Enhancing Flavonoid Production by Promiscuous Activity Of
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Metabolic Engineering of Microbial Cell Factories for Biosynthesis of Flavonoids: a Review
molecules Review Metabolic Engineering of Microbial Cell Factories for Biosynthesis of Flavonoids: A Review Hanghang Lou 1,†, Lifei Hu 2,†, Hongyun Lu 1, Tianyu Wei 1 and Qihe Chen 1,* 1 Department of Food Science and Nutrition, Zhejiang University, Hangzhou 310058, China; [email protected] (H.L.); [email protected] (H.L.); [email protected] (T.W.) 2 Hubei Key Lab of Quality and Safety of Traditional Chinese Medicine & Health Food, Huangshi 435100, China; [email protected] * Correspondence: [email protected]; Tel.: +86-0571-8698-4316 † These authors are equally to this manuscript. Abstract: Flavonoids belong to a class of plant secondary metabolites that have a polyphenol structure. Flavonoids show extensive biological activity, such as antioxidative, anti-inflammatory, anti-mutagenic, anti-cancer, and antibacterial properties, so they are widely used in the food, phar- maceutical, and nutraceutical industries. However, traditional sources of flavonoids are no longer sufficient to meet current demands. In recent years, with the clarification of the biosynthetic pathway of flavonoids and the development of synthetic biology, it has become possible to use synthetic metabolic engineering methods with microorganisms as hosts to produce flavonoids. This article mainly reviews the biosynthetic pathways of flavonoids and the development of microbial expression systems for the production of flavonoids in order to provide a useful reference for further research on synthetic metabolic engineering of flavonoids. Meanwhile, the application of co-culture systems in the biosynthesis of flavonoids is emphasized in this review. Citation: Lou, H.; Hu, L.; Lu, H.; Wei, Keywords: flavonoids; metabolic engineering; co-culture system; biosynthesis; microbial cell factories T.; Chen, Q. -

Herbal Products and Their Active Constituents Used Alone and in Combination with Antifungal Drugs Against Drug-Resistant Candida Sp
antibiotics Review Herbal Products and Their Active Constituents Used Alone and in Combination with Antifungal Drugs against Drug-Resistant Candida sp. Anna Herman 1,* and Andrzej Przemysław Herman 2 1 Faculty of Health Sciences, Warsaw School of Engineering and Health, Bitwy Warszawskiej 1920 18 Street, 02-366 Warsaw, Poland 2 Department of Genetic Engineering, The Kielanowski Institute of Animal Physiology and Nutrition, Polish Academy of Sciences, Instytucka 3 Street, 05-110 Jabłonna, Poland; [email protected] * Correspondence: [email protected]; Tel.: +48-22-856-70-44; Fax: +48-22-646-34-18 Abstract: Clinical isolates of Candida yeast are the most common cause of opportunistic fungal infections resistant to certain antifungal drugs. Therefore, it is necessary to detect more effective anti- fungal agents that would be successful in overcoming such infections. Among them are some herbal products and their active constituents.The purpose of this review is to summarize the current state of knowledge onherbal products and their active constituents havingantifungal activity against drug- resistant Candida sp. used alone and in combination with antifungal drugs.The possible mechanisms of their action on drug-resistant Candida sp. including (1) inhibition of budding yeast transformation into hyphae; (2) inhibition of biofilm formation; (3) inhibition of cell wall or cytoplasmic membrane biosynthesis; (4) ROS production; and (5) over-expression of membrane transporters will be also described. Citation: Herman, A.; Herman, A.P. Herbal Products and Their Active Keywords: herbalproducts; herbal active constituents; drug-resistant Candida sp.; antifungal drug Constituents Used Alone and in Combination with Antifungal Drugs against Drug-Resistant Candida sp. -

