The Role of 4E-T in Translational Regulation and Mrna Decay
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Signature Redacted Signatureredacted
The biochemical basis for the cooperative action of microRNAs S ISTITUTE By By MAssACHUSEOFTECHNOLOGY > Daniel Briskin OCT 0 12019 B.S., Molecular Genetics (2012) LIBRARIES 0 Ohio State University SUBMITTED TO THE DEPARTMENT OF BIOLOGY IN PARTIAL FULFILLMENT OF THE REQUIREMENTS FOR THE DEGREE OF DOCTOR OF PHILOSOPHY AT THE MASSACHUSETTS INSTITUTE OF TECHNOLOGY -SEPTEMBER-2019- -fve e 7-1012 © 2019 Massachusetts Institute of Technology All rights reserved Signature redacted Signature of Author: DanielBriskin Department of Biology 17 September 2019 Signature redacted Certified By:__ David P. Bartel Professor of Biology Thesis Supervisor Accepted By: Signatureredacted -V Stephen Bell Uncas and Helen Whitaker Professor of Biology Investigator, Howard Hughes Medical Institute Co-Director, Biology Graduate Committee 1 2 The biochemical basis for the cooperative action of microRNAs by Daniel Briskin Submitted to the Department of Biology on 17 September 2019 In partial fulfillment of the requirements for the degree of doctor of philosophy Abstract In metazoans, microRNAs (miRNAs) act to repress mRNAs through a combination of translational repression and target degradation. miRNAs predominantly pair within the 3' untranslated region (3' UTR) of the mRNA. In cells, closely spaced miRNA target sites within an mRNA can act cooperatively, leading to more repression of the target mRNA than expected by independent action at each site. This dissertation details the use of purified miRNA-AGO2 complexes, synthetic target RNAs, and a purified domain of TNRC6B that is able to simultaneously bind multiple AGO proteins. We examined the target site occupancy and affinities for miRNA-AGO2 binding in the absence and presence of TNRC6B, for target RNAs with a single miRNA site as well as multiple miRNA sites spaced at varying distances. -

News & Views Research
NEWS & VIEWS RESEARCH may exhibit some tissue specificity in humans. Upstream intron Downstream intron a Sibley et al. found that genes with long introns sequence sequence tend to be expressed in the human nervous Precursor RNA system, and they identified recursively spliced 3' Splice 5' Splice RNAs expressed in the human brain6. Duff site site First step et al. detected some selectivity for recursive splicing in the brain in a screen of 20 human tissues (including fetal brain and adult cerebel- Second step lum), but this may partly reflect the difficulty of detecting recursively spliced RNAs in tis- Mature mRNA sues that express such RNAs at low levels. It will be important to determine whether this b RS exon specificity, if real, results from the tendency of recursively spliced genes to be expressed in the brain, or whether cells in the nervous system have factors that promote recursive Competing 5' splice sites First step splicing. Many genes that have long introns, including those that undergo recursive splicing, are linked to neurological diseases and to Second step 9–11 Second step autism . Whether these conditions are sometimes triggered by errors in the multi- step recursive RNA-splicing process will be an exciting avenue for future studies. ■ NMD Heidi Cook-Andersen and Miles F. Wilkinson are in the Department of Reproductive Medicine, University of Figure 1 | Mechanisms of recursive splicing. a, In recursive splicing, long intron sequences of precursor California, San Diego, La Jolla, California, RNA are removed in a stepwise process mediated by juxtaposed internal 3ʹ and 5ʹ splice sites. In the first step, 92093, USA. -

