CDKN2B Deletion Is Essential for Pancreatic Cancer Development Instead of Unmeaningful Co-Deletion Due to Juxtaposition to CDKN2A
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

CDKN2A and CDKN2B Methylation in Coronary Heart Disease Cases and Controls
EXPERIMENTAL AND THERAPEUTIC MEDICINE 14: 6093-6098, 2016 CDKN2A and CDKN2B methylation in coronary heart disease cases and controls JINYAN ZHONG1,2*, XIAOYING CHEN3*, HUADAN YE3, NAN WU1,3, XIAOMIN CHEN1 and SHIWEI DUAN3 1Cardiology Center, Ningbo First Hospital, Ningbo University; 2Department of Cardiology, Ningbo Second Hospital, Ningbo, Zhejiang 315010; 3Medical Genetics Center, School of Medicine, Ningbo University, Ningbo, Zhejiang 315211, P.R. China Received May 27, 2016; Accepted March 24, 2017 DOI: 10.3892/etm.2017.5310 Abstract. The aim of the present study was to investigate frequencies were significantly associated with age, and there the association between cyclin-dependent kinase inhibitor was a gender dimorphism in CDKN2B methylation. 2A (CDKN2A) and cyclin-dependent kinase inhibitor 2B (CDKN2B) methylation, and coronary heart disease (CHD), Introduction and to explore the interaction between methylation status and CHD clinical characteristics in Han Chinese patients. A total Coronary heart disease (CHD) is a complex chronic disease of 189 CHD (96 males, 93 females) and 190 well-matched that is caused by an imbalance between blood supply and non-CHD controls (96 males, 94 females) were recruited for demand in myocardium. Various environmental and genetic the study. Methylation-specific polymerase chain reaction factors are known to contribute to onset and development of technology was used to examine gene promoter methylation CHD (1). As of 2010, CHD was the leading cause of mortality status. Comparisons of methylation frequencies between CHD globally, resulting in over 7 million cases of mortality (2). and non-CHD patients were carried out using the Chi-square Therefore, association studies for CHD biomarkers have been test. -

Epigenetic Silencing of CDKN1A and CDKN2B by SNHG1 Promotes The
Li et al. Cell Death and Disease (2020) 11:823 https://doi.org/10.1038/s41419-020-03031-6 Cell Death & Disease ARTICLE Open Access Epigenetic silencing of CDKN1A and CDKN2B by SNHG1 promotes the cell cycle, migration and epithelial-mesenchymal transition progression of hepatocellular carcinoma Bei Li1,AngLi2, Zhen You 1,JingchangXu1 and Sha Zhu3 Abstract Enhanced SNHG1 (small nucleolar RNA host gene 1) expression has been found to play a critical role in the initiation and progression of hepatocellular carcinoma (HCC) with its detailed mechanism largely unknown. In this study, we show that SNHG1 promotes the HCC progression through epigenetically silencing CDKN1A and CDKN2B in the nucleus, and competing with CDK4 mRNA for binding miR-140-5p in the cytoplasm. Using bioinformatics analyses, we found hepatocarcinogenesis is particularly associated with dysregulated expression of SNHG1 and activation of the cell cycle pathway. SNHG1 was upregulated in HCC tissues and cells, and its knockdown significantly inhibited HCC cell cycle, growth, metastasis, and epithelial–mesenchymal transition (EMT) both in vitro and in vivo. Chromatin immunoprecipitation and RNA immunoprecipitation assays demonstrate that SNHG1 inhibit the transcription of CDKN1A and CDKN2B through enhancing EZH2 mediated-H3K27me3 in the promoter of CDKN1A and CDKN2B, thus resulting in the de-repression of the cell cycle. Dual-luciferase assay and RNA pulldown revealed that SNHG1 promotes 1234567890():,; 1234567890():,; 1234567890():,; 1234567890():,; the expression of CDK4 by competitively binding to miR-140-5p. In conclusion, we propose that SNHG1 formed a regulatory network to confer an oncogenic function in HCC and SNHG1 may serve as a potential target for HCC diagnosis and treatment. -

