The Tissue-Specific RNA Binding Protein T-STAR Controls Regional
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Atlas Antibodies in Breast Cancer Research Table of Contents
ATLAS ANTIBODIES IN BREAST CANCER RESEARCH TABLE OF CONTENTS The Human Protein Atlas, Triple A Polyclonals and PrecisA Monoclonals (4-5) Clinical markers (6) Antibodies used in breast cancer research (7-13) Antibodies against MammaPrint and other gene expression test proteins (14-16) Antibodies identified in the Human Protein Atlas (17-14) Finding cancer biomarkers, as exemplified by RBM3, granulin and anillin (19-22) Co-Development program (23) Contact (24) Page 2 (24) Page 3 (24) The Human Protein Atlas: a map of the Human Proteome The Human Protein Atlas (HPA) is a The Human Protein Atlas consortium cell types. All the IHC images for Swedish-based program initiated in is mainly funded by the Knut and Alice the normal tissue have undergone 2003 with the aim to map all the human Wallenberg Foundation. pathology-based annotation of proteins in cells, tissues and organs expression levels. using integration of various omics The Human Protein Atlas consists of technologies, including antibody- six separate parts, each focusing on References based imaging, mass spectrometry- a particular aspect of the genome- 1. Sjöstedt E, et al. (2020) An atlas of the based proteomics, transcriptomics wide analysis of the human proteins: protein-coding genes in the human, pig, and and systems biology. mouse brain. Science 367(6482) 2. Thul PJ, et al. (2017) A subcellular map of • The Tissue Atlas shows the the human proteome. Science. 356(6340): All the data in the knowledge resource distribution of proteins across all eaal3321 is open access to allow scientists both major tissues and organs in the 3. -

Genetic Targeting of NRXN2 in Mice Unveils Role in Excitatory Cortical Synapse Function and Social Behaviors
King’s Research Portal DOI: 10.3389/fnsyn.2015.00003 Document Version Publisher's PDF, also known as Version of record Link to publication record in King's Research Portal Citation for published version (APA): Born, G., Grayton, H. M., Langhorst, H., Dudanova, I., Rohlmann, A., Woodward, B. W., Collier, D. A., Fernandes, C., & Missler, M. (2015). Genetic targeting of NRXN2 in mice unveils role in excitatory cortical synapse function and social behaviors. Frontiers in Synaptic Neuroscience, 7(0), [3]. https://doi.org/10.3389/fnsyn.2015.00003 Citing this paper Please note that where the full-text provided on King's Research Portal is the Author Accepted Manuscript or Post-Print version this may differ from the final Published version. If citing, it is advised that you check and use the publisher's definitive version for pagination, volume/issue, and date of publication details. And where the final published version is provided on the Research Portal, if citing you are again advised to check the publisher's website for any subsequent corrections. General rights Copyright and moral rights for the publications made accessible in the Research Portal are retained by the authors and/or other copyright owners and it is a condition of accessing publications that users recognize and abide by the legal requirements associated with these rights. •Users may download and print one copy of any publication from the Research Portal for the purpose of private study or research. •You may not further distribute the material or use it for any profit-making activity or commercial gain •You may freely distribute the URL identifying the publication in the Research Portal Take down policy If you believe that this document breaches copyright please contact [email protected] providing details, and we will remove access to the work immediately and investigate your claim. -