Interpreting Sources and Endocrine Active Components of Trace Organic Contaminant Mixtures in Minnesota Lakes
INTERPRETING SOURCES AND ENDOCRINE ACTIVE COMPONENTS OF TRACE ORGANIC CONTAMINANT MIXTURES IN MINNESOTA LAKES by Meaghan E. Guyader © Copyright by Meaghan E. Guyader, 2018 All Rights Reserved A thesis submitted to the Faculty and the Board of Trustees of the Colorado School of Mines in partial fulfillment of the requirements for the degree of Doctor of Philosophy (Civil and Environmental Engineering). Golden, Colorado Date _____________________________ Signed: _____________________________ Meaghan E. Guyader Signed: _____________________________ Dr. Christopher P. Higgins Thesis Advisor Golden, Colorado Date _____________________________ Signed: _____________________________ Dr. Terri S. Hogue Professor and Department Head Department of Civil and Environmental Engineering ii ABSTRACT On-site wastewater treatment systems (OWTSs) are a suspected source of widespread trace organic contaminant (TOrC) occurrence in Minnesota lakes. TOrCs are a diverse set of synthetic and natural chemicals regularly used as cleaning agents, personal care products, medicinal substances, herbicides and pesticides, and foods or flavorings. Wastewater streams are known to concentrate TOrC discharges to the environment, particularly accumulating these chemicals at outfalls from centralized wastewater treatment plants. Fish inhabiting these effluent dominated environments are also known to display intersex qualities. Concurrent evidence of this phenomenon, known as endocrine disruption, in Minnesota lake fish drives hypotheses that OWTSs, the primary form of wastewater treatment in shoreline residences, may contribute to TOrC occurrence and the endocrine activity in these water bodies. The causative agents specific to fish in this region remain poorly understood. The objective of this dissertation was to investigate OWTSs as sources of TOrCs in Minnesota lakes, and TOrCs as potential causative agents for endocrine disruption in resident fish. -

Introduction Common Gynecologic Ailments Could Be Menstrual Period
สมนุ ไพรสำ หรบั โรคสตรที ใี่ ชโ้ ดยหมอพนื้ บำ้ นในจงั หวดั นครนำยก The Use of Medicinal Plants for Gynecologic Ailments by Thai Traditional Folk Healers in Nakhonnayok Province นิพนธ์ต้นฉบ ับ Original Article วรพรรณ สทิ ธถิ าวร1*, ลลิตา วีระเสถียร1 และ ชไมพร อ ้นสว่าง2 Worapan Sitthithaworn1*, Lalita Weerasathien1 and Chamaiporn Onsawang2 1 สาขาเภสชั เวท คณะเภสชั ศาสตร ์ มหาวิทยาลัยศรีนครินทรวิโรฒ อ.องครักษ์ จ.นครนายก 26120 1 Department of Pharmacognosy, Faculty of Pharmacy, Srinakharinwirot University, Ongkarak, 2 งานแพทย์แผนไทย กลุ่มงานคุม้ ครองผูบ้ รโิ ภคและเภสชั สาธารณสขุ ส านักงานสาธารณสขุ จังหวัดนครนายก Nakonnayok 26120, Thailand 2 อ.เมือง จ.นครนายก 26000 Thai Traditional Medicine Unit, Division of Consumer Protection and Public Health Pharmacy, Nakhonnayok Public Health Administration Office, Muang, Nakonnayok 26000, Thailand * Corresponding author: [email protected] * Corresponding author: [email protected] วำรสำรไทยเภสชั ศำสตรแ์ ละวทิ ยำกำรสุขภำพ 2562;14(3):111-121. Thai Pharmaceutical and Health Science Journal 2019;14(3):111-121. บทค ัดย่อ Abstract วัตถุประสงค์: เพื่อระบุสมุนไพรที่หมอพื้นบ้านในจังหวัดนครนายกใช้รักษาโรค Objective: To determine medicinal plants used by folk healers in สตรีในกลุ่มอาการไข้ทับระดู ปวดประจาเดือน ประจาเดือนมาไม่ปกติ และตกขาว Nakhonnayok province for gynecological ailments including pelvic และศึกษาความสัมพันธ์ของสรรพคุณสมุนไพรกับผลการศึกษาฤทธทิ์ างเภสชั inflammatory disease (menstrual fever), dysmenorrhea, oligomenorrhea and วิทยาที่มีรายงานไว้ วิธีการศึกษา: การวิจัยเชิงคุณภาพนี้เก็บข้อมูลโดยการ -

Therapeutic Effects of Bossenbergia Rotunda
