Table 7. Genes
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Gene Symbol Gene Description ACVR1B Activin a Receptor, Type IB
Table S1. Kinase clones included in human kinase cDNA library for yeast two-hybrid screening Gene Symbol Gene Description ACVR1B activin A receptor, type IB ADCK2 aarF domain containing kinase 2 ADCK4 aarF domain containing kinase 4 AGK multiple substrate lipid kinase;MULK AK1 adenylate kinase 1 AK3 adenylate kinase 3 like 1 AK3L1 adenylate kinase 3 ALDH18A1 aldehyde dehydrogenase 18 family, member A1;ALDH18A1 ALK anaplastic lymphoma kinase (Ki-1) ALPK1 alpha-kinase 1 ALPK2 alpha-kinase 2 AMHR2 anti-Mullerian hormone receptor, type II ARAF v-raf murine sarcoma 3611 viral oncogene homolog 1 ARSG arylsulfatase G;ARSG AURKB aurora kinase B AURKC aurora kinase C BCKDK branched chain alpha-ketoacid dehydrogenase kinase BMPR1A bone morphogenetic protein receptor, type IA BMPR2 bone morphogenetic protein receptor, type II (serine/threonine kinase) BRAF v-raf murine sarcoma viral oncogene homolog B1 BRD3 bromodomain containing 3 BRD4 bromodomain containing 4 BTK Bruton agammaglobulinemia tyrosine kinase BUB1 BUB1 budding uninhibited by benzimidazoles 1 homolog (yeast) BUB1B BUB1 budding uninhibited by benzimidazoles 1 homolog beta (yeast) C9orf98 chromosome 9 open reading frame 98;C9orf98 CABC1 chaperone, ABC1 activity of bc1 complex like (S. pombe) CALM1 calmodulin 1 (phosphorylase kinase, delta) CALM2 calmodulin 2 (phosphorylase kinase, delta) CALM3 calmodulin 3 (phosphorylase kinase, delta) CAMK1 calcium/calmodulin-dependent protein kinase I CAMK2A calcium/calmodulin-dependent protein kinase (CaM kinase) II alpha CAMK2B calcium/calmodulin-dependent -

Association Analyses of Known Genetic Variants with Gene
ASSOCIATION ANALYSES OF KNOWN GENETIC VARIANTS WITH GENE EXPRESSION IN BRAIN by Viktoriya Strumba A dissertation submitted in partial fulfillment of the requirements for the degree of Doctor of Philosophy (Bioinformatics) in The University of Michigan 2009 Doctoral Committee: Professor Margit Burmeister, Chair Professor Huda Akil Professor Brian D. Athey Assistant Professor Zhaohui S. Qin Research Statistician Thomas Blackwell To Sam and Valentina Dmitriy and Elizabeth ii ACKNOWLEDGEMENTS I would like to thank my advisor Professor Margit Burmeister, who tirelessly guided me though seemingly impassable corridors of graduate work. Throughout my thesis writing period she provided sound advice, encouragement and inspiration. Leading by example, her enthusiasm and dedication have been instrumental in my path to becoming a better scientist. I also would like to thank my co-advisor Tom Blackwell. His careful prodding always kept me on my toes and looking for answers, which taught me the depth of careful statistical analysis. His diligence and dedication have been irreplaceable in most difficult of projects. I also would like to thank my other committee members: Huda Akil, Brian Athey and Steve Qin as well as David States. You did not make it easy for me, but I thank you for believing and not giving up. Huda’s eloquence in every subject matter she explained have been particularly inspiring, while both Huda’s and Brian’s valuable advice made the completion of this dissertation possible. I would also like to thank all the members of the Burmeister lab, both past and present: Sandra Villafuerte, Kristine Ito, Cindy Schoen, Karen Majczenko, Ellen Schmidt, Randi Burns, Gang Su, Nan Xiang and Ana Progovac. -