P.1.H.029. Abnormality of Social Behavior and Dysfunction of Autism
P.1.h.029. Abnormality of social behavior and dysfunction of autism related gene expression developing under chronic social defeat stress in male mice Kudryavtseva N.N., Kovalenko I.L., Smagin D.A., Galyamina A.G., Babenko V.N. Federal Research Center, Institute of Cytology and Genetics of SB RAS, Novosibirsk, Russia The ability of people to communicate with each other is a necessary component of social behavior and the normal development of individuals living in a community. An apparent decline in sociability may be the result of a negative social environment or the development of neurological disorders, including autistic spectrum disorders. The behavior of these humans may be characterized by the impairment of socialization, low communication, restricted and repetitive behaviors. This study aimed to analyze changes in the social behaviors induced by daily social defeat stress in male mice and to study the involvement of autism associated genes in the process of the deterioration of social behaviors. sensory contact agonistic interactions aggressive defeated mice Methods.The sensory contact model [1] was mice used to generate repeated experience of social defeats in male mice of C57BL/6J strain. After three days of sensory contact the partition is removed for 10 min to allow agonistic interactions between males. Undoubted superiority of one of the partners is evident The collected brain samples were sequenced at JSC Genoanalytica (www.genoanalytica.ru, within 2-3 daily social encounters with the same Moscow, Russia), where the mRNA was extracted using the Dynabeads mRNA Purification Kit (Ambion, USA). cDNA libraries were constructed using NEBNext mRNA Library PrepReagent Set opponent. -

Neurexin II Alpha (NRXN2) (NM 015080) Human Tagged ORF Clone Product Data
OriGene Technologies, Inc. 9620 Medical Center Drive, Ste 200 Rockville, MD 20850, US Phone: +1-888-267-4436 [email protected] EU: [email protected] CN: [email protected] Product datasheet for RC219788 Neurexin II alpha (NRXN2) (NM_015080) Human Tagged ORF Clone Product data: Product Type: Expression Plasmids Product Name: Neurexin II alpha (NRXN2) (NM_015080) Human Tagged ORF Clone Tag: Myc-DDK Symbol: NRXN2 Vector: pCMV6-Entry (PS100001) E. coli Selection: Kanamycin (25 ug/mL) Cell Selection: Neomycin ORF Nucleotide >RC219788 representing NM_015080 Sequence: Red=Cloning site Blue=ORF Green=Tags(s) TTTTGTAATACGACTCACTATAGGGCGGCCGGGAATTCGTCGACTGGATCCGGTACCGAGGAGATCTGCC GCCGCGATCGCC ATGGCGTCCGGGAGCCGGTGGCGGCCGACACCGCCGCCGCTGCTGTTGCTGCTGCTGCTGGCGCTGGCGG CGCGCGCGGACGGCCTGGAGTTCGGCGGCGGCCCCGGGCAGTGGGCTCGCTACGCGCGCTGGGCGGGCGC GGCGAGCAGCGGCGAGCTCAGCTTCAGCCTGCGCACCAACGCCACGCGCGCGCTGCTGCTCTACCTGGAC GACGGCGGCGACTGCGACTTCCTGGAGCTGCTGCTGGTGGACGGCCGCCTGCGGCTGCGCTTCACGCTTT CGTGCGCCGAGCCGGCCACGCTGCAGCTGGACACGCCGGTGGCCGACGACCGCTGGCACATGGTGCTGCT GACCCGCGACGCGCGCCGCACGGCGCTGGCGGTGGACGGCGAGGCCCGCGCCGCCGAGGTGCGCTCCAAG CGGCGCGAGATGCAGGTGGCCAGCGACCTGTTCGTGGGCGGCATCCCGCCCGACGTGCGCCTCTCGGCGC TTACGCTGAGCACCGTCAAGTACGAGCCGCCCTTCCGCGGCCTCTTGGCCAACCTGAAGCTGGGCGAGCG GCCCCCCGCGCTGCTGGGCAGCCAGGGCCTGCGCGGCGCCACCGCCGACCCGCTGTGCGCGCCCGCGCGC AACCCCTGCGCCAACGGCGGCCTCTGCACCGTGCTGGCCCCCGGCGAGGTGGGCTGCGACTGCAGCCACA CGGGCTTCGGCGGCAAGTTCTGCAGCGAAGAGGAGCACCCCATGGAAGGTCCGGCTCACCTGACGTTAAA CAGCGAAGTAGGGTCCTTACTGTTCTCCGAGGGGGGGGCCGGGAGAGGAGGAGCCGGCGATGTGCACCAG CCAACAAAAGGCAAGGAGGAGTTTGTGGCGACCTTCAAAGGCAATGAGTTCTTCTGCTACGACCTGTCAC -
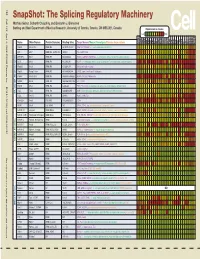
Snapshot: the Splicing Regulatory Machinery Mathieu Gabut, Sidharth Chaudhry, and Benjamin J