International Journal of Science and Research (IJSR) ISSN (Online): 2319-7064 Index Copernicus Value (2015): 78.96 | Impact Factor (2015): 6.391 Therapeutic Effects of Bossenbergia rotunda S. Aishwarya Bachelor of Dental Surgery, Saveetha Dental College and Hospitals Abstract: Boesenbergia rotunda (L.) (Fingerroot), formerly known as Boesenbergia or Kaempferiapandurata (Roxb). Schltr. (Zingiberaceae), is distributed in south-east Asian countries, such as Indonesia, Malaysia and Thailand. The rhizomes of this plant have been used for the treatment of peptic ulcer, as well as colic, oral diseases, urinary disorders, dysentery and inflammation. As people have started to focus more on natural plants species for their curative properties. B. rotunda is a native ingredient in many Asian countries and is used as a condiment in food. It is also used as traditional medicine to treat several illnesses, consumed as traditional tonic especially after childbirth, beauty aid for teenage girls, and as a leukorrhea preventive remedy for women. Its fresh rhizomes are also used to treat inflammatory diseases, in addition to being used as an antifungal, antiparasitic, and aphrodisiac among Thai folks. Moreover, AIDS patients self-medicate themselves with B. rotunda to cure the infection. With the advancement in technology, the ethnomedicinal usages of herbal plants can be explained through in vitro and in vivo studies to prove the activities of the plant extracts. The current state of research on B. rotunda clearly shows that the isolated bioactive compounds have high potential in treating many diseases. Keywords: Zingerberaceae, anti fungal, anti parasitic, Chalcones, flavonoids. 1. Introduction panduratin derivative are prenylated flavonoids from B. pandurata that showed broad range of biological activities, Boesenbergia rotunda is a ginger species that grows in such as strong antibacterial acitivity9-11, anti- inflammatory Southeast Asia, India, Sri Lanka, and Southern China. -

Combinatorial Biosynthesis of Non-Bacterial and Unnatural Flavonoids, Stilbenoids and Curcuminoids by Microorganisms Sueharu Horinouchi
J. Antibiot. 61(12): 709–728, 2008 THE JOURNAL OF REVIEW ARTICLE ANTIBIOTICS Combinatorial Biosynthesis of Non-bacterial and Unnatural Flavonoids, Stilbenoids and Curcuminoids by Microorganisms Sueharu Horinouchi Received: August 1, 2008 / Accepted: October 14, 2008 © Japan Antibiotics Research Association Abstract One of the approaches of combinatorial biosynthesis is combining genes from different organisms and designing a new set of gene clusters to produce bioactive compounds, leading to diversification of both chemical and natural product libraries. This makes efficient use of the potential of the host organisms, especially when microorganisms are used. An Escherichia coli system, in which artificial biosynthetic pathways for production of plant-specific medicinal polyketides, such as flavonoids, stilbenoids, isoflavonoids, and curcuminoids, are assembled, has been designed and expressed. Starting with amino acids tyrosine and phenylalanine as substrates, this system yields naringenin, resveratrol, genistein, and curcumin, for example, all of which are beneficial to human health because of their wide variety of biological activities. Supplementation of unnatural carboxylic acids to the recombinant E. coli cells carrying the artificial pathways by precursor-directed biosynthesis results in production of unnatural compounds. Addition of decorating or modification enzymes to the artificial pathway leads to production of natural and unnatural flavonols, flavones, and methylated resveratrols. This microbial system is promising for construction of larger libraries by employing other polyketide synthases and decorating enzymes of various origins. In addition, the concept of building and expressing artificial biosynthetic pathways for production of non-bacterial and unnatural compounds in microorganisms should be successfully applied to production of not only plant-specific polyketides but also many other useful compound classes. -

