A Resource for Membrane-Embedded Protein Structures and Their Lipid Interactions
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-
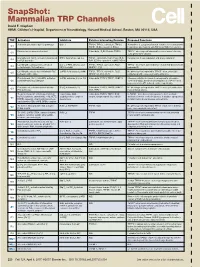
Snapshot: Mammalian TRP Channels David E
SnapShot: Mammalian TRP Channels David E. Clapham HHMI, Children’s Hospital, Department of Neurobiology, Harvard Medical School, Boston, MA 02115, USA TRP Activators Inhibitors Putative Interacting Proteins Proposed Functions Activation potentiated by PLC pathways Gd, La TRPC4, TRPC5, calmodulin, TRPC3, Homodimer is a purported stretch-sensitive ion channel; form C1 TRPP1, IP3Rs, caveolin-1, PMCA heteromeric ion channels with TRPC4 or TRPC5 in neurons -/- Pheromone receptor mechanism? Calmodulin, IP3R3, Enkurin, TRPC6 TRPC2 mice respond abnormally to urine-based olfactory C2 cues; pheromone sensing 2+ Diacylglycerol, [Ca ]I, activation potentiated BTP2, flufenamate, Gd, La TRPC1, calmodulin, PLCβ, PLCγ, IP3R, Potential role in vasoregulation and airway regulation C3 by PLC pathways RyR, SERCA, caveolin-1, αSNAP, NCX1 La (100 µM), calmidazolium, activation [Ca2+] , 2-APB, niflumic acid, TRPC1, TRPC5, calmodulin, PLCβ, TRPC4-/- mice have abnormalities in endothelial-based vessel C4 i potentiated by PLC pathways DIDS, La (mM) NHERF1, IP3R permeability La (100 µM), activation potentiated by PLC 2-APB, flufenamate, La (mM) TRPC1, TRPC4, calmodulin, PLCβ, No phenotype yet reported in TRPC5-/- mice; potentially C5 pathways, nitric oxide NHERF1/2, ZO-1, IP3R regulates growth cones and neurite extension 2+ Diacylglycerol, [Ca ]I, 20-HETE, activation 2-APB, amiloride, Cd, La, Gd Calmodulin, TRPC3, TRPC7, FKBP12 Missense mutation in human focal segmental glomerulo- C6 potentiated by PLC pathways sclerosis (FSGS); abnormal vasoregulation in TRPC6-/- -

Study of Selectivity and Permeation in Voltage-Gated Ion Channels
Study of Selectivity and Permeation in Voltage-Gated Ion Channels By Janhavi Giri, Ph.D. Visiting Research Faculty Division of Molecular Biophysics and Physiology Rush University Medical Center Chicago, IL, USA Email: [email protected] i TABLE OF CONTENTS Page TABLE OF CONTENTS ..................................................................................................... i LIST OF TABLES ............................................................................................................. iii LIST OF FIGURES ........................................................................................................... iv SUMMARY ............................................................................................................... xvi CHAPTER 1. INTRODUCTION ....................................................................................... 1 1.1. Background ................................................................................................... 1 1.2. Overview .................................................................................................... 10 CHAPTER 2. SELF-ORGANIZED MODELS OF SELECTIVITY IN CALCIUM CHANNELS ........................................................................................... 13 2.1. Introduction ................................................................................................ 13 2.2. Methods ...................................................................................................... 19 2.2.1. Model of Channel and Electrolyte ........................................................19 -

An Aquaporin-4/Transient Receptor Potential Vanilloid 4 (AQP4/TRPV4) Complex Is Essential for Cell-Volume Control in Astrocytes
An aquaporin-4/transient receptor potential vanilloid 4 (AQP4/TRPV4) complex is essential for cell-volume control in astrocytes Valentina Benfenatia,1, Marco Caprinib,1, Melania Doviziob, Maria N. Mylonakouc, Stefano Ferronib, Ole P. Ottersenc, and Mahmood Amiry-Moghaddamc,2 aInstitute for Nanostructured Materials, Consiglio Nazionale delle Ricerche, 40129 Bologna, Italy; bDepartment of Human and General Physiology, University of Bologna, 40127 Bologna, Italy; and cCenter for Molecular Biology and Neuroscience and Department of Anatomy, University of Oslo, 0317 Oslo, Norway Edited* by Peter Agre, Johns Hopkins Malaria Research Institute, Baltimore, MD, and approved December 27, 2010 (received for review September 1, 2010) Regulatory volume decrease (RVD) is a key mechanism for volume channels (VRAC). Osmolyte efflux through VRAC is thought to control that serves to prevent detrimental swelling in response to provide the driving force for water exit (6, 7). The factors that hypo-osmotic stress. The molecular basis of RVD is not understood. initiate RVD have received comparatively little attention. Here Here we show that a complex containing aquaporin-4 (AQP4) and we test our hypothesis that activation of astroglial RVD depends transient receptor potential vanilloid 4 (TRPV4) is essential for RVD on a molecular interaction between AQP4 and TRPV4. fi in astrocytes. Astrocytes from AQP4-KO mice and astrocytes treated Our hypothesis rests on our recent nding that TRPV4 is with TRPV4 siRNA fail to respond to hypotonic stress by increased strongly expressed in astrocytic endfeet membranes abutting the – intracellular Ca2+ and RVD. Coimmunoprecipitation and immunohis- pia (including the extension of the pia that lines the Virchow tochemistry analyses show that AQP4 and TRPV4 interact and coloc- Robin spaces) and in endfeet underlying ependyma of the ven- alize. -