192 Cell SnapShot: The Splicing Regulatory Machinery Mathieu Gabut, Sidharth Chaudhry, and Benjamin J. Blencowe 133 Banting and Best Department of Medical Research, University of Toronto, Toronto, ON M5S 3E1, Canada Expression in mouse , April4, 2008©2008Elsevier Inc. Low High Name Other Names Protein Domains Binding Sites Target Genes/Mouse Phenotypes/Disease Associations Amy Ceb Hip Hyp OB Eye SC BM Bo Ht SM Epd Kd Liv Lu Pan Pla Pro Sto Spl Thy Thd Te Ut Ov E6.5 E8.5 E10.5 SRp20 Sfrs3, X16 RRM, RS GCUCCUCUUC SRp20, CT/CGRP; −/− early embryonic lethal E3.5 9G8 Sfrs7 RRM, RS, C2HC Znf (GAC)n Tau, GnRH, 9G8 ASF/SF2 Sfrs1 RRM, RS RGAAGAAC HipK3, CaMKIIδ, HIV RNAs; −/− embryonic lethal, cond. KO cardiomyopathy SC35 Sfrs2 RRM, RS UGCUGUU AChE; −/− embryonic lethal, cond. KO deficient T-cell maturation, cardiomyopathy; LS SRp30c Sfrs9 RRM, RS CUGGAUU Glucocorticoid receptor SRp38 Fusip1, Nssr RRM, RS ACAAAGACAA CREB, type II and type XI collagens SRp40 Sfrs5, HRS RRM, RS AGGAGAAGGGA HipK3, PKCβ-II, Fibronectin SRp55 Sfrs6 RRM, RS GGCAGCACCUG cTnT, CD44 DOI 10.1016/j.cell.2008.03.010 SRp75 Sfrs4 RRM, RS GAAGGA FN1, E1A, CD45; overexpression enhances chondrogenic differentiation Tra2α Tra2a RRM, RS GAAARGARR GnRH; overexpression promotes RA-induced neural differentiation SR and SR-Related Proteins Tra2β Sfrs10 RRM, RS (GAA)n HipK3, SMN, Tau SRm160 Srrm1 RS, PWI AUGAAGAGGA CD44 SWAP Sfrs8 RS, SWAP ND SWAP, CD45, Tau; possible asthma susceptibility gene hnRNP A1 Hnrnpa1 RRM, RGG UAGGGA/U HipK3, SMN2, c-H-ras; rheumatoid arthritis, systemic lupus -

Deletion of Α-Neurexin II Results in Autism-Related
OPEN Citation: Transl Psychiatry (2014) 4, e484; doi:10.1038/tp.2014.123 www.nature.com/tp ORIGINAL ARTICLE Deletion of α-neurexin II results in autism-related behaviors in mice J Dachtler1, J Glasper2, RN Cohen1, JL Ivorra1, DJ Swiffen1, AJ Jackson1, MK Harte2, RJ Rodgers3 and SJ Clapcote1 Autism is a common and frequently disabling neurodevelopmental disorder with a strong genetic basis. Human genetic studies have discovered mutations disrupting exons of the NRXN2 gene, which encodes the synaptic adhesion protein α-neurexin II (Nrxn2α), in two unrelated individuals with autism, but a causal link between NRXN2 and the disorder remains unclear. To begin to test the hypothesis that Nrxn2α deficiency contributes to the symptoms of autism, we employed Nrxn2α knockout (KO) mice that genetically model Nrxn2α deficiency in vivo. We report that Nrxn2α KO mice displayed deficits in sociability and social memory when exposed to novel conspecifics. In tests of exploratory activity, Nrxn2α KO mice displayed an anxiety-like phenotype in comparison with wild-type littermates, with thigmotaxis in an open field, less time spent in the open arms of an elevated plus maze, more time spent in the enclosure of an emergence test and less time spent exploring novel objects. However, Nrxn2α KO mice did not exhibit any obvious changes in prepulse inhibition or in passive avoidance learning. Real-time PCR analysis of the frontal cortex and hippocampus revealed significant decreases in the mRNA levels of genes encoding proteins involved in both excitatory and inhibitory transmission. Quantification of protein expression revealed that Munc18-1, encoded by Stxbp1, was significantly decreased in the hippocampus of Nrxn2α KO mice, which is suggestive of deficiencies in presynaptic vesicular release. -

Structures of Neurexophilin–Neurexin Complexes Reveal a Regulatory Mechanism of Alternative Splicing