(Bsc Zoology and Microbiology) Concept of Introns and Exons
Unit-5 Molecular Biology (BSc Zoology and Microbiology) Concept of introns and exons Most of the portion of a gene in higher eukaryotes consists of noncoding DNA that interrupts the relatively short segments of coding DNA. The coding sequences are called exons. The noncoding sequences are called introns. Intron: An intron is a portion of a gene that does not code for amino acids An intron is any nucleotide sequence within a gene which is represented in the primary transcript of the gene, but not present in the final processed form. In other words, Introns are noncoding regions of an RNA transcript which are eliminated by splicing before translation. Sequences that are joined together in the final mature RNA after RNA splicing are exons. Introns are very large chunks of RNA within a messenger RNA molecule that interfere with the code of the exons. And these introns get removed from the RNA molecule to leave a string of exons attached to each other so that the appropriate amino acids can be encoded for. Introns are rare in genes of prokaryotes. #Look carefully at the diagram above, we have already discussed about the modification and processing of eukaryotic RNA. In which 5’ guanine cap and 3’poly A tail is added. So at that time, noncoding regions i.e. introns are removed. We hv done ths already. Ok Exon: The coding sequences are called Exon. An exon is the portion of a gene that codes for amino acids. In the cells of plants and animals, most gene sequences are broken up by one or more DNA sequences called introns. -

Current Perspectives in Intronic Micro Rnas (Mirnas)
Journal of Biomedical Science (2006) 13:5–15 5 DOI 10.1007/s11373-005-9036-8 Current perspectives in intronic micro RNAs (miRNAs) Shao-Yao Ying & Shi-Lung Lin Department of Cell & Neurobiology, Keck School of Medicine, BMT-403, University of Southern California, 1333 San Pablo Street, Los Angeles, CA, 90033, USA Received 27 May 2005; accepted 14 September 2005 Ó 2005 National Science Council, Taipei Key words: fine-tuning of gene function, functional/structural genomics, gene expression, genetic regula- tion, intronic microRNA, miRNA biogenesis, miRNA, post-translational modification, regulatory gene Summary MicroRNAs (miRNAs), small single-stranded regulatory RNAs capable of interfering with intracellular messenger RNAs (mRNAs) that contain either complete or partial complementarity, are useful for the design of new therapies against cancer polymorphism and viral mutation. Numerous miRNAs have been reported to induce RNA interference (RNAi), a post-transcriptional gene silencing mechanism. Intronic miRNAs, derived from introns by RNA splicing and Dicer processing, can interfere with intracellular mRNAs to silence that gene expression. The intronic miRNAs differ uniquely from previously described intergenic miRNAs in the requirement of type II RNA polymerases (Pol-II) and spliceosomal components for its biogenesis. Several kinds of intronic miRNAs have been identified in Caenorhabditis elegans, mouse and human cells; however, neither their function nor application has been reported. To this day, the computer searching program for miRNA seldom include the intronic portion of protein-coding RNAs. The functional significance of artificially generated intronic miRNAs has been successfully ascertained in several biological systems such as zebrafishes, chicken embryos and adult mice, indicating the evolutionary pres- ervation of this gene regulation system in vivo. -

A Set of Regulatory Genes Co-Expressed in Embryonic Human Brain Is Implicated in Disrupted Speech Development
Molecular Psychiatry https://doi.org/10.1038/s41380-018-0020-x ARTICLE A set of regulatory genes co-expressed in embryonic human brain is implicated in disrupted speech development 1 1 1 2 3 Else Eising ● Amaia Carrion-Castillo ● Arianna Vino ● Edythe A. Strand ● Kathy J. Jakielski ● 4,5 6 7 8 9 Thomas S. Scerri ● Michael S. Hildebrand ● Richard Webster ● Alan Ma ● Bernard Mazoyer ● 1,10 4,5 6,11 6,12 13 Clyde Francks ● Melanie Bahlo ● Ingrid E. Scheffer ● Angela T. Morgan ● Lawrence D. Shriberg ● Simon E. Fisher 1,10 Received: 22 September 2017 / Revised: 3 December 2017 / Accepted: 2 January 2018 © The Author(s) 2018. This article is published with open access Abstract Genetic investigations of people with impaired development of spoken language provide windows into key aspects of human biology. Over 15 years after FOXP2 was identified, most speech and language impairments remain unexplained at the molecular level. We sequenced whole genomes of nineteen unrelated individuals diagnosed with childhood apraxia of speech, a rare disorder enriched for causative mutations of large effect. Where DNA was available from unaffected parents, CHD3 SETD1A WDR5 fi 1234567890();,: we discovered de novo mutations, implicating genes, including , and . In other probands, we identi ed novel loss-of-function variants affecting KAT6A, SETBP1, ZFHX4, TNRC6B and MKL2, regulatory genes with links to neurodevelopment. Several of the new candidates interact with each other or with known speech-related genes. Moreover, they show significant clustering within a single co-expression module of genes highly expressed during early human brain development. This study highlights gene regulatory pathways in the developing brain that may contribute to acquisition of proficient speech. -