Gadolinium Chloride Restores the Function of the Gap Junctional Intercellular Communication Between Hepatocytes in a Liver Injury
Article Gadolinium Chloride Restores the Function of the Gap Junctional Intercellular Communication between Hepatocytes in a Liver Injury Le Yang, Chengbin Dong, Lei Tian, Xiaofang Ji, Lin Yang and Liying Li * Department of Cell Biology, Municipal Laboratory for Liver Protection and Regulation of Regeneration, Capital Medical University, Beijing 100069, China * Correspondence: [email protected]; Tel.: +0086-10-83950468; Fax: +0086-10-83950468; Received: 3 July 2019; Accepted: 30 July 2019; Published: 31 July 2019 Abstract: Background: Gadolinium chloride (GdCl3) has been reported to attenuate liver injury caused by a variety of toxicants. Gap junctional intercellular communication (GJIC) is thought to be essential in controlling liver homeostasis and pathology. Here we evaluate the effects of GdCl3 on functional GJIC and connexin expression in mouse models and primary hepatocytes. Methods: Mice were administered GdCl3 intraperitoneally the day before a carbon tetrachloride (CCl4) injection or bile duct ligation (BDL) operation. Primary hepatocytes were treated with CCl4 or lipopolysaccharides (LPS), with or without GdCl3. A scrape loading/dye transfer assay was performed to assess the GJIC function. The expression of connexins was examined by real-time reverse transcription polymerase chain reaction (RT-PCR), western blot and immunofluorescent staining. Results: CCl4 treatment or the BDL operation led to the dysfunction of GJIC and a down-regulation of Cx32 and Cx26 in injured liver. GdCl3 administration restored GJIC function between hepatocytes by facilitating the transfer of fluorescent dye from one cell into adjacent cells via GJIC, and markedly prevented the decrease of Cx32 and Cx26 in injured liver. In primary hepatocytes, CCl4 or LPS treatment induced an obvious decline of Cx32 and Cx26, whereas GdCl3 pretreatment prevented the down-regulation of connexins. -

Labned.Com Bovine
Antibodies Predicted to Bind Homologous Bovine Targets This is a catalog of all antibodies predicted to react with bovine proteins in addition to their validated reactivities. This catalog was assembled by identifying proteins targeted by LabNed antibodies that share 99% sequence homology withtheir bovine counterparts. Table of Contents High Affinity Antibodies .................................................................................... 1 High Sensitivity ELISA Kits .................................................................................... 31 High Affinity Antibodies LabNed supplies only the highest quality antibodies. Our high-affinity polyclonal and monoclonal antibodies are thoroughly validated by Western Blotting, Immunohistochemistry and ELISA. This is our comprehensive catalog of our antibody products predicted to react with bovine proteins, sorted in alphabetical order by target gene name. Applications Gene Name Product Name Reactivity WB ABAT Anti-ABA Human, Mouse, Rat WB ABCA1 Anti-ABCA1 Human, Mouse, Rat WB ABCA1 Anti-ABCA1 Human, Mouse, Rat IHC-P, WB ABCB11 Anti-ABCB11 Human, Mouse, Rat IHC-P, WB ABCC1 Anti-MRP1 Human, Mouse, Rat WB ABCC4 Anti-MRP4 Human, Mouse, Rat IHC-P, WB ABCD3 Anti-PMP70 Human, Mouse, Rat WB ABCE1 Anti-ABCE1 Human, Mouse, Rat WB ABCG1 Anti-ABCG1 Human, Mouse, Rat WB ABCG2 Anti-ABCG2 Human, Mouse, Rat WB ABCG4 Anti-ABCG4 Human, Mouse, Rat WB ABCG5 Anti-ABCG5 Human IHC-P, ICC, WB ABI1 Anti-ABI1 Human, Mouse, Rat WB ABI2 Anti-ABI2 Human, Mouse, Rat WB ACO2 Anti-Aconitase 2 Human, Mouse, Rat -