Nucleocytoplasmic Shuttling of the Glucocorticoid Receptor Is Influenced by Tetratricopeptide Repeat-Containing Proteins Gisela I
© 2020. Published by The Company of Biologists Ltd | Journal of Cell Science (2020) 133, jcs238873. doi:10.1242/jcs.238873 RESEARCH ARTICLE Nucleocytoplasmic shuttling of the glucocorticoid receptor is influenced by tetratricopeptide repeat-containing proteins Gisela I. Mazaira1,*, Pablo C. Echeverria2,* and Mario D. Galigniana1,3,‡ ABSTRACT FKBP52 (also known as FKBP4), FKBP51 (FKBP5) and CyP40 It has been demonstrated that tetratricopeptide-repeat (TPR) domain (PPID), and the immunophilin-like proteins PP5, FKBPL (WISp39) proteins regulate the subcellular localization of glucocorticoid receptor and FKBP37 (ARA9/XAP2/AIP) are counted among the best (GR). This study analyses the influence of the TPR domain of high characterized members of the TPR domain family of proteins molecular weight immunophilins in the retrograde transport and associated with nuclear receptor family members via Hsp90 (Pratt nuclear retention of GR. Overexpression of the TPR peptide et al., 2004a; Storer et al., 2011). TPR domains are composed of prevented efficient nuclear accumulation of the GR by disrupting the loosely conserved 34 amino acid sequence motifs that are found in formation of complexes with the dynein-associated immunophilin arrays comprising between 2 and 20 repeats, wherein the repeats pack FKBP52 (also known as FKBP4), the adaptor transporter importin-β1 against each other giving rise to coiled and superhelical structures (KPNB1), the nuclear pore-associated glycoprotein Nup62 and nuclear (Perez-Riba and Itzhaki, 2019). Originally identified in components of matrix-associated structures. We also show that nuclear import of GR the anaphase-promoting complex (Sikorski et al., 1991), TPR was impaired, whereas GR nuclear export was enhanced. domains are now known to mediate specific protein interactions in Interestingly, the CRM1 (exportin-1) inhibitor leptomycin-B abolished numerous cellular contexts. -
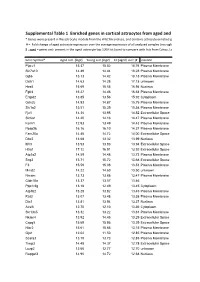
Supplemental Table 1 Enriched Genes in Cortical Astrocytes from Aged
Supplemental Table 1 Enriched genes in cortical astrocytes from aged and young-adult mice * Genes were present in the astrocyte module from the WGCNA analysis, and contains astrocyte enriched genes compared to microglia and oligodendrocytes # = Fold change of aged astrocyte expression over the average expression of all analyzed samples (microglia, astrocytes: young, old, with and without myelin contamination) $ ; aged = genes only present in the aged astrocyte top 1000 list (used to compare with lists from Cahoy, Lovatt, Doyle; see Fig. 4B), all = genes present in all astrocyte top 1000 lists Gene Symbol* Aged astr. (log2) Young astr.(log2) FC (aged/ aver.)