Article Structures of neurexophilin–neurexin complexes reveal a regulatory mechanism of alternative splicing Steven C Wilson1 , K Ian White1 , Qiangjun Zhou1, Richard A Pfuetzner1, Ucheor B Choi1, Thomas C Südhof1,2 & Axel T Brunger1,2,* Abstract Neurexins (Nrxns) are a family of presynaptic cell-adhesion mole- cules that are important for specifying the functional properties of Neurexins are presynaptic, cell-adhesion molecules that specify synapses (Su¨dhof, 2017). Through their interactions with various the functional properties of synapses via interactions with trans- binding partners, neurexins constitute part of a molecular code and synaptic ligands. Neurexins are extensively alternatively spliced at act as hub molecules to organize and regulate synaptic function six canonical sites that regulate multifarious ligand interactions, (Su¨dhof, 2017). but the structural mechanisms underlying alternative splicing- Neurexins are encoded by three genes (Nrxn1–3 in mice), each dependent neurexin regulation are largely unknown. Here, we of which contains multiple promoters to drive expression of determined high-resolution structures of the complex of neurex- Nrxn1a–3a, Nrxn1b–3b, and Nrxn1c. The extracellular sequences of ophilin-1 and the second laminin/neurexin/sex-hormone-binding a-neurexins contain six LNS domains (LNS1–LNS6) interspersed globulin domain (LNS2) of neurexin-1 and examined how alterna- with epidermal growth factor-like domains (EGFA–EGFC), followed tive splicing at splice site #2 (SS2) regulates the complex. Our data by a cysteine loop and a highly glycosylated stalk region (Fig 1A). reveal a unique, extensive, neurexophilin–neurexin binding inter- The b-neurexins include only the C-terminal region of a-neurexins face that extends the jelly-roll b-sandwich of LNS2 of neurexin-1 from the LNS6 domain onward. -

Logistic Regression Analysis for the Identification of the Metastasis-Associated Signaling Pathways of Osteosarcoma
INTERNATIONAL JOURNAL OF MOLECULAR MEDICINE 41: 1233-1244, 2018 Logistic regression analysis for the identification of the metastasis-associated signaling pathways of osteosarcoma YANG LIU1, WEI SUN2, XIAOJUN MA2, YUEDONG HAO3, GANG LIU3, XIAOHUI HU3, HOULAI SHANG3, PENGFEI WU3, ZEXUE ZHAO3 and WEIDONG LIU3 1Department of Orthopedics, Affiliated Hospital of Inner Mongolia University for The Nationalities, Tongliao, Inner Mongolia 028007; 2Department of Orthopaedics, Shanghai General Hospital, School of Medicine, Shanghai Jiao Tong University, Shanghai 200080; 3Department of Orthopedics, Huai'an First People's Hospital, Nanjing Medical University, Huai'an, Jiangsu 223300, P.R. China Received February 20, 2017; Accepted September 26, 2017 DOI: 10.3892/ijmm.2018.3360 Abstract. Osteosarcoma (OS) is the most common histological migration were identified as the terms that were significantly type of primary bone cancer. The present study was designed different. ICAM1, PDGFB, PDGFRB and TGFB1 were iden- to identify the key genes and signaling pathways involved in the tified to be enriched in cell migration and regulation of cell metastasis of OS. Microarray data of GSE39055 were down- migration. Nucleotide excision repair and the Wnt signaling loaded from the Gene Expression Omnibus database, which pathway were the metastasis-associated pathways of OS. In included 19 OS biopsy specimens before metastasis (control addition, ICAM1, PDGFB, PDGFRB and TGFB1, which were group) and 18 OS biopsy specimens after metastasis (case involved in cell migration and regulation of cell migration may group). After the differentially expressed genes (DEGs) were affect the metastasis of OS. identified using the Linear Models for Microarray Analysis package, hierarchical clustering analysis and unsupervised Introduction clustering analysis were performed separately, using orange software and the self-organization map method. -

Autism-Associated Mir-873 Regulates ARID1B, SHANK3 and NRXN2
