DTNB Antibody Cat
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Individual Protomers of a G Protein-Coupled Receptor Dimer Integrate Distinct Functional Modules
OPEN Citation: Cell Discovery (2015) 1, 15011; doi:10.1038/celldisc.2015.11 © 2015 SIBS, CAS All rights reserved 2056-5968/15 ARTICLE www.nature.com/celldisc Individual protomers of a G protein-coupled receptor dimer integrate distinct functional modules Nathan D Camp1, Kyung-Soon Lee2, Jennifer L Wacker-Mhyre2, Timothy S Kountz2, Ji-Min Park2, Dorathy-Ann Harris2, Marianne Estrada2, Aaron Stewart2, Alejandro Wolf-Yadlin1, Chris Hague2 1Department of Genome Sciences, University of Washington School of Medicine, Seattle, WA, USA; 2Department of Pharmacology, University of Washington School of Medicine, Seattle, WA, USA Recent advances in proteomic technology reveal G-protein-coupled receptors (GPCRs) are organized as large, macromolecular protein complexes in cell membranes, adding a new layer of intricacy to GPCR signaling. We previously reported the α1D-adrenergic receptor (ADRA1D)—a key regulator of cardiovascular, urinary and CNS function—binds the syntrophin family of PDZ domain proteins (SNTA, SNTB1, and SNTB2) through a C-terminal PDZ ligand inter- action, ensuring receptor plasma membrane localization and G-protein coupling. To assess the uniqueness of this novel GPCR complex, 23 human GPCRs containing Type I PDZ ligands were subjected to TAP/MS proteomic analysis. Syntrophins did not interact with any other GPCRs. Unexpectedly, a second PDZ domain protein, scribble (SCRIB), was detected in ADRA1D complexes. Biochemical, proteomic, and dynamic mass redistribution analyses indicate syntrophins and SCRIB compete for the PDZ ligand, simultaneously exist within an ADRA1D multimer, and impart divergent pharmacological properties to the complex. Our results reveal an unprecedented modular dimeric architecture for the ADRA1D in the cell membrane, providing unexpected opportunities for fine-tuning receptor function through novel protein interactions in vivo, and for intervening in signal transduction with small molecules that can stabilize or disrupt unique GPCR:PDZ protein interfaces. -

The Amino Acid Sequence of Troponin C from Chicken Skeletal Muscle
View metadata, citation and similar papers at core.ac.uk brought to you by CORE provided by Elsevier - Publisher Connector Volume 70, number 1 FEBS LETTERS November 1976 THE AMINO ACID SEQUENCE OF TROPONIN C FROM CHICKEN SKELETAL MUSCLE J. M. WILKINSON Department of Biochemistry, University of Birmingham, P. 0. Box 363, Birmingham, BI5 2TT. England Received 21 September 1976 1. Introduction Hirabayshi and Perry [5] have shown that chicken troponin C isolated from a mixture of breast and leg Troponin C is the component of the troponin muscle is a single antigen and hence, by inference the complex which binds Ca2+ and thereby triggers the troponin C components present in both tissues are activation of the actomyosin ATPase and hence the very similar if not identical. The present paper reports onset of contraction. The amino acid sequence of the amino acid sequence of chicken troponin C and troponin C from rabbit fast skeletal muscle has been discusses its relationship with rabbit troponin C to determined by Collins et al. [l ] and that from which it is very similar. No evidence of heterogeneity bovine cardiac muscle by van Eerd and Takahashi in the sequence has been found. [2]. These proteins have been shown to be homolo- gous not only with the calcium binding parvalbumins 2. Experimental but also with both the DTNB and alkali light chains of myosin [a]. Based on the homology with the Troponin C was prepared from mixed breast and parvalbumins, four binding sites for Ca2+ have been leg muscle and purified by chromatography on DEAE- proposed for rabbit troponin C and a three dimensio- cellulose as described by Perry and Cole 171. -