Micropropagation-An in Vitro Technique for the Conservation of Alpinia Galanga
Available online a t www.pelagiaresearchlibrary.com Pelagia Research Library Advances in Applied Science Research, 2014, 5(3):259-263 ISSN: 0976-8610 CODEN (USA): AASRFC Micropropagation-an in vitro technique for the conservation of Alpinia galanga Nongmaithem M. Singh 1, Lukram A. Chanu 1, Yendrembam P. Devi 1, Wahengbam R.C. Singh 2 and Heigrujam B. Singh 2 1DBT-Institutional Biotech Hub, Pettigrew College, Ukhrul, Manipur 2DBT- Institutional Biotech Hub, Deptt. of Biotechnology, S.K. Women’s College, Nambol, Manipur _____________________________________________________________________________________________ ABSTRACT This study was conducted to develop an efficient protocol for mass propagation of Alpinia galanga L. Explants from rhizome buds were cultured on Murashige and Skoog (MS) medium supplemented with 6-Benzylaminopurine (BAP) alone (0 to 5 mg/l) or a combination of BAP (0 to 5 mg/l) and indole 3-acetic acid (IAA) (0 to 2 mg/l). MS medium supplemented with a combination of 5.0 mg/l BAP and 2.0 mg/l IAA or 3.0 mg/l BAP and 0.5 mg/l IAA produced the highest mean number of shoots per explant as compared to other concentrations. The best shoot length was obtained on the medium containing 1.0 mg/l of BAP and 2.0 mg/l IAA. Thus, combined effects of BAP and IAA improved significantly the shoot growth and proliferation. MS medium supplemented with a combination of 5.0 mg/l BAP and 2 mg/l IAA gave the highest number of roots. However, longest roots per explant were obtained with 1.0 mg/l BAP alone. -

Development and Optimization of a High Sensitivity LC-MS/MS Method for the Determination of Hesperidin and Naringenin in Rat Plasma: Pharmacokinetic Approach
molecules Article Development and Optimization of a High Sensitivity LC-MS/MS Method for the Determination of Hesperidin and Naringenin in Rat Plasma: Pharmacokinetic Approach Jesús Alfredo Araujo-León 1,* , Rolffy Ortiz-Andrade 2, Rivelino Armando Vera-Sánchez 1, Julio Enrique Oney-Montalvo 1, Tania Isolina Coral-Martínez 1 and Zulema Cantillo-Ciau 3 1 Chromatography Laboratory, Chemistry Faculty, Autonomous University of Yucatan, Mérida 97069, Mexico; [email protected] (R.A.V.-S.); [email protected] (J.E.O.-M.); [email protected] (T.I.C.-M.) 2 Pharmacology Laboratory, Chemistry Faculty, Autonomous University of Yucatan, Mérida 97069, Mexico; rolff[email protected] 3 Medicinal Chemistry Laboratory, Chemistry Faculty, Autonomous University of Yucatan, Mérida 97069, Mexico; [email protected] * Correspondence: [email protected]; Tel.: +52-999-922-5708 Academic Editor: Derek J. McPhee Received: 6 August 2020; Accepted: 12 September 2020; Published: 16 September 2020 Abstract: The purpose of this study was to develop, optimize, and fully validate a high-sensitivity methodology using UHPLC-MS/MS to simultaneously quantify hesperidin and naringenin in microsamples (100 µL) of murine plasma after intragastric administration of single pure flavonoids and a mixture. The optimization process allowed for high sensitivity with detection limits of approximately picogram order using an electrospray ionization (ESI) source in negative mode and an experiment based on multiple reaction monitoring (MRM). The validation parameters showed excellent linearity and detection limits, with a precision of less than 8% and a recovery of over 90%. This methodology was applied to compare the pharmacokinetic parameters for the administration of hesperidin and naringenin in individual form or in the form of a mixture. -

C-23 Phytochemical of Kaempferia Plant And