Transyt, the Transport Systems Tracker
bioRxiv preprint doi: https://doi.org/10.1101/2021.04.29.441738; this version posted April 30, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-ND 4.0 International license. TranSyT, the Transport Systems Tracker Davide Lagoa1,2, José P. Faria2, Filipe Liu2, Emanuel Cunha1, Christopher S. Henry2 and Oscar Dias1,* 1 Centre of Biological Engineering, University of Minho, Braga, 4704-553, Portugal 2 Computing, Environment, and Life Sciences Division, Argonne National Laboratory, Lemont, IL 60439, USA * To whom correspondence should be addressed. Tel: +351 253 604422; Fax: ; Email: [email protected] ABSTRACT The importance and rate of development of GSM models have been growing for the last years, increasing the demand for software solutions that automatise several steps of this process. However, since TRIAGE’s release, software development for automatic integration of transport reactions into models has stalled. Here we present the Transport Systems Tracker (TranSyT), the next iteration of TRIAGE. Unlike its predecessor, TranSyT does not rely on manual curation to expand its internal database, derived from highly-curated records retrieved from TCDB, and complemented with information from other data sources. TranSyT compiles information regarding, TC families, transport proteins, and derives reactions into its internal database, making it available for rapid annotation of complete genomes. All transport reactions have GPR associations and can be exported with identifiers from four different metabolite databases. TranSyT is currently available as a plugin for merlin v4.0 and an app for KBase. -

X-Ray Structure of Human Aquaporin 2 and Its Implications for Nephrogenic Diabetes Insipidus and Trafficking
X-ray structure of human aquaporin 2 and its implications for nephrogenic diabetes insipidus and trafficking Anna Fricka, Urszula Kosinska Erikssona, Fabrizio de Mattiab, Fredrik Öberga, Kristina Hedfalka, Richard Neutzea, Willem J. de Gripc, Peter M. T. Deenb, and Susanna Törnroth-Horsefielda,d,1 aDepartment of Chemistry and Molecular Biology, University of Gothenburg, 405 30 Gothenburg, Sweden; bDepartment of Physiology, Radboud University Medical Center, 6500 HB, Nijmegen, The Netherlands; cDepartment of Biochemistry, Radboud Institute for Molecular Life Sciences, Radboud University Medical Center, G525 GA, Nijmegen, The Netherlands; and dDepartment of Biochemistry and Structural Biology, Centre for Molecular Protein Science, Lund University, 221 00 Lund, Sweden Edited* by Robert M. Stroud, University of California, San Francisco, CA, and approved March 20, 2014 (received for review November 15, 2013) Human aquaporin 2 (AQP2) is a water channel found in the kidney Because of its central role in water homeostasis, dysregulation collecting duct, where it plays a key role in concentrating urine. of AQP2 is implicated in several human disease states including Water reabsorption is regulated by AQP2 trafficking between congestive heart failure, liver cirrhosis, and preeclampsia (13). intracellular storage vesicles and the apical membrane. This pro- Failure to recruit AQP2 to the apical membrane underlies ac- cess is tightly controlled by the pituitary hormone arginine vaso- quired and congenital nephrogenic diabetes insipidus (NDI), pressin and defective trafficking results in nephrogenic diabetes a water balance disorder in which patients lack the ability to insipidus (NDI). Here we present the X-ray structure of human concentrate urine, leading to severe dehydration (14). Although AQP2 at 2.75 Å resolution. -