Muscular Dystrophy in PTFR/Cavin-1 Null Mice
Muscular dystrophy in PTFR/cavin-1 null mice Shi-Ying Ding, … , Libin Liu, Paul F. Pilch JCI Insight. 2017;2(5):e91023. https://doi.org/10.1172/jci.insight.91023. Research Article Cell biology Muscle biology Mice and humans lacking the caveolae component polymerase I transcription release factor (PTRF, also known as cavin- 1) exhibit lipo- and muscular dystrophy. Here we describe the molecular features underlying the muscle phenotype for PTRF/cavin-1 null mice. These animals had a decreased ability to exercise, and exhibited muscle hypertrophy with increased muscle fiber size and muscle mass due, in part, to constitutive activation of the Akt pathway. Their muscles were fibrotic and exhibited impaired membrane integrity accompanied by an apparent compensatory activation of the dystrophin-glycoprotein complex along with elevated expression of proteins involved in muscle repair function. Ptrf deletion also caused decreased mitochondrial function, oxygen consumption, and altered myofiber composition. Thus, in addition to compromised adipocyte-related physiology, the absence of PTRF/cavin-1 in mice caused a unique form of muscular dystrophy with a phenotype similar or identical to that seen in humans lacking this protein. Further understanding of this muscular dystrophy model will provide information relevant to the human situation and guidance for potential therapies. Find the latest version: https://jci.me/91023/pdf RESEARCH ARTICLE Muscular dystrophy in PTFR/cavin-1 null mice Shi-Ying Ding,1 Libin Liu,1 and Paul F. Pilch1,2 1Department of Biochemistry, 2Department of Medicine, Boston University School of Medicine, Boston, Massachusetts, USA. Mice and humans lacking the caveolae component polymerase I transcription release factor (PTRF, also known as cavin-1) exhibit lipo- and muscular dystrophy. -

Analysis of the Dystrophin Interactome
Analysis of the dystrophin interactome Dissertation In fulfillment of the requirements for the degree “Doctor rerum naturalium (Dr. rer. nat.)” integrated in the International Graduate School for Myology MyoGrad in the Department for Biology, Chemistry and Pharmacy at the Freie Universität Berlin in Cotutelle Agreement with the Ecole Doctorale 515 “Complexité du Vivant” at the Université Pierre et Marie Curie Paris Submitted by Matthew Thorley born in Scunthorpe, United Kingdom Berlin, 2016 Supervisor: Simone Spuler Second examiner: Sigmar Stricker Date of defense: 7th December 2016 Dedicated to My mother, Joy Thorley My father, David Thorley My sister, Alexandra Thorley My fiancée, Vera Sakhno-Cortesi Acknowledgements First and foremost, I would like to thank my supervisors William Duddy and Stephanie Duguez who gave me this research opportunity. Through their combined knowledge of computational and practical expertise within the field and constant availability for any and all assistance I required, have made the research possible. Their overarching support, approachability and upbeat nature throughout, while granting me freedom have made this year project very enjoyable. The additional guidance and supported offered by Matthias Selbach and his team whenever required along with a constant welcoming invitation within their lab has been greatly appreciated. I thank MyoGrad for the collaboration established between UPMC and Freie University, creating the collaboration within this research project possible, and offering research experience in both the Institute of Myology in Paris and the Max Delbruck Centre in Berlin. Vital to this process have been Gisele Bonne, Heike Pascal, Lidia Dolle and Susanne Wissler who have aided in the often complex processes that I am still not sure I fully understand. -

