C. Elegans Meeting
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Caenorhabditis Microbiota: Worm Guts Get Populated Laura C
Clark and Hodgkin BMC Biology (2016) 14:37 DOI 10.1186/s12915-016-0260-7 COMMENTARY Open Access Caenorhabditis microbiota: worm guts get populated Laura C. Clark and Jonathan Hodgkin* Please see related Research article: The native microbiome of the nematode Caenorhabditis elegans: Gateway to a new host-microbiome model, http://dx.doi.org/10.1186/s12915-016-0258-1 effects on the life history of the worm are often profound Abstract [2]. It has been increasingly recognized that the worm Until recently, almost nothing has been known about microbiota is an important consideration in achieving a the natural microbiota of the model nematode naturalistic experimental model in which to study, for Caenorhabditis elegans. Reporting their research in instance, host–pathogen interactions or worm behavior. BMC Biology, Dirksen and colleagues describe the first Dirksen et al [3] present the first step towards under- sequencing effort to characterize the gut microbiota standing understanding the complex interactions of the of environmentally isolated C. elegans and the related natural worm microbiota by reporting a 16S rDNA-based taxa Caenorhabditis briggsae and Caenorhabditis “head count” of the bacterial population present in wild remanei In contrast to the monoxenic, microbiota-free nematode isolates (Fig. 1). Interestingly, it appears that cultures that are studied in hundreds of laboratories, it nematodes isolated from diverse natural environment- appears that natural populations of Caenorhabditis s—and even those that have been maintained for a short harbor distinct microbiotas. time on E. coli following isolation—share a “core” host- defined microbiota. This finding is in agreement with work by Berg et al. -

Novel Driver Strength Index Highlights Important Cancer Genes in TCGA Pancanatlas Patients
medRxiv preprint doi: https://doi.org/10.1101/2021.08.01.21261447; this version posted August 5, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted medRxiv a license to display the preprint in perpetuity. It is made available under a CC-BY-NC-ND 4.0 International license . Novel Driver Strength Index highlights important cancer genes in TCGA PanCanAtlas patients Aleksey V. Belikov*, Danila V. Otnyukov, Alexey D. Vyatkin and Sergey V. Leonov Laboratory of Innovative Medicine, School of Biological and Medical Physics, Moscow Institute of Physics and Technology, 141701 Dolgoprudny, Moscow Region, Russia *Corresponding author: [email protected] NOTE: This preprint reports new research that has not been certified by peer review and should not be used to guide clinical practice. 1 medRxiv preprint doi: https://doi.org/10.1101/2021.08.01.21261447; this version posted August 5, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted medRxiv a license to display the preprint in perpetuity. It is made available under a CC-BY-NC-ND 4.0 International license . Abstract Elucidating crucial driver genes is paramount for understanding the cancer origins and mechanisms of progression, as well as selecting targets for molecular therapy. Cancer genes are usually ranked by the frequency of mutation, which, however, does not necessarily reflect their driver strength. Here we hypothesize that driver strength is higher for genes that are preferentially mutated in patients with few driver mutations overall, because these few mutations should be strong enough to initiate cancer. -

Antigen-Specific Memory CD4 T Cells Coordinated Changes in DNA