(10) Patent No.: US 7857978 B2
US007857978B2 (12) United States Patent (10) Patent No.: US 7,857,978 B2 Jensen et a]. (45) Date of Patent: Dec. 28, 2010 (54) MEMBRANE FOR FILTERING OF WATER (58) Field of Classi?cation Search .......... .. 210/ 500.35, 210/500.36, 500.42, 641, 321.75, 257.2, (75) Inventors: Peter Holme Jensen, Copenhagen (DK); 210/500.2, 7, 490, 6504653, 655, 500.1, Danielle Keller, Odense (DK); Claus 210/500.29, 900; 435/17.1, 4; 436/524, 436/172, 82.05; 530/402; 425/450, 1.21, HéliX Nielsen, Taastrup (DK) 425/9, 32, 417, 489; 264/4.1, 4.3, 4.6, 41; 424/450; 977/7l3i7l4, 718; 422/101 (73) Assignee: Aquaporin A/S, Copehhagen N (DK) See application ?le for complete search history. ( * ) Notice: Subject to any disclaimer, the term of this (56) References Cited patent is extended or adjusted under 35 U.S.C. 154(b) by 0 days. U.S. PATENT DOCUMENTS 3,906,250 A 9/1975 Loeb (21) Appl. No.: 11/915,121 (Continued) (22) PCT Filed: May 19, 2006 FOREIGN PATENT DOCUMENTS EP 1885477 Bl 2/2010 (86) PCT No.: PCT/DK2006/000278 W0 WO 02/13955 2/2002 W0 WO 2004/099088 11/2004 § 371 (0X1)’ W0 WO 2007/033675 3/2007 (2), (4) Date: May 12, 2008 OTHER PUBLICATIONS Coury et al., Reconstintution of Water channel function of aquaporins (87) PCT Pub. No.: WO2006/122566 l and 2 by expression in yeast secretory vesicles, 1998, The American Physiological Society, pp. F34 -F42 .* PCT Pub. Date: Nov. 23, 2006 (Continued) (65) Prior Publication Data Primary ExamineriTony G Soohoo Assistant ExamineriDavid C Mellon US 2009/0120874 A1 May 14, 2009 (74) Attorney, Agent, or FirmiKristina Bieker-Brady; Clark & Elbing LLP Related U.S. -

Phosphate-Selective Porins from the Outer Membranes of Fluorescent Pseudomonas Sp
Phosphate-selective porins from the outer membranes of fluorescent Pseudomonas sp. KEITHPOOLE, THOMAS R. PARR,JR., AND ROBERTE. W. HANCOCK Department of Microbiology, University of British Columbia, Vancouver, B.C., Canada V6T I W5 Received July 22, 1986 Accepted October 14, 1986 POOLE,K., T. R. PARR,JR., and R. E. W. HANCOCK.1987. Phosphate-selective porins from the outer membranes of fluorescent Pseudomonas sp. Can. J. Microbiol . 33: 63-69. Phosphate starvation induced oligomeric proteins from the outer membranes of Pseudomonas Juorescens, Pseudomonas putida, Pseudomonas aureofaciens, and Pseudomonas chlororaphis were purified to homogeneity. The incorporation of the purified proteins into planar lipid bilayer membranes resulted in stepwise increases in membrane conductance. Single channel conductance experiments demonstrated that these proteins were all capable of forming small channels, similar to the Pseudo- monas aeruginosa phosphate porin protein P, with average single channel conductances in I M KC1 of between 233 and 252 pS. Single channel conductance measurements made in salts of varying cation or anion size indicated that the channels were uni- formly anion selective. The measurement of single channel conductance as a function of KC1 concentration revealed that all channels saturated at higher salt concentrations, consistent with the presence of an anion-binding site in the channel. Apparent Kd values for C1- binding were calculated and shown to vary only twofold (180-297 mM) among all channels, including protein P channels. Phosphate competitively inhibited chloride conductance through these channels with apparent 150 values of between 0.59 and 2.5 mM phosphate at 40 mM C1- and between 9.7 and 27 mM phosphate at 1 M C1-. -

The Importance of Porins and Β-Lactamase in Outer Membrane Vesicles on the Hydrolysis of Β-Lactam Antibiotics
International Journal of Molecular Sciences Article The Importance of Porins and β-Lactamase in Outer Membrane Vesicles on the Hydrolysis of β-Lactam Antibiotics Si Won Kim 1, Jung Seok Lee 1, Seong Bin Park 2, Ae Rin Lee 1, Jae Wook Jung 1, Jin Hong Chun 1, Jassy Mary S. Lazarte 1, Jaesung Kim 1, Jong-Su Seo 3, Jong-Hwan Kim 3, Jong-Wook Song 3, Min Woo Ha 4, Kim D. Thompson 5, Chang-Ro Lee 6 , Myunghwan Jung 7 and Tae Sung Jung 1,8,* 1 Laboratory of Aquatic Animal Diseases, Institute of Animal Medicine, College of Veterinary Medicine, Gyeongsang National University, Jinju 52828, Korea; [email protected] (S.W.K.); [email protected] (J.S.L.); gladofl[email protected] (A.R.L.); [email protected] (J.W.J.); [email protected] (J.H.C.); [email protected] (J.M.S.L.); [email protected] (J.K.) 2 Coastal Research & Extension Center, Mississippi State University, Starkville, MS 39567, USA; [email protected] 3 Environmental Chemistry Research Center, Korea Institute of Toxicology Gyeongnam Department of Environmental Toxicology and Chemistry, Jinju 52834, Korea; [email protected] (J.-S.S.); [email protected] (J.-H.K.); [email protected] (J.-W.S.) 4 College of Pharmacy, Jeju National University, 102, Jejudaehak-ro, Jeju-si, Jeju-do 63243, Korea; [email protected] 5 Moredun Research Institute, Pentlands Science Park, Penicuik, Midlothian EH26 0PZ, UK; [email protected] 6 Department of Biological Sciences, Myongji University, Yongin, Gyeonggido 449-728, Korea; [email protected] 7 Department of Microbiology, Research Institute of Life Sciences, College of Medicine, Gyeongsang National University, Jinju 52727, Korea; [email protected] 8 Centre for Marine Bioproducts Development, College of Medicine and Public Health, Flinders, University, Bedford Park, Adelaide, SA 5042, Australia * Correspondence: [email protected]; Tel.: +82-10-8545-9310; Fax: +82-55-762-6733 Received: 13 February 2020; Accepted: 16 April 2020; Published: 17 April 2020 Abstract: Gram-negative bacteria have an outer membrane inhibiting the entry of antibiotics. -