Infrequent Methylation of CDKN2A(Mts1p16) and Rare Mutation of Both CDKN2A and CDKN2B(Mts2ip15) in Primary Astrocytic Tumours
British Joumal of Cancer (1997) 75(1), 2-8 © 1997 Cancer Research Campaign Infrequent methylation of CDKN2A(MTS1p16) and rare mutation of both CDKN2A and CDKN2B(MTS2Ip15) in primary astrocytic tumours EE Schmidt, K Ichimura, KR Messerle, HM Goike and VP Collins Institute for Oncology and Pathology, Division of Tumour Pathology, and Ludwig Institute for Cancer Research, Stockholm Branch, Karolinska Hospital, S-171 76 Stockholm, Sweden Summary In a series of 46 glioblastomas, 16 anaplastic astrocytomas and eight astrocytomas, all tumours retaining one or both alleles of CDKN2A (48 tumours) and CDKN2B (49 tumours) were subjected to sequence analysis (entire coding region and splice acceptor and donor sites). One glioblastoma with hemizygous deletion of CDKN2A showed a missense mutation in exon 2 (codon 83) that would result in the substitution of tyrosine for histidine in the protein. None of the tumours retaining alleles of CDKN2B showed mutations of this gene. Glioblastomas with retention of both alleles of CDKN2A (14 tumours) and CDKN2B (16 tumours) expressed transcripts for these genes. In contrast, 7/13 glioblastomas with hemizygous deletions of CDKN2A and 8/11 glioblastomas with hemizygous deletions of CDKN2B showed no or weak expression. Anaplastic astrocytomas and astrocytomas showed a considerable variation in the expression of both genes, regardless of whether they retained one or two copies of the genes. The methylation status of the 5' CpG island of the CDKN2A gene was studied in all 15 tumours retaining only one allele of CDKN2A as well as in the six tumours showing no significant expression of transcript despite their retaining both CDKN2A alleles. -

A Variant at 9P21.3 Functionally Implicates CDKN2B in Paediatric B-Cell Precursor Acute Lymphoblastic Leukaemia Aetiology
UCSF UC San Francisco Previously Published Works Title A variant at 9p21.3 functionally implicates CDKN2B in paediatric B-cell precursor acute lymphoblastic leukaemia aetiology. Permalink https://escholarship.org/uc/item/3bz9m1pp Journal Nature communications, 7(1) ISSN 2041-1723 Authors Hungate, Eric A Vora, Sapana R Gamazon, Eric R et al. Publication Date 2016-02-12 DOI 10.1038/ncomms10635 Peer reviewed eScholarship.org Powered by the California Digital Library University of California ARTICLE Received 23 Apr 2015 | Accepted 7 Jan 2016 | Published 12 Feb 2016 DOI: 10.1038/ncomms10635 OPEN A variant at 9p21.3 functionally implicates CDKN2B in paediatric B-cell precursor acute lymphoblastic leukaemia aetiology Eric A. Hungate1,*, Sapana R. Vora2,*, Eric R. Gamazon3,4, Takaya Moriyama5, Timothy Best2, Imge Hulur6, Younghee Lee3, Tiffany-Jane Evans7, Eva Ellinghaus8, Martin Stanulla9,Je´remie Rudant10,11, Laurent Orsi10,11, Jacqueline Clavel10,11,12, Elizabeth Milne13, Rodney J. Scott7,14, Ching-Hon Pui15, Nancy J. Cox3, Mignon L. Loh16, Jun J. Yang5, Andrew D. Skol1 & Kenan Onel1 Paediatric B-cell precursor acute lymphoblastic leukaemia (BCP-ALL) is the most common cancer of childhood, yet little is known about BCP-ALL predisposition. In this study, in 2,187 cases of European ancestry and 5,543 controls, we discover and replicate a locus indexed by À 15 rs77728904 at 9p21.3 associated with BCP-ALL susceptibility (Pcombined ¼ 3.32 Â 10 , OR ¼ 1.72) and independent from rs3731217, the previously reported ALL-associated variant in this region. Of correlated SNPs tagged by this locus, only rs662463 is significant in African Americans, suggesting it is a plausible causative variant. -
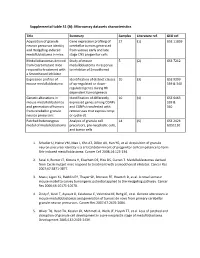
Supplemental Table S1 (A): Microarray Datasets Characteristics
Supplemental table S1 (A): Microarray datasets characteristics Title Summary Samples Literature ref. GEO ref. Acquisition of granule Gene expression profiling of 27 (1) GSE 11859 neuron precursor identity cerebellar tumors generated and Hedgehog‐induced from various early and late medulloblastoma in mice. stage CNS progenitor cells Medulloblastomas derived Study of mouse 5 (2) GSE 7212 from Cxcr6 mutant mice medulloblastoma in response respond to treatment with to inhibitor of Smoothened a Smoothened inhibitor Expression profiles of Identification of distinct classes 10 (3) GSE 9299 mouse medulloblastoma of up‐regulated or down‐ 339 & 340 regulated genes during Hh dependent tumorigenesis Genetic alterations in Identification of differently 10 (4) GSE 6463 mouse medulloblastomas expressed genes among CGNPs 339 & and generation of tumors and CGNPs transfected with 340 from cerebellar granule retroviruses that express nmyc neuron precursors or cyclin‐d1 Patched heterozygous Analysis of granule cell 14 (5) GSE 2426 model of medulloblastoma precursors, pre‐neoplastic cells, GDS1110 and tumor cells 1. Schuller U, Heine VM, Mao J, Kho AT, Dillon AK, Han YG, et al. Acquisition of granule neuron precursor identity is a critical determinant of progenitor cell competence to form Shh‐induced medulloblastoma. Cancer Cell 2008;14:123‐134. 2. Sasai K, Romer JT, Kimura H, Eberhart DE, Rice DS, Curran T. Medulloblastomas derived from Cxcr6 mutant mice respond to treatment with a smoothened inhibitor. Cancer Res 2007;67:3871‐3877. 3. Mao J, Ligon KL, Rakhlin EY, Thayer SP, Bronson RT, Rowitch D, et al. A novel somatic mouse model to survey tumorigenic potential applied to the Hedgehog pathway. Cancer Res 2006;66:10171‐10178. -