Downloaded from http://www.jimmunol.org/ by guest on September 24, 2021 is online at: average * The Journal of Immunology The Journal of Immunology published online 18 March 2013 from submission to initial decision 4 weeks from acceptance to publication http://www.jimmunol.org/content/early/2013/03/17/jimmun ol.1202267 Coordinated Changes in DNA Methylation in Antigen-Specific Memory CD4 T Cells Shin-ichi Hashimoto, Katsumi Ogoshi, Atsushi Sasaki, Jun Abe, Wei Qu, Yoichiro Nakatani, Budrul Ahsan, Kenshiro Oshima, Francis H. W. Shand, Akio Ametani, Yutaka Suzuki, Shuichi Kaneko, Takashi Wada, Masahira Hattori, Sumio Sugano, Shinichi Morishita and Kouji Matsushima J Immunol Submit online. Every submission reviewed by practicing scientists ? is published twice each month by Author Choice option Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts http://jimmunol.org/subscription Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html Freely available online through http://www.jimmunol.org/content/suppl/2013/03/18/jimmunol.120226 7.DC1 Information about subscribing to The JI No Triage! Fast Publication! Rapid Reviews! 30 days* Why • • • Material Permissions Email Alerts Subscription Author Choice Supplementary The Journal of Immunology The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2013 by The American Association of Immunologists, Inc. All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. This information is current as of September 24, 2021. Published March 18, 2013, doi:10.4049/jimmunol.1202267 The Journal of Immunology Coordinated Changes in DNA Methylation in Antigen-Specific Memory CD4 T Cells Shin-ichi Hashimoto,*,†,‡ Katsumi Ogoshi,* Atsushi Sasaki,† Jun Abe,* Wei Qu,† Yoichiro Nakatani,† Budrul Ahsan,x Kenshiro Oshima,† Francis H. -

Figure S1. HAEC ROS Production and ML090 NOX5-Inhibition
Figure S1. HAEC ROS production and ML090 NOX5-inhibition. (a) Extracellular H2O2 production in HAEC treated with ML090 at different concentrations and 24 h after being infected with GFP and NOX5-β adenoviruses (MOI 100). **p< 0.01, and ****p< 0.0001 vs control NOX5-β-infected cells (ML090, 0 nM). Results expressed as mean ± SEM. Fold increase vs GFP-infected cells with 0 nM of ML090. n= 6. (b) NOX5-β overexpression and DHE oxidation in HAEC. Representative images from three experiments are shown. Intracellular superoxide anion production of HAEC 24 h after infection with GFP and NOX5-β adenoviruses at different MOIs treated or not with ML090 (10 nM). MOI: Multiplicity of infection. Figure S2. Ontology analysis of HAEC infected with NOX5-β. Ontology analysis shows that the response to unfolded protein is the most relevant. Figure S3. UPR mRNA expression in heart of infarcted transgenic mice. n= 12-13. Results expressed as mean ± SEM. Table S1: Altered gene expression due to NOX5-β expression at 12 h (bold, highlighted in yellow). N12hvsG12h N18hvsG18h N24hvsG24h GeneName GeneDescription TranscriptID logFC p-value logFC p-value logFC p-value family with sequence similarity NM_052966 1.45 1.20E-17 2.44 3.27E-19 2.96 6.24E-21 FAM129A 129. member A DnaJ (Hsp40) homolog. NM_001130182 2.19 9.83E-20 2.94 2.90E-19 3.01 1.68E-19 DNAJA4 subfamily A. member 4 phorbol-12-myristate-13-acetate- NM_021127 0.93 1.84E-12 2.41 1.32E-17 2.69 1.43E-18 PMAIP1 induced protein 1 E2F7 E2F transcription factor 7 NM_203394 0.71 8.35E-11 2.20 2.21E-17 2.48 1.84E-18 DnaJ (Hsp40) homolog. -

Genetic Analyses of Signal Transduction Pathways