Proceeding of International Conference On Research, Implementation And Education Of Mathematics And Sciences 2014, Yogyakarta State University, 18-20 May 2014 C-23 PHYTOCHEMICAL OF KAEMPFERIA PLANT AND BIOPROSPECTING FOR CANCER TREATMENT Sri Atun Chemistry education Faculty of Mathematical and Natural Science, Yogyakarta State University, Jl. Colombo No. 1 Yogyakarta, Indonesia, 55281 e-mail : [email protected] ABSTRACT Kaempferia genus is perennial member of the Zingiberaceae family and is cultivated in Indonesia and other parts of Southeast Asia. Number of studies has been conducted, providing information related to Kaempferia as antioxidant; antimutagenic; and chemopreventive agent. This paper reports some isolated compounds from this plant, biological activity, and bioprospecting for cancer treatment. Keyword: Cancer treatment; Kaempferia; Zingiberaceae INTRODUCTION Kaempferia is a genus, belong to family of Zingiberaceae. This plant grows in Southeast Asia, India, Sri Lanka, Indonesia, and Southem China. Kaempferia genus sinonim with Boesenbergia genus by Baker. This plant has 8 different botanical names which are Boesenbergia cochinchinensis (Gagnep.) Loes., Boesenbergia pandurata (Roxb.) Schltr., Curcuma rotunda L., Gastrochilus panduratus (Roxb.) Ridl., Gastrochilus rotundus (L.) Alston, Kaempferia cochinchinensis Gagnep., Kaempferia ovate Roscoe, Kaempferia galanga, Kaempferia rotunda, and Kaempferia pandurata Roxb nonetheless it is currently known as Boesenbergia rotunda (L.)Mansf (Tan Eng-Chong, et. al, 2012). The plants grown naturally in damp, shaded parts of the lowland or on hill slopes, as scattered plants or thickets. Economically important species among the plant families, the Zingiberaceae, which are perennial rhizomatous herbs, contain volatile oil and other important compounds of enormous medicinal values (Singh C.B., 2013). Phytochemical and biologycal activities of some species of Kaempferia Phytochemical and biologycal some species of plants of the genus Kaempferia reported by many researchers, among others: 1. -

Vasorelaxant Effect of Boesenbergia Rotunda and Its Active Ingredients
plants Article Vasorelaxant Effect of Boesenbergia rotunda and Its Active Ingredients on an Isolated Coronary Artery 1, 1, 1 1 2 Deepak Adhikari y, Dal-Seong Gong y, Se Hee Oh , Eun Hee Sung , Seung On Lee , Dong-Wook Kim 2, Min-Ho Oak 1,* and Hyun Jung Kim 1,* 1 College of Pharmacy and Natural Medicine Research Institute, Mokpo National University, Muan-gun 58554, Korea; [email protected] (D.A.); [email protected] (D.-S.G.); [email protected] (S.H.O.); [email protected] (E.H.S.) 2 Department of Oriental Medicine Resources, Mokpo National University, Muan-gun 58554, Korea; [email protected] (S.O.L.); [email protected] (D.-W.K.) * Correspondence: [email protected] (M.-H.O.); [email protected] (H.J.K.); Tel.: +82-61-450-2681 (M.-H.O.); +82-61-450-2686 (H.J.K.) The authors contributed to this work equally. y Received: 4 November 2020; Accepted: 27 November 2020; Published: 1 December 2020 Abstract: Cardiovascular diseases are a major cause of death in developed countries. The regulation of vascular tone is a major approach to prevent and ameliorate vascular diseases. As part of our ongoing screening for cardioprotective natural compounds, we investigated the vasorelaxant effect of rhizomes from Boesenbergia rotunda (L.) Mansf. [Boesenbergia pandurata (Roxb.) Schltr.] used as a spice and herbal medicine in Asian countries. The methanol extract of B. rotunda rhizomes (BRE) exhibited significant vasorelaxation effects ex vivo at EC values of 13.4 6.1 µg/mL and 40.9 7.9 µg/mL, 50 ± ± respectively, with and without endothelium in the porcine coronary artery ring. -
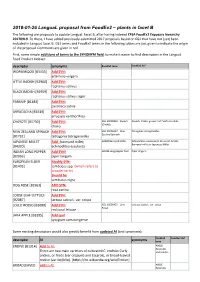
2018-01-26 Langual Proposal from Foodex2 – Plants in Facet B