Deregulation of the P14arf/MDM2/P53 Pathway Is A
[CANCER RESEARCH 60, 417–424, January 15, 2000] Deregulation of the p14ARF/MDM2/p53 Pathway Is a Prerequisite for Human 1 Astrocytic Gliomas with G1-S Transition Control Gene Abnormalities Koichi Ichimura, Maria Bondesson Bolin,2 Helena M. Goike, Esther E. Schmidt, Ahmad Moshref, and V. Peter Collins3 Department of Pathology, Division of Molecular Histopathology, University of Cambridge, Addenbrooke’s Hospital, Cambridge CB2 2QQ, United Kingdom [K. I., E. E. S., V. P. C], and Ludwig Institute for Cancer Research and Unit of Tumorpathology, Department of Oncology and Pathology, Karolinska Hospital, 171 76 Stockholm, Sweden [K. I., M. B. B., H. M. G., E. E. S., A. M., V. P. C.] ABSTRACT There are several lines of evidence to indicate that deregulation of G1-S transition control alone may be detrimental for cell survival: (a) Deregulation of G -S transition control in cell cycle is one of the 1 transfection of rodent cells with the adenoviral E1A and E1B genes important mechanisms in the development of human tumors including resulted in transformation due to the viral proteins binding to and astrocytic gliomas. We have previously reported that approximately two- inactivating pRB and p53. Transfection with E1A alone induced p53 thirds of glioblastomas (GBs) had abnormalities of G1-S transition control either by mutation/homozygous deletion of RB1 or CDKN2A (p16INK4A), dependent apoptosis (10); (b) when E2F1 was ectopically expressed in or amplification of CDK4 (K. Ichimura et al., Oncogene, 13: 1065–1072, rodent embryo fibroblasts, although the cell indeed entered S phase 1996). However, abnormalities of G1-S transition control genes may in- (11), apoptosis was induced in a p53-dependent manner (12); (c) loss duce p53-dependent apoptosis in cells. -