GENETIC ANALYSES OF SIGNAL TRANSDUCTION PATHWAYS INVOLVED IN NEUROMUSCULAR EXCITABILITY AND NEURODEGENERATION IN CAENORHABTIDIS ELEGANS by BWARENABA KAUTU GUY A. CALDWELL, COMMITTEE CHAIR KIM A. CALDWELL JANIS M. O’DONNELL STEVAN MARCUS KATRINA RAMONELL ANDREW WEST A DISSERTATION Submitted in partial fulfillment of the requirements for the degree of Doctor of Philosophy in the Department of Biological Sciences in the Graduate School of The University of Alabama TUSCALOOSA, ALABAMA 2012 Copyright Bwarenaba Kautu 2012 ALL RIGHTS RESERVED ABSTRACT Signal transduction pathways regulate many cellular and molecular aspects of the brain including neurotransmission and cell survival. Defects in neuronal signaling can lead to a variety of cognitive and affective disorders such as epilepsy and Parkinson’s disease. Here, I use a genetically tractable organism, Caenorhabditis elegans (C. elegans), as a model system to study the impact of two canonical signaling pathways on neuronal activity and survival. Using molecular and genetic tools, pharmacological assays, and microscopy techniques, I showed that the canonical Rac GTPase pathway regulates neuronal synchrony in the GABAergic neurons of C. elegans. In our experiments we observed that Rac GTPase mutants exhibited behavioral responses to a GABAA receptor antagonist, pentylenetetrazole. These mutants also exhibited hypersensitivities to an acetylcholinesterase inhibitor, aldicarb, suggesting deficiencies in GABA transmission. Knockdown of selected cytoskeletal genes in Rac hypomorph mutants revealed synergistic interactions, particularly between the dynein motor complex and some members of the canonical Rac-signaling pathway. Examination of the nerve cords of C. elegans revealed that these genetic factors function to regulate vesicle transport in the GABAergic neurons of C. elegans. In my second project, I characterized the role of the heterotrimeric G protein Gαq in the context of neuronal survival, using a C. -

Genetics and Pathogenesis of Polycystic Kidney Disease
DISEASE OF THE MONTH J Am Soc Nephrol 13: 2384–2398, 2002 Eberhard Ritz, Feature Editor Genetics and Pathogenesis of Polycystic Kidney Disease PETER IGARASHI* and STEFAN SOMLO† *Department of Internal Medicine, The University of Texas Southwestern Medical Center, Dallas, Texas; and †Departments of Internal Medicine and Genetics, Yale University School of Medicine, New Haven, Connecticut. Polycystic kidney disease (PKD), a common genetic cause of Autosomal Dominant Polycystic Kidney Disease chronic renal failure in children and adults, is characterized by ADPKD is one of the most common genetic diseases in the accumulation of fluid-filled cysts in the kidney and other humans affecting all ethnic groups worldwide with an inci- organs. The renal cysts originate from the epithelia of the dence of 1 in 500 to 1 in 1,000 (7). The clinical manifestations nephrons and renal collecting system and are lined by a single include abdominal mass, chronic flank or back pain, gross layer of cells that have higher rates of cellular proliferation and hematuria, urinary tract infection, and urolithiasis. Affected are less differentiated than normal tubular cells (1). Abnormal- individuals typically present in the third and fourth decade, and ities in gene expression, cell polarity, fluid secretion, apoptosis, ESRD usually occurs within 5 to 10 yr after the development and extracellular matrix have also been described in PKD, but of renal insufficiency. However, presentation in infancy or the mechanism of cyst formation remains incompletely under- childhood has also been reported (8,9). In addition to causing stood (2–6). In recent months, there have been several ad- progressive renal failure, renal cysts can be complicated by vances in our understanding of the genetics and pathogenesis hemorrhage, rupture, infection, nephrolithiasis, and intractable of PKD. -