# Location Ptprz1 15.37 15.02 18.76 Plasma Membrane Slc7a10 14.49 14.44 18.28 Plasma Membrane Gjb6 15.13 14.42 18.18 Plasma Membrane Dclk1 14.63 14.28 17.18 unknown Hes5 15.69 15.55 16.94 Nucleus Fgfr3 15.27 14.46 16.54 Plasma Membrane Entpd2 13.85 13.56 15.92 Cytoplasm Grin2c 14.93 14.87 15.75 Plasma Membrane Slc1a2 15.51 15.39 15.58 Plasma Membrane Fjx1 14.36 13.98 14.52 Extracellular Space Slc6a1 14.20 14.16 14.47 Plasma Membrane Kcnk1 12.93 13.49 14.43 Plasma Membrane Ppap2b 16.16 16.10 14.37 Plasma Membrane Fam20a 14.48 14.72 14.00 Extracellular Space Dbx2 13.68 13.32 13.99 Nucleus Itih3 13.93 13.93 13.94 Extracellular Space Htra1 17.12 16.91 13.92 Extracellular Space Atp1a2 14.59 14.48 13.73 Plasma Membrane Scg3 15.71 15.72 13.68 Extracellular Space F3 15.59 15.08 13.51 Plasma Membrane Mmd2 14.22 14.60 13.50 unknown Nrcam 13.73 13.88 13.47 Plasma Membrane Cldn10a 13.37 13.57 13.46 -

Table 2. Significant
Table 2. Significant (Q < 0.05 and |d | > 0.5) transcripts from the meta-analysis Gene Chr Mb Gene Name Affy ProbeSet cDNA_IDs d HAP/LAP d HAP/LAP d d IS Average d Ztest P values Q-value Symbol ID (study #5) 1 2 STS B2m 2 122 beta-2 microglobulin 1452428_a_at AI848245 1.75334941 4 3.2 4 3.2316485 1.07398E-09 5.69E-08 Man2b1 8 84.4 mannosidase 2, alpha B1 1416340_a_at H4049B01 3.75722111 3.87309653 2.1 1.6 2.84852656 5.32443E-07 1.58E-05 1110032A03Rik 9 50.9 RIKEN cDNA 1110032A03 gene 1417211_a_at H4035E05 4 1.66015788 4 1.7 2.82772795 2.94266E-05 0.000527 NA 9 48.5 --- 1456111_at 3.43701477 1.85785922 4 2 2.8237185 9.97969E-08 3.48E-06 Scn4b 9 45.3 Sodium channel, type IV, beta 1434008_at AI844796 3.79536664 1.63774235 3.3 2.3 2.75319499 1.48057E-08 6.21E-07 polypeptide Gadd45gip1 8 84.1 RIKEN cDNA 2310040G17 gene 1417619_at 4 3.38875643 1.4 2 2.69163229 8.84279E-06 0.0001904 BC056474 15 12.1 Mus musculus cDNA clone 1424117_at H3030A06 3.95752801 2.42838452 1.9 2.2 2.62132809 1.3344E-08 5.66E-07 MGC:67360 IMAGE:6823629, complete cds NA 4 153 guanine nucleotide binding protein, 1454696_at -3.46081884 -4 -1.3 -1.6 -2.6026947 8.58458E-05 0.0012617 beta 1 Gnb1 4 153 guanine nucleotide binding protein, 1417432_a_at H3094D02 -3.13334396 -4 -1.6 -1.7 -2.5946297 1.04542E-05 0.0002202 beta 1 Gadd45gip1 8 84.1 RAD23a homolog (S. -

Characterization and Stress Response of the Jmjc Domain-Containing Histone Demethylase Gene Family in the Allotetraploid Cotton Species Gossypium Hirsutum
plants Article Characterization and Stress Response of the JmjC Domain-Containing Histone Demethylase Gene Family in the Allotetraploid Cotton Species Gossypium hirsutum 1, 2, 2, 3 3 3 Jie Zhang y, Junping Feng y, Wei Liu *, Zhongying Ren , Junjie Zhao , Xiaoyu Pei , Yangai Liu 3, Daigang Yang 3 and Xiongfeng Ma 1,3,* 1 Zhengzhou Research Base, State Key Laboratory of Cotton Biology, School of Life Sciences, Zhengzhou University, Zhengzhou 450001, China; [email protected] 2 Collaborative Innovation Center of Henan Grain Crops, Agronomy College, Henan Agricultural University, Zhengzhou 450002, China; [email protected] 3 State Key Laboratory of Cotton Biology, Institute of Cotton Research, Chinese Academy of Agricultural Sciences, Anyang 455000, China; [email protected] (Z.R.); [email protected] (J.Z.); [email protected] (X.P.); [email protected] (Y.L.); [email