Cyclin E1 and Rb Modulation As Common Events at Time of Resistance to Palbociclib in Hormone Receptor-Positive Breast Cancer
www.nature.com/npjbcancer ARTICLE OPEN Cyclin E1 and Rb modulation as common events at time of resistance to palbociclib in hormone receptor-positive breast cancer Cristina Guarducci1, Martina Bonechi1, Matteo Benelli2, Chiara Biagioni2,3, Giulia Boccalini1, Dario Romagnoli2, Roberto Verardo4, Rachel Schiff5,6,7,8, C. Kent Osborne5,6,7,8, Carmine De Angelis5,6,8, Angelo Di Leo3, Luca Malorni1,3 and Ilenia Migliaccio1 CDK4/6 inhibitors represent a new treatment standard for hormone receptor-positive (HR+), HER2-negative advanced breast cancer (BC) patients. Although efficacious, resistance to these agents is universal. Here, we profiled a large panel of HR+ BC cell lines with conditioned resistance to the CDK4/6 inhibitor palbociclib, and analyzed cell cycle-related markers by gene expression profiles (GEP) and western blot (WB). GEP showed high molecular heterogeneity among the models, with E2F targets being significantly enriched both during treatment and at the time of resistance. By both WB and GEP, a common molecular feature at the time of palbociclib resistance was the concomitant overexpression of cyclin E1 and down-regulation of Rb. CCNE1 was the only significantly up-regulated gene among E2F targets at resistance with CCNE1 genomic amplification being observed in two resistant models. Rb was downregulated in all resistant models; a reduction of RB1 copy number was observed in three resistant cell lines. In silico analyses showed that CCNE1/RB1 ratio correlated with palbociclib IC50 in different datasets of both breast and non-breast cancer cell lines, performing better than CCNE1 or RB1 taken separately. Finally, the CCNE1/RB1 ratio was shown to be an adverse prognostic factor in patients with ER+ BC and to be able to discriminate palbociclib-sensitive versus resistant among patients enrolled in the NeoPalAna trial, a neoadjuvant trial testing palbociclib, performing better than CCNE1 or RB1 alone. -

Neddylation Mediates Ventricular Chamber Maturation PNAS PLUS Through Repression of Hippo Signaling
Neddylation mediates ventricular chamber maturation PNAS PLUS through repression of Hippo signaling Jianqiu Zoua, Wenxia Maa, Jie Lia, Rodney Littlejohna, Hongyi Zhoub, Il-man Kima, David J. R. Fultona, Weiqin Chenb, Neal L. Weintrauba, Jiliang Zhouc, and Huabo Sua,c,d,1 aVascular Biology Center, Medical College of Georgia, Augusta University, Augusta, GA 30912; bDepartment of Physiology, Medical College of Georgia, Augusta University, Augusta, GA 30912; cDepartment of Pharmacology and Toxicology, Medical College of Georgia, Augusta University, Augusta, GA 30912; and dProtein Modification and Degradation Lab, School of Basic Medical Sciences, Guangzhou Medical University, Guangzhou, Guangdong 510260, China Edited by Deepak Srivastava, Gladstone Institute of Cardiovascular Disease, San Francisco, CA, and accepted by Editorial Board Member Brenda A. Schulman March 23, 2018 (received for review November 7, 2017) During development, ventricular chamber maturation is a crucial maturation of the ventricular wall is regulated by a complex sig- step in the formation of a functionally competent postnatal heart. naling network comprised of growth factors, transcription factors, Defects in this process can lead to left ventricular noncompaction and epigenetic regulators (3, 4), such as NRG1/ERBB2 (8), cardiomyopathy and heart failure. However, molecular mecha- NOTCH (9, 10), TGFβ (11, 12), Smad7 (13), and HDAC1/2 (14). nisms underlying ventricular chamber development remain However, the importance of novel ubiquitin-like protein modifiers incompletely understood. Neddylation is a posttranslational mod- in this process has not been demonstrated. ification that attaches ubiquitin-like protein NEDD8 to protein tar- Neural precursor cell expressed, developmentally down-regulated gets via NEDD8-specific E1-E2-E3 enzymes. Here, we report that 8 (NEDD8) is a ubiquitin-like protein that shares a high degree of neddylation is temporally regulated in the heart and plays a key homology with ubiquitin (15). -