RNA Splice Sites Classification Using Convolutional Neural Network Models
RNA Splice Sites Classification Using Convolutional Neural Network Models Thanyathorn Thanapattheerakul Worrawat Engchuan Daniele Merico School of Information Technology, The Centre for Applied Genomics, Molecular Diagnostics, King Mongkut’s University of Genetics and Genome Biology, The Deep Genomics, Technology Thonburi, Hospital for Sick Children, Toronto, Ontario, Canada Bangkok, Thailand Toronto, Ontario, Canada [email protected] [email protected] [email protected] Narumol Doungpan Kiyota Hashimoto Jonathan H. Chan Faculty of Engineering, Faculty of Technology and School of Information Technology, King Mongkut’s University of Environment, King Mongkut’s University of Technology Thonburi, Prince of Songkla University, Technology Thonburi, Bangkok, Thailand Phuket, Thailand Bangkok, Thailand [email protected] [email protected] [email protected] Abstract—RNA splicing refers to the elimination of non- completely make it loss of function. The alternative splicing coding region on transcribed pre-messenger ribonucleic acid can produce different functional proteins, which could lead (RNA). Identifying splicing site is an essential step which can to causing abnormal states in human [3]. be used to gain novel insights of alternative splicing as well as Many studies have proposed models to recognize the splicing defects, potentially cause malfunction of protein splice sites to reveal which splice sites contain a mutation resulting from mutations at splice site. In this work, we that may cause a splicing error. One common method to propose a data preprocessing step applying to RNA sequences recognize binding sites in motif sequences is called Position- and the models leveraging Convolutional Neural Network (CNN). The preprocessing step includes reducing sequence Weight-Matrix (PWM). -

The CTD Role in Cotranscriptional RNA Processing and Surveillance
CORE Metadata, citation and similar papers at core.ac.uk Provided by Elsevier - Publisher Connector FEBS Letters 582 (2008) 1971–1976 Minireview The CTD role in cotranscriptional RNA processing and surveillance Se´rgio F. de Almeida, Maria Carmo-Fonseca* Instituto de Medicina Molecular, Faculdade de Medicina, Universidade de Lisboa, 1649-028 Lisboa, Portugal Received 4 April 2008; revised 13 April 2008; accepted 14 April 2008 Available online 22 April 2008 Edited by Ulrike Kutay proteins specific to each snRNP. The selection of specific splice Abstract In higher eukaryotes, the production of mature mes- senger RNA that exits the nucleus to be translated into protein sites (ss) on a particular pre-mRNA substrate relies on an intri- requires precise and extensive processing of the nascent tran- cate interplay involving the cooperative binding of trans-acting script. The processing steps include 50-end capping, splicing, splicing proteins to cis-acting sequence elements in the pre- and 30-end formation. Pre-mRNA processing is coupled to tran- mRNA. In mammals, an intron is defined by four short and scription by mechanisms that are not well understood but involve poorly conserved consensus sequences: the exon–intron junc- the carboxyl-terminal domain (CTD) of the largest subunit of tions (50ss and 30ss); the branch point sequence; and the poly- RNA polymerase II. This review focuses on recent findings that pyrimidine tract (Table 1). These sequences are recognized by provide novel insight into the role of the CTD in promoting RNA base pairing with the spliceosomal snRNAs (Fig. 1). In addi- processing and surveillance. tion, both exons and introns contain weak binding sites for a Ó 2008 Federation of European Biochemical Societies. -