protected] (D.Y.) * Correspondence: [email protected] (W.L.); [email protected] (X.M.) These authors contributed equally to this work. y Received: 12 October 2020; Accepted: 18 November 2020; Published: 20 November 2020 Abstract: Histone modification is an important epigenetic modification that controls gene transcriptional regulation in eukaryotes. Histone methylation is accomplished by histone methyltransferase and can occur on two amino acid residues, arginine and lysine. JumonjiC (JmjC) domain-containing histone demethylase regulates gene transcription and chromatin structure by changing the methylation state of the lysine residue site and plays an important role in plant growth and development. In this study, we carried out genome-wide identification and comprehensive analysis of JmjC genes in the allotetraploid cotton species Gossypium hirsutum. In total, 50 JmjC genes were identified and in G. -

PDF Download
MEK-7 Polyclonal Antibody Catalog No : YT2725 Reactivity : Human,Mouse,Rat Applications : WB,ELISA Gene Name : MAP2K7 Protein Name : Dual specificity mitogen-activated protein kinase kinase 7 Human Gene Id : 5609 Human Swiss Prot O14733 No : Mouse Gene Id : 26400 Mouse Swiss Prot Q8CE90 No : Rat Gene Id : 363855 Rat Swiss Prot No : Q4KSH7 Immunogen : The antiserum was produced against synthesized peptide derived from human MAP2K7. AA range:241-290 Specificity : MEK-7 Polyclonal Antibody detects endogenous levels of MEK-7 protein. Formulation : Liquid in PBS containing 50% glycerol, 0.5% BSA and 0.02% sodium azide. Source : Rabbit Dilution : Western Blot: 1/500 - 1/2000. ELISA: 1/20000. Not yet tested in other applications. Purification : The antibody was affinity-purified from rabbit antiserum by affinity- chromatography using epitope-specific immunogen. Concentration : 1 mg/ml 1 / 3 Storage Stability : -20°C/1 year Molecularweight : 47485 Observed Band : 47 Cell Pathway : MAPK_ERK_Growth,MAPK_G_Protein,ErbB_HER,Toll_Like,T_Cell_Receptor, Fc epsilon RI,Neurotrophin,GnRH, Background : mitogen-activated protein kinase kinase 7(MAP2K7) Homo sapiens The protein encoded by this gene is a dual specificity protein kinase that belongs to the MAP kinase kinase family. This kinase specifically activates MAPK8/JNK1 and MAPK9/JNK2, and this kinase itself is phosphorylated and activated by MAP kinase kinase kinases including MAP3K1/MEKK1, MAP3K2/MEKK2,MAP3K3/MEKK5, and MAP4K2/GCK. This kinase is involved in the signal transduction mediating the cell -

Proteome Cold-Shock Response in the Extremely Acidophilic Archaeon, Cuniculiplasma Divulgatum
microorganisms Article Proteome Cold-Shock Response in the Extremely Acidophilic Archaeon, Cuniculiplasma divulgatum Rafael Bargiela 1 , Karin Lanthaler 1,2, Colin M. Potter 1,2 , Manuel Ferrer 3 , Alexander F. Yakunin 1,2, Bela Paizs 1,2, Peter N. Golyshin 1,2 and Olga V. Golyshina 1,2,* 1 School of Natural Sciences, Bangor University, Deiniol Rd, Bangor LL57 2UW, UK; [email protected] (R.B.); [email protected] (K.L.); [email protected] (C.M.P.); [email protected] (A.F.Y.); [email protected] (B.P.); [email protected] (P.N.G.) 2 Centre for Environmental Biotechnology, Bangor University, Deiniol Rd, Bangor LL57 2UW, UK 