Clonal Intratumor Heterogeneity of Promoter Hypermethylation In
Modern Pathology (2014) 27, 869–874 & 2014 USCAP, Inc. All rights reserved 0893-3952/14 $32.00 869 Clonal intratumor heterogeneity of promoter hypermethylation in breast cancer by MS-MLPA Cathy B Moelans, Jolien S de Groot, Xiaojuan Pan, Elsken van der Wall and Paul J van Diest Department of Pathology, University Medical Center Utrecht, Utrecht, The Netherlands Intratumor heterogeneity may lead to sampling bias and may present major challenges to personalized medicine and biomarker development. Despite many studies investigating genetic heterogeneity, epigenetic intratumor heterogeneity of promoter hypermethylation has only rarely been examined in breast cancer. To examine clonal intratumor heterogeneity of promotor hypermethylation, we performed methylation-specific multiplex ligation-dependent probe amplification (MS-MLPA) for 24 established tumor-suppressor genes on multiple spatially separated samples obtained from 21 primary breast carcinomas. Multiregion analysis was performed, representing at least two and a maximum of five tumor blocks per patient and four areas per tumor block. Methylation results were heterogeneous at one or more genetic loci between different tumor regions in 95% of breast carcinomas. The most heterogeneous loci in decreasing frequency were RASSF1A (62%), CDKN2B (43%), APC (38%), GSTP1 (33%), CDH13 (24%), DAPK1 (19%), and CDKN1B (5%). Heterogeneity lead to a methylation status change in at least one locus in 65% of the tumors. For most genes, the relative contribution of between-patients and between-block variability to the total variation in methylation results was similar. Regardless of the gene, contribution of within-block variability was of little importance. In conclusion, although most variation in methylation status is present between individual breast cancers, clonal epigenetic heterogeneity is seen within most primary breast carcinomas, indicating that methylation results from a single random sample may not be representative of the whole tumor. -

Aberrations of the P53 Pathway Components P53, MDM2 And
Leukemia (1999) 13, 453–459 1999 Stockton Press All rights reserved 0887-6924/99 $12.00 http://www.stockton-press.co.uk/leu Aberrations of the p53 pathway components p53, MDM2 and CDKN2A appear independent in diffuse large B cell lymphoma MB Møller1, Y Ino2, A-M Gerdes3, K Skjødt4, DN Louis2 and NT Pedersen1 Departments of 1Pathology, 3Clinical Genetics, and 4Medical Microbiology, Odense University, Denmark; and 2Molecular Neuro-Oncology Laboratory, Department of Pathology (Neuropathology) and Neurosurgical Service, Massachusetts General Hospital and Harvard Medical School, Boston, MA, USA The two gene products of the CDKN2A gene, p16 and p19ARF, ways together through the 9p21 locus.5,6 In this locus resides have recently been linked to each of two major tumour sup- two cyclin-dependent kinase inhibitor genes, CDKN2A and pressor pathways in human carcinogenesis, the RB1 pathway CDKN2B. The CDKN2A gene has the rare ability to encode and the p53 pathway. p16 inhibits the phosphorylation of the ARF retinoblastoma gene product by cyclin D-dependent kinases, two different proteins, p16 and p19 . This is due to the whereas p19ARF targets MDM2, a p53 inhibitory protein, for unusual organisation of the gene with two alternative first degradation. A deletion of CDKN2A would therefore disturb exons, exon 1␣ and exon 1, and common exons 2 and 3.7,8 both pathways. To explore the p53 pathway genes as a func- Because they are translated in different reading frames p16 tional unit in diffuse large B cell non-Hodgkin’s lymphomas and p19ARF are structurally unrelated. Despite this difference (DLCL), we wanted to see whether there exists mutually exclus- iveness of aberrations of CDKN2A, MDM2 and p53, since this both can arrest the cell cycle in G1, albeit through different has not been analysed previously. -