Lu et al. Translational Psychiatry (2020) 10:418 https://doi.org/10.1038/s41398-020-01106-8 Translational Psychiatry ARTICLE Open Access Autism-associated miR-873 regulates ARID1B, SHANK3 and NRXN2 involved in neurodevelopment Jing Lu 1, Yan Zhu 1, Sarah Williams2, Michelle Watts 3,MaryA.Tonta1, Harold A. Coleman1, Helena C. Parkington1 and Charles Claudianos 1,4 Abstract Autism spectrum disorders (ASD) are highly heritable neurodevelopmental disorders with significant genetic heterogeneity. Noncoding microRNAs (miRNAs) are recognised as playing key roles in development of ASD albeit the function of these regulatory genes remains unclear. We previously conducted whole-exome sequencing of Australian families with ASD and identified four novel single nucleotide variations in mature miRNA sequences. A pull-down transcriptome analysis using transfected SH-SY5Y cells proposed a mechanistic model to examine changes in binding affinity associated with a unique mutation found in the conserved ‘seed’ region of miR-873-5p (rs777143952: T > A). Results suggested several ASD-risk genes were differentially targeted by wild-type and mutant miR-873 variants. In the current study, a dual-luciferase reporter assay confirmed miR-873 variants have a 20-30% inhibition/dysregulation effect on candidate autism risk genes ARID1B, SHANK3 and NRXN2 and also confirmed the affected expression with qPCR. In vitro mouse hippocampal neurons transfected with mutant miR-873 showed less morphological complexity and enhanced sodium currents and excitatory neurotransmission compared to cells transfected with wild-type miR- 873. A second in vitro study showed CRISPR/Cas9 miR-873 disrupted SH-SY5Y neuroblastoma cells acquired a neuronal-like morphology and increased expression of ASD important genes ARID1B, SHANK3, ADNP2, ANK2 and CHD8. -

Analysis of Alternative Splicing Associated with Aging and Neurodegeneration in the Human Brain
Downloaded from genome.cshlp.org on October 2, 2021 - Published by Cold Spring Harbor Laboratory Press Research Analysis of alternative splicing associated with aging and neurodegeneration in the human brain James R. Tollervey,1 Zhen Wang,1 Tibor Hortoba´gyi,2 Joshua T. Witten,1 Kathi Zarnack,3 Melis Kayikci,1 Tyson A. Clark,4 Anthony C. Schweitzer,4 Gregor Rot,5 Tomazˇ Curk,5 Blazˇ Zupan,5 Boris Rogelj,2 Christopher E. Shaw,2 and Jernej Ule1,6 1MRC Laboratory of Molecular Biology, Hills Road, Cambridge CB2 0QH, United Kingdom; 2MRC Centre for Neurodegeneration Research, King’s College London, Institute of Psychiatry, De Crespigny Park, London SE5 8AF, United Kingdom; 3EMBL–European Bioinformatics Institute, Wellcome Trust Genome Campus, Hinxton, Cambridge CB10 1SD, United Kingdom; 4Expression Research, Affymetrix, Inc., Santa Clara, California 95051, USA; 5Faculty of Computer and Information Science, University of Ljubljana, Trzˇasˇka 25, SI-1000 Ljubljana, Slovenia Age is the most important risk factor for neurodegeneration; however, the effects of aging and neurodegeneration on gene expression in the human brain have most often been studied separately. Here, we analyzed changes in transcript levels and alternative splicing in the temporal cortex of individuals of different ages who were cognitively normal, affected by frontotemporal lobar degeneration (FTLD), or affected by Alzheimer’s disease (AD). We identified age-related splicing changes in cognitively normal individuals and found that these were present also in 95% of individuals with FTLD or AD, independent of their age. These changes were consistent with increased polypyrimidine tract binding protein (PTB)– dependent splicing activity. We also identified disease-specific splicing changes that were present in individuals with FTLD or AD, but not in cognitively normal individuals. -

Dynamic Changes in RNA–Protein Interactions and RNA Secondary Structure in Mammalian Erythropoiesis