Selective Permeability of Gap Junction Channels
View metadata, citation and similar papers at core.ac.uk brought to you by CORE provided by Elsevier - Publisher Connector Biochimica et Biophysica Acta 1662 (2004) 96–101 www.bba-direct.com Review Selective permeability of gap junction channels Gary S. Goldberg*, Virginijus Valiunas, Peter R. Brink Department of Physiology and Biophysics, School of Medicine, State University of New York at Stony Brook, Basic Science Tower L 6, Health Science Complex, Stony Brook, NY 11794-8661, USA Received 23 June 2003; accepted 21 November 2003 Abstract Gap junctions mediate the transfer of small cytoplasmic molecules between adjacent cells. A family of gap junction proteins exist that form channels with unique properties, and differ in their ability to mediate the transfer of specific molecules. Mutations in a number of individual gap junction proteins, called connexins, cause specific human diseases. Therefore, it is important to understand how gap junctions selectively move molecules between cells. Rules that dictate the ability of a molecule to travel through gap junction channels are complex. In addition to molecular weight and size, the ability of a solute to transverse these channels depends on its net charge, shape, and interactions with specific connexins that constitute gap junctions in particular cells. This review presents some data and interpretations pertaining to mechanisms that govern the differential transfer of signals through gap junction channels. D 2004 Elsevier B.V. All rights reserved. Keywords: Gap junction; Connexin; Cell communication; Gap junctional communication; Permselectivity; Intercellular channel; Membrane channel; Ion channel 1. Introduction ions through gap junctions appears to be paramount to coordinated function [10–12]. -

Energetics of Discrete Selectivity Bands and Mutation-Induced Transitions in the Calcium-Sodium Ion Channels Family
PHYSICAL REVIEW E 88, 052712 (2013) Energetics of discrete selectivity bands and mutation-induced transitions in the calcium-sodium ion channels family I. Kaufman,1 D. G. Luchinsky,1,2 R. Tindjong,1 P. V. E. McClintock,1,* and R. S. Eisenberg3 1Department of Physics, Lancaster University, Lancaster LA1 4YB, United Kingdom 2Mission Critical Technologies Inc., 2041 Rosecrans Ave. Suite 225 El Segundo, California 90245, USA 3Department of Molecular Biophysics and Physiology, Rush Medical College, 1750 West Harrison, Chicago, Illinois 60612, USA (Received 8 May 2013; published 19 November 2013) We use Brownian dynamics (BD) simulations to study the ionic conduction and valence selectivity of a generic electrostatic model of a biological ion channel as functions of the fixed charge Qf at its selectivity filter. We are thus able to reconcile the discrete calcium conduction bands recently revealed in our BD simulations, M0 (Qf = 1e), M1 (3e), M2 (5e), with a set of sodium conduction bands L0 (0.5e), L1 (1.5e), thereby obtaining a completed pattern of conduction and selectivity bands vs Qf for the sodium-calcium channels family. An increase of Qf leads to an increase of calcium selectivity: L0 (sodium-selective, nonblocking channel) → M0 (nonselective channel) → L1 (sodium-selective channel with divalent block) → M1 (calcium-selective channel exhibiting the anomalous mole fraction effect). We create a consistent identification scheme where the L0 band is putatively identified with the eukaryotic sodium channel The scheme created is able to account for the experimentally observed mutation-induced transformations between nonselective channels, sodium- selective channels, and calcium-selective channels, which we interpret as transitions between different rows of the identification table.