Calcium Sensitivity of Actomyosin Atpase
Calcium Sensitivity of Actomyosin ATPase: Its Modification by Substitution of Myosin Sulfhydryl Groups Peter Dancker Max-Planck-Institut für medizinische Forschung, Abteilung Physiologie, Heidelberg (Z. Naturforsch. 30 c, 586 — 592 [1975]; received June 30, 1975) Actomyosin ATPase, Calcium Sensitivity, Sulfhydryl Groups SH group substitution by DTNB enabled natural actomyosin to split ATP (in the presence of Mg2+) also in the absence of Ca2*, when assayed at low ionic strength. At higher KC1 concentra tions the ATPase activity of SH group substituted actomyosin was still Ca-dependent. Addition of unsubstituted myosin to natural actomyosin whose SH groups had been substituted increased the ATPase activity. This increase was Ca-insensitive indicating that SH group substitution of myosin in actomyosin can make the interaction of additional myosin molecules Ca-independent. In natural actomyosin Ca-insensitivity of ATPase activity was attained at a lower degree of SH group sub stitution when substitution was performed in the presence of EDTA. The part of ATPase activity which still remained Ca-sensitive after DTNB treatment could be activated by lower concentrations of free Ca2+ than the Ca-sensitive ATPase of untreated actomyosin. In reconstituted actomyosin the Ca-sensitivity of ATPase activity could more easily be reduced when the myosin-actin ratio was high. For demonstrating remaining Ca-sensitivity in SH group substituted reconstituted acto myosin more tropomyosin-troponin was needed than for sensitizing unsubstituted actomyosin to Ca2+. — The similarities between the ATPase activity of SH group substituted actomyosin on the one hand and that of actomyosin at low concentrations of ATP on the other hand suggest that SH group substitution modifies actin-myosin interaction in a similar way as does nucleotide-free myosin (rigor myosin). -

Dystrobrevin in Nonmuscle Dystrophin-Associated Protein Complex-Like Complexes in Kidney and Liver
Role of β-Dystrobrevin in Nonmuscle Dystrophin-Associated Protein Complex-Like Complexes in Kidney and Liver Nellie Y. Loh, Daniela Nebenius-Oosthuizen, Derek J. Blake, Andrew J. H. Smith and Kay E. Davies Mol. Cell. Biol. 2001, 21(21):7442. DOI: 10.1128/MCB.21.21.7442-7448.2001. Downloaded from Updated information and services can be found at: http://mcb.asm.org/content/21/21/7442 These include: http://mcb.asm.org/ REFERENCES This article cites 18 articles, 11 of which can be accessed free at: http://mcb.asm.org/content/21/21/7442#ref-list-1 CONTENT ALERTS Receive: RSS Feeds, eTOCs, free email alerts (when new articles cite this article), more» on February 25, 2014 by Cardiff Univ Information about commercial reprint orders: http://journals.asm.org/site/misc/reprints.xhtml To subscribe to to another ASM Journal go to: http://journals.asm.org/site/subscriptions/ MOLECULAR AND CELLULAR BIOLOGY, Nov. 2001, p. 7442–7448 Vol. 21, No. 21 0270-7306/01/$04.00ϩ0 DOI: 10.1128/MCB.21.21.7442–7448.2001 Copyright © 2001, American Society for Microbiology. All Rights Reserved. Role of -Dystrobrevin in Nonmuscle Dystrophin-Associated Protein Complex-Like Complexes in Kidney and Liver NELLIE Y. LOH,1† DANIELA NEBENIUS-OOSTHUIZEN,2‡ DEREK J. BLAKE,1§ 2 1,3 ANDREW J. H. SMITH, AND KAY E. DAVIES * MRC Functional Genetics Unit,3 Department of Human Anatomy and Genetics,1 University of Oxford, Oxford OX1 3QX, and Centre for Genome Research, The University of Edinburgh, Edinburgh EH9 3JQ,2 United Kingdom Received 27 July 2001/Accepted 31 July 2001 Downloaded from -Dystrobrevin is a dystrophin-related and -associated protein that is highly expressed in brain, kidney, and liver. -