Caenorhabditis Elegans and Caenorhabditis Briggsae
Mol Gen Genomics (2005) 273: 299–310 DOI 10.1007/s00438-004-1105-6 ORIGINAL PAPER Richard Jovelin Æ Patrick C. Phillips Functional constraint and divergence in the G protein family in Caenorhabditis elegans and Caenorhabditis briggsae Received: 2 July 2004 / Accepted: 9 December 2004 / Published online: 27 April 2005 Ó Springer-Verlag 2005 Abstract Part of the challenge of the post-genomic Keywords Caenorhabditis elegans Æ Caenorhabditis world is to identify functional elements within the wide briggsae Æ G protein Æ Divergence Æ Gene regulation array of information generated by genome sequencing. Although cross-species comparisons and investigation of rates of sequence divergence are an efficient approach, the relationship between sequence divergence and func- Introduction tional conservation is not clear. Here, we use a com- parative approach to examine questions of evolutionary Recent whole genome sequencing projects have revealed rates and conserved function within the guanine nucle- that a substantial portion of genome evolution consists otide-binding protein (G protein) gene family in nema- of divergence and diversification of gene families (e.g., todes of the genus Caenorhabditis. In particular, we Chervitz et al. 1998; Lander et al. 2001; Venter et al. show that, in cases where the Caenorhabditis elegans 2001; Zdobnov et al. 2002). One of the primary chal- ortholog shows a loss-of-function phenotype, G protein lenges in this emerging field is to use information on genes of C. elegans and Caenorhabditis briggsae diverge sequence similarity and divergence among genomes to on average three times more slowly than G protein genes infer gene function. Very low rates of change might that do not exhibit any phenotype when mutated in C. -

Supplementary Table 2
Supplementary Table 2. Differentially Expressed Genes following Sham treatment relative to Untreated Controls Fold Change Accession Name Symbol 3 h 12 h NM_013121 CD28 antigen Cd28 12.82 BG665360 FMS-like tyrosine kinase 1 Flt1 9.63 NM_012701 Adrenergic receptor, beta 1 Adrb1 8.24 0.46 U20796 Nuclear receptor subfamily 1, group D, member 2 Nr1d2 7.22 NM_017116 Calpain 2 Capn2 6.41 BE097282 Guanine nucleotide binding protein, alpha 12 Gna12 6.21 NM_053328 Basic helix-loop-helix domain containing, class B2 Bhlhb2 5.79 NM_053831 Guanylate cyclase 2f Gucy2f 5.71 AW251703 Tumor necrosis factor receptor superfamily, member 12a Tnfrsf12a 5.57 NM_021691 Twist homolog 2 (Drosophila) Twist2 5.42 NM_133550 Fc receptor, IgE, low affinity II, alpha polypeptide Fcer2a 4.93 NM_031120 Signal sequence receptor, gamma Ssr3 4.84 NM_053544 Secreted frizzled-related protein 4 Sfrp4 4.73 NM_053910 Pleckstrin homology, Sec7 and coiled/coil domains 1 Pscd1 4.69 BE113233 Suppressor of cytokine signaling 2 Socs2 4.68 NM_053949 Potassium voltage-gated channel, subfamily H (eag- Kcnh2 4.60 related), member 2 NM_017305 Glutamate cysteine ligase, modifier subunit Gclm 4.59 NM_017309 Protein phospatase 3, regulatory subunit B, alpha Ppp3r1 4.54 isoform,type 1 NM_012765 5-hydroxytryptamine (serotonin) receptor 2C Htr2c 4.46 NM_017218 V-erb-b2 erythroblastic leukemia viral oncogene homolog Erbb3 4.42 3 (avian) AW918369 Zinc finger protein 191 Zfp191 4.38 NM_031034 Guanine nucleotide binding protein, alpha 12 Gna12 4.38 NM_017020 Interleukin 6 receptor Il6r 4.37 AJ002942 -
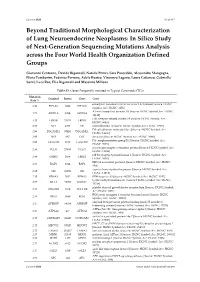
Beyond Traditional Morphological Characterization of Lung