2018-01-26 LanguaL proposal from FoodEx2 – plants in facet B The following are proposals to update LanguaL Facet B, after having indexed EFSA FoodEx2 Exposure hierarchy 20170919. To these, I have added previously-submitted 2017 proposals based on GS1 that have not (yet) been included in LanguaL facet B. GS1 terms and FoodEx2 terms in the following tables are just given to indicate the origin of the proposal. Comments are given in red. First, some simple additions of terms to the SYNONYM field, to make it easier to find descriptors in the LanguaL Food Product Indexer: descriptor synonyms FoodEx2 term FoodEx2 def WORMWOOD [B3433] Add SYN: artemisia vulgaris LITTLE RADISH [B2960] Add SYN: raphanus sativus BLACK RADISH [B2959] Add SYN: raphanus sativus niger PARSNIP [B1483] Add SYN: pastinaca sativa ARRACACHA [B3439] Add SYN: arracacia xanthorrhiza CHAYOTE [B1730] Add SYN: GS1 10006356 - Squash Squash, Choko, grown from Sechium edule (Choko) choko NEW ZEALAND SPINACH Add SYN: GS1 10006427 - New- Tetragonia tetragonoides Zealand Spinach [B1732] tetragonia tetragonoides JAPANESE MILLET Add : barnyard millet; A000Z Barnyard millet Echinochloa esculenta (A. Braun) H. Scholz, Barnyard millet or Japanese Millet. [B4320] echinochloa esculenta INDIAN LONG PEPPER Add SYN! A019B Long pepper fruit Piper longum [B2956] piper longum EUROPEAN ELDER Modify SYN: [B1403] sambucus spp. (which refers to broader term) Should be sambucus nigra DOG ROSE [B2961] ADD SYN: rosa canina LOOSE LEAF LETTUCE Add SYN: [B2087] lactusa sativa L. var. crispa LOLLO ROSSO [B2088] Add SYN: GS1 10006425 - Lollo Lactuca sativa L. var. crispa Rosso red coral lettuce JAVA APPLE [B3395] Add syn! syzygium samarangense Some existing descriptors would also greatly benefit from updated AI (and synonyms): FoodEx2 FoodEx2 def descriptor AI synonyms term ENDIVE [B1314] Add to AI: A00LD Escaroles There are two main varieties of cultivated C. -

Cushnie TPT, Lamb AJ. Antimicrobial Activity of Flavonoids. International Journal of Antimicrobial Agents, 2005. 26(5):343-356
Cushnie TPT, Lamb AJ. Antimicrobial activity of flavonoids. International Journal of Antimicrobial Agents, 2005. 26(5):343-356. PMID: 16323269 DOI: 10.1016/j.ijantimicag.2005.09.002 The journal article above is freely available from the publishers at: http://www.idpublications.com/journals/PDFs/IJAA/ANTAGE_MostCited_1.pdf and also... http://www.ijaaonline.com/article/S0924-8579(05)00255-4/fulltext Errata for the article (typesetting errors by Elsevier Ireland) are freely available from the publishers at: http://www.ijaaonline.com/article/S0924-8579(05)00352-3/fulltext and also... http://www.sciencedirect.com/science/article/pii/S0924857905003523 International Journal of Antimicrobial Agents 26 (2005) 343–356 Review Antimicrobial activity of flavonoids T.P. Tim Cushnie, Andrew J. Lamb ∗ School of Pharmacy, The Robert Gordon University, Schoolhill, Aberdeen AB10 1FR, UK Abstract Flavonoids are ubiquitous in photosynthesising cells and are commonly found in fruit, vegetables, nuts, seeds, stems, flowers, tea, wine, propolis and honey. For centuries, preparations containing these compounds as the principal physiologically active constituents have been used to treat human diseases. Increasingly, this class of natural products is becoming the subject of anti-infective research, and many groups have isolated and identified the structures of flavonoids possessing antifungal, antiviral and antibacterial activity. Moreover, several groups have demonstrated synergy between active flavonoids as well as between flavonoids and existing chemotherapeutics. Reports of activity in the field of antibacterial flavonoid research are widely conflicting, probably owing to inter- and intra-assay variation in susceptibility testing. However, several high-quality investigations have examined the relationship between flavonoid structure and antibacterial activity and these are in close agreement.