Autism and Cancer Share Risk Genes, Pathways, and Drug Targets
TIGS 1255 No. of Pages 8 Forum Table 1 summarizes the characteristics of unclear whether this is related to its signal- Autism and Cancer risk genes for ASD that are also risk genes ing function or a consequence of a second for cancers, extending the original finding independent PTEN activity, but this dual Share Risk Genes, that the PI3K-Akt-mTOR signaling axis function may provide the rationale for the (involving PTEN, FMR1, NF1, TSC1, and dominant role of PTEN in cancer and Pathways, and Drug TSC2) was associated with inherited risk autism. Other genes encoding common Targets for both cancer and ASD [6–9]. Recent tumor signaling pathways include MET8[1_TD$IF],[2_TD$IF] genome-wide exome-sequencing studies PTK7, and HRAS, while p53, AKT, mTOR, Jacqueline N. Crawley,1,2,* of de novo variants in ASD and cancer WNT, NOTCH, and MAPK are compo- Wolf-Dietrich Heyer,3,4 and have begun to uncover considerable addi- nents of signaling pathways regulating Janine M. LaSalle1,4,5 tional overlap. What is surprising about the the nuclear factors described above. genes in Table 1 is not necessarily the Autism is a neurodevelopmental number of risk genes found in both autism Autism is comorbid with several mono- and cancer, but the shared functions of genic neurodevelopmental disorders, disorder, diagnosed behaviorally genes in chromatin remodeling and including Fragile X (FMR1), Rett syndrome by social and communication genome maintenance, transcription fac- (MECP2), Phelan-McDermid (SHANK3), fi de cits, repetitive behaviors, tors, and signal transduction pathways 15q duplication syndrome (UBE3A), and restricted interests. Recent leading to nuclear changes [7,8]. -

HHS Public Access Author Manuscript
HHS Public Access Author manuscript Author Manuscript Author ManuscriptOncogene Author Manuscript. Author manuscript; Author Manuscript available in PMC 2015 July 02. Published in final edited form as: Oncogene. 2015 January 2; 34(1): 94–103. doi:10.1038/onc.2013.523. Upregulation of the microRNA cluster at the Dlk1-Dio3 locus in lung adenocarcinoma Paul N. Valdmanis1,2, Biswajoy Roy-Chaudhuri1,2, Hak Kyun Kim1,2, Leanne C. Sayles3, Yanyan Zheng3, Chen-Hua Chuang2,4,5, Deborah R. Caswell2,4,5, Kirk Chu1,2, Monte M. Winslow2,4,5, E. Alejandro Sweet-Cordero1,3,5, and Mark A. Kay1,2 1Department of Pediatrics, Stanford University, Stanford, CA, USA 2Department of Genetics, Stanford University, Stanford, CA, USA 3Division of Hematology/Oncology, Department of Pediatrics, Stanford University, Stanford, CA, USA 4Cancer Biology Program, Stanford University, Stanford, CA, USA 5Stanford Cancer Institute, Stanford University, Stanford, CA, USA Abstract Mice in which lung epithelial cells can be induced to express an oncogenic KrasG12D develop lung adenocarcinomas in a manner analogous to humans. A myriad of genetic changes accompany lung adenocarcinomas, many of which are poorly understood. To get a comprehensive understanding of both the transcriptional and post-transcriptional changes that accompany lung adenocarcinomas, we took an omics approach in profiling both the coding genes and the non- coding small RNAs in an induced mouse model of lung adenocarcinoma. RNAseq transcriptome analysis of KrasG12D tumors from F1 hybrid mice revealed features specific to tumor samples. This includes the repression of a network of GTPase related genes (Prkg1, Gnao1 and Rgs9) in tumor samples and an enrichment of Apobec1-mediated cytosine to uridine RNA editing. -