Cancers 2020 S1 of S15 Beyond Traditional Morphological Characterization of Lung Neuroendocrine Neoplasms: In Silico Study of Next-Generation Sequencing Mutations Analysis across the Four World Health Organization Defined Groups Giovanni Centonze, Davide Biganzoli, Natalie Prinzi, Sara Pusceddu, Alessandro Mangogna, Elena Tamborini, Federica Perrone, Adele Busico, Vincenzo Lagano, Laura Cattaneo, Gabriella Sozzi, Luca Roz, Elia Biganzoli and Massimo Milione Table S1. Genes Frequently mutated in Typical Carcinoids (TCs). Mutation Original Entrez Gene Gene Rate % eukaryotic translation initiation factor 1A X-linked [Source: HGNC 4.84 EIF1AX 1964 EIF1AX Symbol; Acc: HGNC: 3250] AT-rich interaction domain 1A [Source: HGNC Symbol;Acc: HGNC: 4.71 ARID1A 8289 ARID1A 11110] LDL receptor related protein 1B [Source: HGNC Symbol; Acc: 4.35 LRP1B 53353 LRP1B HGNC: 6693] 3.53 NF1 4763 NF1 neurofibromin 1 [Source: HGNC Symbol;Acc: HGNC: 7765] DS cell adhesion molecule like 1 [Source: HGNC Symbol; Acc: 2.90 DSCAML1 57453 DSCAML1 HGNC: 14656] 2.90 DST 667 DST dystonin [Source: HGNC Symbol;Acc: HGNC: 1090] FA complementation group D2 [Source: HGNC Symbol; Acc: 2.90 FANCD2 2177 FANCD2 HGNC: 3585] piccolo presynaptic cytomatrix protein [Source: HGNC Symbol; Acc: 2.90 PCLO 27445 PCLO HGNC: 13406] erb-b2 receptor tyrosine kinase 2 [Source: HGNC Symbol; Acc: 2.44 ERBB2 2064 ERBB2 HGNC: 3430] BRCA1 associated protein 1 [Source: HGNC Symbol; Acc: HGNC: 2.35 BAP1 8314 BAP1 950] capicua transcriptional repressor [Source: HGNC Symbol; Acc: 2.35 CIC 23152 CIC HGNC: -

Zootaxa,Comparison of the Cryptic Nematode Species Caenorhabditis
Zootaxa 1456: 45–62 (2007) ISSN 1175-5326 (print edition) www.mapress.com/zootaxa/ ZOOTAXA Copyright © 2007 · Magnolia Press ISSN 1175-5334 (online edition) Comparison of the cryptic nematode species Caenorhabditis brenneri sp. n. and C. remanei (Nematoda: Rhabditidae) with the stem species pattern of the Caenorhabditis Elegans group WALTER SUDHAUS1 & KARIN KIONTKE2 1Institut für Biologie/Zoologie, AG Evolutionsbiologie, Freie Universität Berlin, Königin-Luise Straße 1-3, 14195 Berlin, Germany. [email protected] 2Department of Biology, New York University, 100 Washington Square E., New York, NY10003, USA. [email protected] Abstract The new gonochoristic member of the Caenorhabditis Elegans group, C. brenneri sp. n., is described. This species is reproductively isolated at the postmating level from its sibling species, C. remanei. Between these species, only minute morphological differences are found, but there are substantial genetic differences. The stem species pattern of the Ele- gans group is reconstructed. C. brenneri sp. n. deviates from this character pattern only in small diagnostic characters. In mating tests of C. brenneri sp. n. females with C. remanei males, fertilization takes place and juveniles occasionally hatch. In the reverse combination, no offspring were observed. Individuals from widely separated populations of each species can be crossed successfully (e.g. C. brenneri sp. n. populations from Guadeloupe and Sumatra, or C. remanei populations from Japan and Germany). Both species have been isolated only from anthropogenic habitats, rich in decom- posing organic material. C. brenneri sp. n. is distributed circumtropically, C. remanei is only found in northern temperate regions. To date, no overlap of the ranges was found. -

Non-Classical Ligand-Independent Regulation of Go Protein by An