Perkinelmer Genomics to Request the Saliva Swab Collection Kit for Patients That Cannot Provide a Blood Sample As Whole Blood Is the Preferred Sample
Autism and Intellectual Disability TRIO Panel Test Code TR002 Test Summary This test analyzes 2429 genes that have been associated with Autism and Intellectual Disability and/or disorders associated with Autism and Intellectual Disability with the analysis being performed as a TRIO Turn-Around-Time (TAT)* 3 - 5 weeks Acceptable Sample Types Whole Blood (EDTA) (Preferred sample type) DNA, Isolated Dried Blood Spots Saliva Acceptable Billing Types Self (patient) Payment Institutional Billing Commercial Insurance Indications for Testing Comprehensive test for patients with intellectual disability or global developmental delays (Moeschler et al 2014 PMID: 25157020). Comprehensive test for individuals with multiple congenital anomalies (Miller et al. 2010 PMID 20466091). Patients with autism/autism spectrum disorders (ASDs). Suspected autosomal recessive condition due to close familial relations Previously negative karyotyping and/or chromosomal microarray results. Test Description This panel analyzes 2429 genes that have been associated with Autism and ID and/or disorders associated with Autism and ID. Both sequencing and deletion/duplication (CNV) analysis will be performed on the coding regions of all genes included (unless otherwise marked). All analysis is performed utilizing Next Generation Sequencing (NGS) technology. CNV analysis is designed to detect the majority of deletions and duplications of three exons or greater in size. Smaller CNV events may also be detected and reported, but additional follow-up testing is recommended if a smaller CNV is suspected. All variants are classified according to ACMG guidelines. Condition Description Autism Spectrum Disorder (ASD) refers to a group of developmental disabilities that are typically associated with challenges of varying severity in the areas of social interaction, communication, and repetitive/restricted behaviors. -

Modulation of Titin-Based Stiffness by Disulfide Bonding in the Cardiac
View metadata, citation and similar papers at core.ac.uk brought to you by CORE provided by Elsevier - Publisher Connector Biophysical Journal Volume 97 August 2009 825–834 825 Modulation of Titin-Based Stiffness by Disulfide Bonding in the Cardiac Titin N2-B Unique Sequence Anika Gru¨tzner,† Sergi Garcia-Manyes,‡ Sebastian Ko¨tter,† Carmen L. Badilla,‡ Julio M. Fernandez,‡ and Wolfgang A. Linke†§* †Physiology and Biophysics Unit, University of Mu¨nster, Mu¨nster, Germany; ‡Department of Biological Sciences, Columbia University, New York, New York; and §Institute of Physiology, Department of Cardiovascular Physiology, Ruhr University Bochum, Bochum, Germany ABSTRACT The giant protein titin is responsible for the elasticity of nonactivated muscle sarcomeres. Titin-based passive stiff- ness in myocardium is modulated by titin-isoform switching and protein-kinase (PK)A- or PKG-dependent titin phosphorylation. Additional modulatory effects on titin stiffness may arise from disulfide bonding under oxidant stress, as many immunoglobulin- like (Ig-)domains in titin’s spring region have a potential for S-S formation. Using single-molecule atomic force microscopy (AFM) force-extension measurements on recombinant Ig-domain polyprotein constructs, we show that titin Ig-modules contain no stabi- lizing disulfide bridge, contrary to previous belief. However, we demonstrate that the human N2-B-unique sequence (N2-Bus), a cardiac-specific, physiologically extensible titin segment comprising 572 amino-acid residues, contains up to three disulfide bridges under oxidizing conditions. AFM force spectroscopy on recombinant N2-Bus molecules demonstrated a much shorter contour length in the absence of a reducing agent than in its presence, consistent with intramolecular S-S bonding. -