Published Online: 27 July, 2021 | Supp Info: http://doi.org/10.26508/lsa.202000659 Downloaded from life-science-alliance.org on 30 September, 2021 Resource Dynamic changes in RNA–protein interactions and RNA secondary structure in mammalian erythropoiesis Mengge Shan1,2 , Xinjun Ji3, Kevin Janssen5 , Ian M Silverman3 , Jesse Humenik3, Ben A Garcia5, Stephen A Liebhaber3,4, Brian D Gregory1,2 Two features of eukaryotic RNA molecules that regulate their and RNA secondary structure. These techniques generally either post-transcriptional fates are RNA secondary structure and RNA- use chemical probing agents or structure-specific RNases (single- binding protein (RBP) interaction sites. However, a comprehen- stranded RNases (ssRNases) and double-stranded RNases [dsRNa- sive global overview of the dynamic nature of these sequence ses]) to provide site-specific evidence for a region being in single- or features during erythropoiesis has never been obtained. Here, we double-stranded configurations (4, 5). use our ribonuclease-mediated structure and RBP-binding site To date, the known repertoire of RBP–RNA interaction sites has mapping approach to reveal the global landscape of RNA sec- been built on a protein-by-protein basis, with studies identifying ondary structure and RBP–RNA interaction sites and the dynamics the targets of a single protein of interest, often through the use of of these features during this important developmental process. techniques such as crosslinking and immunoprecipitation se- We identify dynamic patterns of RNA secondary structure and RBP quencing (CLIP-seq) (6). In CLIP-seq, samples are irradiated with UV binding throughout the process and determine a set of corre- to induce the cross-linking of proteins to their RNA targets. -

Deciphering the Molecular Profile of Plaques, Memory Decline And
ORIGINAL RESEARCH ARTICLE published: 16 April 2014 AGING NEUROSCIENCE doi: 10.3389/fnagi.2014.00075 Deciphering the molecular profile of plaques, memory decline and neuron loss in two mouse models for Alzheimer’s disease by deep sequencing Yvonne Bouter 1†,Tim Kacprowski 2,3†, Robert Weissmann4, Katharina Dietrich1, Henning Borgers 1, Andreas Brauß1, Christian Sperling 4, Oliver Wirths 1, Mario Albrecht 2,5, Lars R. Jensen4, Andreas W. Kuss 4* andThomas A. Bayer 1* 1 Division of Molecular Psychiatry, Georg-August-University Goettingen, University Medicine Goettingen, Goettingen, Germany 2 Department of Bioinformatics, Institute of Biometrics and Medical Informatics, University Medicine Greifswald, Greifswald, Germany 3 Department of Functional Genomics, Interfaculty Institute for Genetics and Functional Genomics, University Medicine Greifswald, Greifswald, Germany 4 Human Molecular Genetics, Department for Human Genetics of the Institute for Genetics and Functional Genomics, Institute for Human Genetics, University Medicine Greifswald, Ernst-Moritz-Arndt University Greifswald, Greifswald, Germany 5 Institute for Knowledge Discovery, Graz University of Technology, Graz, Austria Edited by: One of the central research questions on the etiology of Alzheimer’s disease (AD) is the Isidro Ferrer, University of Barcelona, elucidation of the molecular signatures triggered by the amyloid cascade of pathological Spain events. Next-generation sequencing allows the identification of genes involved in disease Reviewed by: Isidro Ferrer, University of Barcelona, processes in an unbiased manner. We have combined this technique with the analysis of Spain two AD mouse models: (1) The 5XFAD model develops early plaque formation, intraneu- Dietmar R. Thal, University of Ulm, ronal Ab aggregation, neuron loss, and behavioral deficits. (2)TheTg4–42 model expresses Germany N-truncated Ab4–42 and develops neuron loss and behavioral deficits albeit without plaque *Correspondence: formation.