Non-classical ligand-independent regulation of Go protein by an orphan Class C GPCR Mariana Hajj, Teresa de Vita, Claire Vol, Charlotte Renassia, Jean-Charles Bologna, Isabelle Brabet, Magali Cazade, Manuela Pastore, Jaroslav Blahos, Gilles Labesse, et al. To cite this version: Mariana Hajj, Teresa de Vita, Claire Vol, Charlotte Renassia, Jean-Charles Bologna, et al.. Non- classical ligand-independent regulation of Go protein by an orphan Class C GPCR. Molecular Pharma- cology, American Society for Pharmacology and Experimental Therapeutics, 2019, 96 (2), pp.233-246. 10.1124/mol.118.113019. hal-02396282 HAL Id: hal-02396282 https://hal.archives-ouvertes.fr/hal-02396282 Submitted on 5 Dec 2019 HAL is a multi-disciplinary open access L’archive ouverte pluridisciplinaire HAL, est archive for the deposit and dissemination of sci- destinée au dépôt et à la diffusion de documents entific research documents, whether they are pub- scientifiques de niveau recherche, publiés ou non, lished or not. The documents may come from émanant des établissements d’enseignement et de teaching and research institutions in France or recherche français ou étrangers, des laboratoires abroad, or from public or private research centers. publics ou privés. MOL #113019 Non-classical ligand-independent regulation of Go protein by an orphan Class C GPCR Mariana Hajj, Teresa De Vita, Claire Vol, Charlotte Renassia, Jean-Charles Bologna, Isabelle Brabet, Magali Cazade, Manuela Pastore, Jaroslav Blahos, Gilles Labesse, Jean-Philippe Pin, Laurent Prézeau IGF, Univ. Montpellier, CNRS, INSERM, Montpellier, France: MH, TDV, CV, CR, JCB, IB, MC, MP, JPP, LP Institute of Molecular Genetics, Academy of Sciences of the Czech Republic and Department of Pharmacology, 2nd Medical School, Charles University, Prague, Czech Republic: JB CBS, Univ. -

Reproductive Mode and the Evolution of Genome Size and Structure in Caenorhabditis Nematodes
RESEARCH ARTICLE Reproductive Mode and the Evolution of Genome Size and Structure in Caenorhabditis Nematodes Janna L. Fierst1¤a, John H. Willis1, Cristel G. Thomas2, Wei Wang2, Rose M. Reynolds1¤b, Timothy E. Ahearne1, Asher D. Cutter2, Patrick C. Phillips1* 1 Institute of Ecology and Evolution, University of Oregon, Eugene, Oregon, United States of America, 2 Department of Ecology and Evolutionary Biology and Centre for the Analysis of Genome Evolution and Function, University of Toronto, Ontario, Canada a11111 ¤a Current Address: Department of Biological Sciences, The University of Alabama, Tuscaloosa, Alabama, USA ¤b Current Address: Department of Biology, William Jewell College, Liberty, Missouri USA * [email protected] Abstract OPEN ACCESS Citation: Fierst JL, Willis JH, Thomas CG, Wang W, The self-fertile nematode worms Caenorhabditis elegans, C. briggsae, and C. tropicalis Reynolds RM, Ahearne TE, et al. (2015) evolved independently from outcrossing male-female ancestors and have genomes 20- Reproductive Mode and the Evolution of Genome 40% smaller than closely related outcrossing relatives. This pattern of smaller genomes for Size and Structure in Caenorhabditis Nematodes. PLoS Genet 11(6): e1005323. doi:10.1371/journal. selfing species and larger genomes for closely related outcrossing species is also seen in pgen.1005323 plants. We use comparative genomics, including the first high quality genome assembly for Editor: Mark Blaxter, University of Edinburgh, an outcrossing member of the genus (C. remanei) to test several hypotheses for the evolu- UNITED KINGDOM tion of genome reduction under a change in mating system. Unlike plants, it does not appear Received: July 29, 2014 that reductions in the number of repetitive elements, such as transposable elements, are an important contributor to the change in genome size.