Supplementary Table 1 Double Treatment Vs Single Treatment
Supplementary table 1 Double treatment vs single treatment Probe ID Symbol Gene name P value Fold change TC0500007292.hg.1 NIM1K NIM1 serine/threonine protein kinase 1.05E-04 5.02 HTA2-neg-47424007_st NA NA 3.44E-03 4.11 HTA2-pos-3475282_st NA NA 3.30E-03 3.24 TC0X00007013.hg.1 MPC1L mitochondrial pyruvate carrier 1-like 5.22E-03 3.21 TC0200010447.hg.1 CASP8 caspase 8, apoptosis-related cysteine peptidase 3.54E-03 2.46 TC0400008390.hg.1 LRIT3 leucine-rich repeat, immunoglobulin-like and transmembrane domains 3 1.86E-03 2.41 TC1700011905.hg.1 DNAH17 dynein, axonemal, heavy chain 17 1.81E-04 2.40 TC0600012064.hg.1 GCM1 glial cells missing homolog 1 (Drosophila) 2.81E-03 2.39 TC0100015789.hg.1 POGZ Transcript Identified by AceView, Entrez Gene ID(s) 23126 3.64E-04 2.38 TC1300010039.hg.1 NEK5 NIMA-related kinase 5 3.39E-03 2.36 TC0900008222.hg.1 STX17 syntaxin 17 1.08E-03 2.29 TC1700012355.hg.1 KRBA2 KRAB-A domain containing 2 5.98E-03 2.28 HTA2-neg-47424044_st NA NA 5.94E-03 2.24 HTA2-neg-47424360_st NA NA 2.12E-03 2.22 TC0800010802.hg.1 C8orf89 chromosome 8 open reading frame 89 6.51E-04 2.20 TC1500010745.hg.1 POLR2M polymerase (RNA) II (DNA directed) polypeptide M 5.19E-03 2.20 TC1500007409.hg.1 GCNT3 glucosaminyl (N-acetyl) transferase 3, mucin type 6.48E-03 2.17 TC2200007132.hg.1 RFPL3 ret finger protein-like 3 5.91E-05 2.17 HTA2-neg-47424024_st NA NA 2.45E-03 2.16 TC0200010474.hg.1 KIAA2012 KIAA2012 5.20E-03 2.16 TC1100007216.hg.1 PRRG4 proline rich Gla (G-carboxyglutamic acid) 4 (transmembrane) 7.43E-03 2.15 TC0400012977.hg.1 SH3D19 -

The Role of Titin and Nebulin in Myofibril Assembly in Cultured Embryonic Chick Muscle Cells
Iowa State University Capstones, Theses and Retrospective Theses and Dissertations Dissertations 1988 The oler of titin and nebulin in myofibril assembly in cultured embryonic chick muscle cells Michelle Ann Kurpakus Iowa State University Follow this and additional works at: https://lib.dr.iastate.edu/rtd Part of the Biochemistry Commons Recommended Citation Kurpakus, Michelle Ann, "The or le of titin and nebulin in myofibril assembly in cultured embryonic chick muscle cells " (1988). Retrospective Theses and Dissertations. 9689. https://lib.dr.iastate.edu/rtd/9689 This Dissertation is brought to you for free and open access by the Iowa State University Capstones, Theses and Dissertations at Iowa State University Digital Repository. It has been accepted for inclusion in Retrospective Theses and Dissertations by an authorized administrator of Iowa State University Digital Repository. For more information, please contact [email protected]. INFORMATION TO USERS The most advanced technology has been used to photo graph and reproduce this manuscript from the microfilm master. UMI films the original text directly firom the copy submitted. Thus, some dissertation copies are in typewriter face, while others may be from a computer printer. In the unlikely event that the author did not send UMI a complete manuscript and there are missing pages, these will be noted. Also, if unauthorized copyrighted material had to be removed, a note will indicate the deletion. Oversize materials (e.g., maps, drawings, charts) are re produced by sectioning the original, beginning at the upper left-hand comer and continuing from left to right in equal sections with small overlaps. Each oversize page is available as one exposure on a standard 35 mm slide or as a 17" x 23" black and white photographic print for an additional charge. -

