Finally, utilizing joint-effects analysis and comprehensive prognosis analysis to investigate the prognostic value of HEPACAM2 and related genes. Results: Differential analyses of multiple databases showed that the HEPACAM2 expression level in COAD tumor tissue was signicantly lower than that of adjacent normal tissues. The diagnostic ROC curve results indicated that HEPACAM2 gene had a higher diagnostic value in COAD. -

SAMD9-Related Phenotypes in Humans
RESEARCH ARTICLE The Journal of Clinical Investigation Somatic mutations and progressive monosomy modify SAMD9-related phenotypes in humans Federica Buonocore,1 Peter Kühnen,2 Jenifer P. Suntharalingham,1 Ignacio Del Valle,1 Martin Digweed,3 Harald Stachelscheid,4 Noushafarin Khajavi,2 Mohammed Didi,5 Angela F. Brady,6 Oliver Blankenstein,2 Annie M. Procter,7 Paul Dimitri,8 Jerry K.H. Wales,9 Paolo Ghirri,10 Dieter Knöbl,11 Brigitte Strahm,12 Miriam Erlacher,12,13 Marcin W. Wlodarski,12,13 Wei Chen,14 George K. Kokai,15 Glenn Anderson,16 Deborah Morrogh,17 Dale A. Moulding,18 Shane A. McKee,19 Charlotte M. Niemeyer,12,13 Annette Grüters,2 and John C. Achermann1 1Genetics and Genomic Medicine, University College London (UCL) Great Ormond Street Institute of Child Health, London, United Kingdom. 2Institute of Experimental Pediatric Endocrinology and Department of Pediatric Endocrinology, Charité, Berlin, Germany. 3Department of Human and Medical Genetics, Charité, Berlin, Germany. 4Berlin Institute of Health, Berlin, Germany, and Berlin-Brandenburg Centrum for Regenerative Therapies, Charité, Berlin, Germany. 5Department of Paediatric Endocrinology, Alder Hey Children’s NHS Foundation Trust, Liverpool, United Kingdom. 6North West Thames Regional Genetics Service, Northwick Park Hospital, Harrow, United Kingdom. 7Institute of Medical Genetics, University Hospital of Wales, Cardiff, United Kingdom. 8Academic Unit of Child Health, University of Sheffield, Sheffield, United Kingdom. 9Department of Endocrinology, Children’s Health Queensland Clinical Unit, University of Queensland, Brisbane, Australia. 10Department of Neonatology, University of Pisa, Pisa, Italy. 11Pediatric Endocrinology, Karlsruhe, Germany. 12Department of Pediatrics and Adolescent Medicine, Division of Pediatric Hematology and Oncology, Medical Center University of Freiburg, Faculty of Medicine, University of Freiburg, Freiburg, Germany. -

Surgical and Molecular Characterization of Primary and Metastatic Disease in a Neuroendocrine Tumor Arising in a Tailgut Cyst
Downloaded from molecularcasestudies.cshlp.org on September 26, 2021 - Published by Cold Spring Harbor Laboratory Press TITLE Surgical and molecular characterization of primary and metastatic disease in a neuroendocrine tumor arising in a tailgut cyst RUNNING TITLE Neuroendocrine tumor in a tailgut cyst AUTHORS J Erdrich*, KB Schaberg*, MS Khodadoust, L Zhou, AA Shelton, BC Visser, JM Ford, AA Alizadeh, SR Quake, PL Kunz, JF Beausang** *contributed equally; ** corresponding author: ([email protected]) AUTHOR AFFILIATIONS and FUNDING *Jennifer Erdrich, MD <[email protected]>: Department of Surgery, Stanford University School of Medicine Current position: Department of Surgery, Cedars Sinai Medical Center *Kurt B. Schaberg, MD <[email protected]>: Department of Pathology, Stanford University School of Medicine Current position: Department of Pathology, University of Kentucky Michael S. Khodadoust, MD, PhD <[email protected]>: Department of Medicine/Oncology, Stanford University School of Medicine Li Zhou <[email protected]> Stanford Cancer Institute, Stanford University, Stanford, California, USA Andrew A. Shelton, MD <[email protected]>: Department of Surgery, Stanford University School of Medicine Brendan C Visser, MD <[email protected]>: Department of Surgery, Stanford University School of Medicine James M Ford, MD <[email protected]>: Department of medicine/oncology and Department of Genetics; Stanford University School of Medicine Ash A. Alizadeh MD, PhD <[email protected]>: Department of Medicine/Oncology; Stanford University School of Medicine Stephen R. Quake DPhil<[email protected]>: Department of Bioengineering and Department of Applied Physics; Stanford University and Chan Zuckerberg Biohub Funding: Howard Hughes Medical Institute Pamela L. Kunz MD <[email protected]>: Department of Medicine/Oncology; Stanford University School of Medicine 1 of 13 Downloaded from molecularcasestudies.cshlp.org on September 26, 2021 - Published by Cold Spring Harbor Laboratory Press **John F.