3 Systems Biotechnology Group, Department of Applied Biocatalysis, CSIC—Institute of Catalysis, Marie Curie 2, 28049 Madrid, Spain; [email protected] * Correspondence: [email protected]; Tel.: +44-1248-388607; Fax: +44-1248-382569 Received: 27 April 2020; Accepted: 15 May 2020; Published: 19 May 2020 Abstract: The archaeon Cuniculiplasma divulgatum is ubiquitous in acidic environments with low-to-moderate temperatures. However, molecular mechanisms underlying its ability to thrive at lower temperatures remain unexplored. Using mass spectrometry (MS)-based proteomics, we analysed the effect of short-term (3 h) exposure to cold. The C. divulgatum genome encodes 2016 protein-coding genes, from which 819 proteins were identified in the cells grown under optimal conditions. In line with the peptidolytic lifestyle of C. divulgatum, its intracellular proteome revealed the abundance of proteases, ABC transporters and cytochrome C oxidase. From 747 quantifiable polypeptides, the levels of 582 proteins showed no change after the cold shock, whereas 104 proteins were upregulated suggesting that they might be contributing to cold adaptation. -

The ARID Domain Protein Dril1 Is Necessary for Tgfh Signaling in Xenopus Embryos$
Developmental Biology 278 (2005) 542–559 www.elsevier.com/locate/ydbio Genomes & Developmental Control The ARID domain protein dril1 is necessary for TGFh signaling in Xenopus embryos$ Elizabeth M. Callerya,*, James C. Smithb, Gerald H. Thomsena aDepartment of Biochemistry and Cell Biology and Center for Developmental Genetics, Stony Brook University, Stony Brook, NY 11794-5215, USA bWellcome Trust/Cancer Research UK Gordon Institute and Department of Zoology, University of Cambridge, Tennis Court Road, Cambridge CB2 1QR, UK Received for publication 14 July 2004, revised 30 October 2004, accepted 11 November 2004 Available online 15 December 2004 Abstract ARID domain proteins are members of a highly conserved family involved in chromatin remodeling and cell-fate determination. Dril1 is the founding member of the ARID family and is involved in developmental processes in both Drosophila and Caenorhabditis elegans.We describe the first embryological characterization of this gene in chordates. Dril1 mRNA expression is spatiotemporally regulated and is detected in the involuting mesoderm during gastrulation. Inhibition of dril1 by either a morpholino or an engrailed repressor–dril1 DNA binding domain fusion construct inhibits gastrulation and perturbs induction of the zygotic mesodermal marker Xbra and the organizer markers chordin, noggin, and Xlim1. Xenopus tropicalis dril1 morphants also exhibit impaired gastrulation and axial deficiencies, which can be rescued by coinjection of Xenopus laevis dril1 mRNA. Loss of dril1 inhibits the response of animal caps to activin and secondary axis induction by smad2. Dril1 depletion in animal caps prevents both the smad2-mediated induction of dorsal mesodermal and endodermal markers and the induction of ventral mesoderm by smad1. -

MAP3K12 (Human) Recombinant Protein