The Roles of the Dystrophin-Associated Glycoprotein Complex at the Synapse
Mol Neurobiol (2010) 41:1–21 DOI 10.1007/s12035-009-8089-5 The Roles of the Dystrophin-Associated Glycoprotein Complex at the Synapse Gonneke S. K. Pilgram & Saranyapin Potikanond & Richard A. Baines & Lee G. Fradkin & Jasprina N. Noordermeer Received: 24 August 2009 /Accepted: 15 October 2009 /Published online: 9 November 2009 # The Author(s) 2009. This article is published with open access at Springerlink.com Abstract Duchenne muscular dystrophy is caused by muta- Keywords Dystrophin . DGC . NMJ . CNS . PNS . Retina . tions in the dystrophin gene and is characterized by Duchenne muscular dystrophy . Mammals . Drosophila . progressive muscle wasting. A number of Duchenne patients C. elegans also present with mental retardation. The dystrophin protein is part of the highly conserved dystrophin-associated glyco- Abbreviations protein complex (DGC) which accumulates at the neuromus- ACh Acetylcholine cular junction (NMJ) and at a variety of synapses in the AChR Acetylcholine receptor peripheral and central nervous systems. Many years of Akt Protein kinase B research into the roles of the DGC in muscle have revealed BMP Bone morphogenetic protein its structural function in stabilizing the sarcolemma. In Ca2+ Calcium ion addition, the DGC also acts as a scaffold for various signaling CaMKII Ca2+/calmodulin-dependent protein kinase II pathways. Here, we discuss recent advances in understanding cGMP Cyclic guanosine monophosphate DGC roles in the nervous system, gained from studies in both CNS Central nervous system vertebrate and invertebrate model systems. From these DGC Dystrophin-associated glycoprotein complex studies, it has become clear that the DGC is important for DMD Duchenne muscular dystrophy the maturation of neurotransmitter receptor complexes and for EPSP Excitatory postsynaptic potential the regulation of neurotransmitter release at the NMJ and ERG Electroretinogram central synapses. -
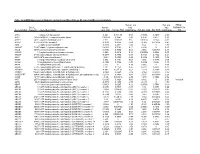
Mrna Expression in Human Leiomyoma and Eker Rats As Measured by Microarray Analysis
Table 3S: mRNA Expression in Human Leiomyoma and Eker Rats as Measured by Microarray Analysis Human_avg Rat_avg_ PENG_ Entrez. Human_ log2_ log2_ RAPAMYCIN Gene.Symbol Gene.ID Gene Description avg_tstat Human_FDR foldChange Rat_avg_tstat Rat_FDR foldChange _DN A1BG 1 alpha-1-B glycoprotein 4.982 9.52E-05 0.68 -0.8346 0.4639 -0.38 A1CF 29974 APOBEC1 complementation factor -0.08024 0.9541 -0.02 0.9141 0.421 0.10 A2BP1 54715 ataxin 2-binding protein 1 2.811 0.01093 0.65 0.07114 0.954 -0.01 A2LD1 87769 AIG2-like domain 1 -0.3033 0.8056 -0.09 -3.365 0.005704 -0.42 A2M 2 alpha-2-macroglobulin -0.8113 0.4691 -0.03 6.02 0 1.75 A4GALT 53947 alpha 1,4-galactosyltransferase 0.4383 0.7128 0.11 6.304 0 2.30 AACS 65985 acetoacetyl-CoA synthetase 0.3595 0.7664 0.03 3.534 0.00388 0.38 AADAC 13 arylacetamide deacetylase (esterase) 0.569 0.6216 0.16 0.005588 0.9968 0.00 AADAT 51166 aminoadipate aminotransferase -0.9577 0.3876 -0.11 0.8123 0.4752 0.24 AAK1 22848 AP2 associated kinase 1 -1.261 0.2505 -0.25 0.8232 0.4689 0.12 AAMP 14 angio-associated, migratory cell protein 0.873 0.4351 0.07 1.656 0.1476 0.06 AANAT 15 arylalkylamine N-acetyltransferase -0.3998 0.7394 -0.08 0.8486 0.456 0.18 AARS 16 alanyl-tRNA synthetase 5.517 0 0.34 8.616 0 0.69 AARS2 57505 alanyl-tRNA synthetase 2, mitochondrial (putative) 1.701 0.1158 0.35 0.5011 0.6622 0.07 AARSD1 80755 alanyl-tRNA synthetase domain containing 1 4.403 9.52E-05 0.52 1.279 0.2609 0.13 AASDH 132949 aminoadipate-semialdehyde dehydrogenase -0.8921 0.4247 -0.12 -2.564 0.02993 -0.32 AASDHPPT 60496 aminoadipate-semialdehyde