Early Adaptation and Acquired Resistance to CDK4/6 Inhibition In
Published OnlineFirst March 28, 2016; DOI: 10.1158/0008-5472.CAN-15-0728 Cancer Therapeutics, Targets, and Chemical Biology Research Early Adaptation and Acquired Resistance to CDK4/6 Inhibition in Estrogen Receptor–Positive Breast Cancer Maria Teresa Herrera-Abreu1, Marta Palafox2, Uzma Asghar1, Martín A. Rivas3, Rosalind J. Cutts1, Isaac Garcia-Murillas1, Alex Pearson1, Marta Guzman2, Olga Rodriguez2, Judit Grueso2, Meritxell Bellet4, Javier Cortes 4, Richard Elliott1, Sunil Pancholi1, Jose Baselga5, Mitch Dowsett1,6, Lesley-Ann Martin1, Nicholas C. Turner1,7, and Violeta Serra2 Abstract Small-molecule inhibitors of the CDK4/6 cell-cycle kinases apy, CDK4/6, and PI3K inhibition was more effective than have shown clinical efficacy in estrogen receptor (ER)-positive paired combinations, provoking rapid tumor regressions in a metastatic breast cancer, although their cytostatic effects are PDX model. Mechanistic investigations showed that acquired limited by primary and acquired resistance. Here we report that resistance to CDK4/6 inhibition resulted from bypass of cyclin ER-positive breast cancer cells can adapt quickly to CDK4/6 D1–CDK4/6 dependency through selection of CCNE1 ampli- inhibition and evade cytostasis, in part, via noncanonical cyclin fication or RB1 loss. Notably, although PI3K inhibitors could D1-CDK2–mediated S-phase entry. This adaptation was pre- prevent resistance to CDK4/6 inhibitors, they failed to resen- vented by cotreatment with hormone therapies or PI3K inhi- sitize cells once resistance had been acquired. However, we bitors, which reduced the levels of cyclin D1 (CCND1)and found that cells acquiring resistance to CDK4/6 inhibitors due other G1–S cyclins, abolished pRb phosphorylation, and inhib- to CCNE1 amplification could be resensitized by targeting ited activation of S-phase transcriptional programs. -

Analysis of the CDKN2A, CDKN2B and CDK4 Genes in 48 Australian Melanoma Kindreds
Oncogene (1997) 15, 2999 ± 3005 1997 Stockton Press All rights reserved 0950 ± 9232/97 $12.00 SHORT REPORT Analysis of the CDKN2A, CDKN2B and CDK4 genes in 48 Australian melanoma kindreds Jose F Flores1, Pamela M Pollock2, Graeme J Walker1,2, J Michael Glendening1, Amy H-T Lin1, Jane M Palmer2, Marilyn K Walters2, Nicholas K Hayward2 and Jane W Fountain1 1University of Southern California, Institute for Genetic Medical, Department of Biochemistry and Molecular Biology, 2011 Zonal Ave (HMR601), Los Angeles, CA, 90033, USA; 2Queensland Cancer Fund Research Unit, Queensland Institute of Medical Research, Herston, 4029, Australia Germline mutations within the cyclin-dependent kinase 1994, 1996; Walker et al., 1994; Luca et al., 1995; inhibitor 2A (CDKN2A) gene and one of its targets, the Platz et al., 1996; Flores et al., 1996) and is genetically cyclin dependent kinase 4 (CDK4) gene, have been linked to familial melanoma (Cannon-Albright et al., identi®ed in a proportion of melanoma kindreds. In the 1992; Nancarrow et al., 1993; Gruis et al., 1993, case of CDK4, only one speci®c mutation, resulting in the 1995a; Goldstein et al., 1994; MacGeoch et al., 1994; substitution of a cysteine for an arginine at codon 24 Holland et al., 1995). p16 is a pivotal cell cycle (R24C), has been found to be associated with melanoma. regulator that binds to the cyclin-dependent kinases 4 We have previously reported the identi®cation of germline (CDK4) and 6 (CDK6) and inhibits their ability to CDKN2A mutations in 7/18 Australian melanoma phosphorylate the retinoblastoma protein (pRb) kindreds and the absence of the R24C CDK4 mutation (Serrano et al., 1993).