Molecular Architecture Underlying Fluid Absorption by the Developing Inner
RESEARCH ARTICLE Molecular architecture underlying fluid absorption by the developing inner ear Keiji Honda1†, Sung Huhn Kim2‡, Michael C Kelly3, Joseph C Burns3§, Laura Constance2, Xiangming Li2#, Fei Zhou2, Michael Hoa4, Matthew W Kelley3, Philine Wangemann2*, Robert J Morell5, Andrew J Griffith1* 1Molecular Biology and Genetics Section, National Institute on Deafness and Other Communication Disorders, National Institutes of Health, Bethesda, United States; 2Anatomy and Physiology Department, Kansas State University, Manhattan, United States; 3Developmental Neuroscience Section, National Institute on Deafness and Other Communication Disorders, National Institutes of Health, Bethesda, United States; 4Auditory Development and Restoration Program, National Institute on *For correspondence: Deafness and Other Communication Disorders, National Institutes of Health, [email protected] (PW); Bethesda, United States; 5Genomics and Computational Biology Core, National [email protected] (AJG) Institute on Deafness and Other Communication Disorders, National Institutes of Present address: Health, Bethesda, United States †Otolaryngology Department, Tsuchiura Kyodo General Hospital, Tsuchiura, Japan; ‡Department of Abstract Mutations of SLC26A4 are a common cause of hearing loss associated with Otorhinolaryngology, Head and enlargement of the endolymphatic sac (EES). Slc26a4 expression in the developing mouse Neck Surgery, Yonsei University College of Medicine, Seoul, endolymphatic sac is required for acquisition of normal inner ear structure and function. Here, we Korea; §Decibel Therapeutics, show that the mouse endolymphatic sac absorbs fluid in an SLC26A4-dependent fashion. Fluid Cambridge, United States; absorption was sensitive to ouabain and gadolinium but insensitive to benzamil, bafilomycin and #Technique R and D-Drug S3226. Single-cell RNA-seq analysis of pre- and postnatal endolymphatic sacs demonstrates two Substance, GlaxoSmithKline types of differentiated cells. -
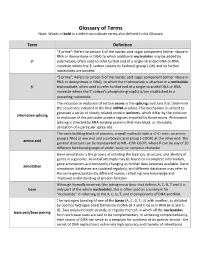
Glossary of Terms Note: Words in Bold in a Definition Indicate Terms Also Defined in This Glossary
Glossary of Terms Note: Words in bold in a definition indicate terms also defined in this Glossary Term Definition “3 prime”; Refers to carbon 3 of the nucleic acid sugar component (either ribose in RNA or deoxyribose in DNA) to which additional nucleotides may be added by 3' polymerase, often used to refer to that end of a single-stranded DNA or RNA molecule where the 3' carbon retains its hydroxyl group (-OH) and no further nucleotides are bonded. “5 prime”; Refers to carbon 5 of the nucleic acid sugar component (either ribose in RNA or deoxyribose in DNA), to which the triphosphate is attached in a nucleotide 5' triphosphate, often used to refer to that end of a single-stranded DNA or RNA molecule where the 5' carbon's phosphate group(s) is/are unattached to a preceding nucleotide. The inclusion or exclusion of certain exons in the splicing reactions that determine the sequences included in the final mRNA product. This mechanism is utilized to generate a series of closely related protein isoforms, which differ by the inclusion alternative splicing or exclusion of the particular protein regions encoded by those exons. Alternative splicing is directed by RNA-binding proteins that may block, or stimulate, utilization of a particular splice site. The basic building block of proteins, a small molecule with a -C-C- core, an amine group (-NH2) at one end and a carboxylic acid group (-COOH) at the other end. The amino acid general structure can be represented as NH2-CHR-COOH, where R can be any of 20 different functional groups of acidic, basic, or nonpolar character. -

Protein-Synthesis.Pdf
Protein Synthesis Contents Introduction ........................................................................................................................ 1 DNA Transcription ............................................................................................................... 1 The Primary Transcript ........................................................................................................ 4 Translation .......................................................................................................................... 7 Protein Structure............................................................................................................... 11 References ........................................................................................................................ 11 Resources .......................................................................................................................... 11 Introduction The genetic message carried on the DNA molecule is a code. This code is ultimately translated into a sequence of amino acids that, when complete, becomes a protein. Proteins carry out the “business” of the cell. Some proteins are used as structural components of cells, some are used to transport other molecules, still others are charged with directing chemical reactions. The latter class of proteins is the enzymes. Regardless of the role played by a protein in the cell one aspect is the same, they are all encoded in the base sequences of DNA. The path from DNA sequence to protein