MAP3K12 (Human) Recombinant Gene Alias: DLK, MUK, ZPK, ZPKP1 Protein Gene Summary: The protein encoded by this gene is a member of serine/threonine protein kinase family. This Catalog Number: P5535 kinase contains a leucine-zipper domain, and is predominately expressed in neuronal cells. The Regulation Status: For research use only (RUO) phosphorylation state of this kinase in synaptic terminals Product Description: Human MAP3K12 (NP_006292.3, was shown to be regulated by membrane depolarization 1 a.a. - 520 a.a.) partial recombinant protein with GST via calcineurin. This kinase forms heterodimers with tag expressed in baculovirus infected Sf21 cells. leucine zipper containing transcription factors, such as cAMP responsive element binding protein (CREB) and Host: Insect MYC, and thus may play a regulatory role in PKA or retinoic acid induced neuronal differentiation. [provided Theoretical MW (kDa): 86 by RefSeq] Applications: Func, SDS-PAGE (See our web site product page for detailed applications information) Protocols: See our web site at http://www.abnova.com/support/protocols.asp or product page for detailed protocols Form: Liquid Preparation Method: Baculovirus infected insect cell (Sf21) expression system Purification: Glutathione sepharose chromatography Purity: 55 % by SDS-PAGE/CBB staining Activity: The activity was determined by ELISA. The enzyme was incubated with GST-fused substrate protein, and after stopping kinase reaction by EDTA, the reaction solution was transferred into glutathione-coated plate. Phosphorylation was detected by anti-phospho antibody and HRP-labeled anti-rabbit IgG (or HRP-labeled anti-mouse IgG). Substrate: MAP2K7 [inactive mutant]. ATP: 100 uM. Storage Buffer: In 50 mM Tris-HCl, 150 mM NaCl, pH 7.5 (0.1% CHAPS, 1 mM DTT, 10% glycerol) Storage Instruction: Store at -80°C. -

Table S1. List of Oligonucleotide Primers Used
Table S1. List of oligonucleotide primers used. Cla4 LF-5' GTAGGATCCGCTCTGTCAAGCCTCCGACC M629Arev CCTCCCTCCATGTACTCcgcGATGACCCAgAGCTCGTTG M629Afwd CAACGAGCTcTGGGTCATCgcgGAGTACATGGAGGGAGG LF-3' GTAGGCCATCTAGGCCGCAATCTCGTCAAGTAAAGTCG RF-5' GTAGGCCTGAGTGGCCCGAGATTGCAACGTGTAACC RF-3' GTAGGATCCCGTACGCTGCGATCGCTTGC Ukc1 LF-5' GCAATATTATGTCTACTTTGAGCG M398Arev CCGCCGGGCAAgAAtTCcgcGAGAAGGTACAGATACGc M398Afwd gCGTATCTGTACCTTCTCgcgGAaTTcTTGCCCGGCGG LF-3' GAGGCCATCTAGGCCATTTACGATGGCAGACAAAGG RF-5' GTGGCCTGAGTGGCCATTGGTTTGGGCGAATGGC RF-3' GCAATATTCGTACGTCAACAGCGCG Nrc2 LF-5' GCAATATTTCGAAAAGGGTCGTTCC M454Grev GCCACCCATGCAGTAcTCgccGCAGAGGTAGAGGTAATC M454Gfwd GATTACCTCTACCTCTGCggcGAgTACTGCATGGGTGGC LF-3' GAGGCCATCTAGGCCGACGAGTGAAGCTTTCGAGCG RF-5' GAGGCCTGAGTGGCCTAAGCATCTTGGCTTCTGC RF-3' GCAATATTCGGTCAACGCTTTTCAGATACC Ipl1 LF-5' GTCAATATTCTACTTTGTGAAGACGCTGC M629Arev GCTCCCCACGACCAGCgAATTCGATagcGAGGAAGACTCGGCCCTCATC M629Afwd GATGAGGGCCGAGTCTTCCTCgctATCGAATTcGCTGGTCGTGGGGAGC LF-3' TGAGGCCATCTAGGCCGGTGCCTTAGATTCCGTATAGC RF-5' CATGGCCTGAGTGGCCGATTCTTCTTCTGTCATCGAC RF-3' GACAATATTGCTGACCTTGTCTACTTGG Ire1 LF-5' GCAATATTAAAGCACAACTCAACGC D1014Arev CCGTAGCCAAGCACCTCGgCCGAtATcGTGAGCGAAG D1014Afwd CTTCGCTCACgATaTCGGcCGAGGTGCTTGGCTACGG LF-3' GAGGCCATCTAGGCCAACTGGGCAAAGGAGATGGA RF-5' GAGGCCTGAGTGGCCGTGCGCCTGTGTATCTCTTTG RF-3' GCAATATTGGCCATCTGAGGGCTGAC Kin28 LF-5' GACAATATTCATCTTTCACCCTTCCAAAG L94Arev TGATGAGTGCTTCTAGATTGGTGTCggcGAAcTCgAGCACCAGGTTG L94Afwd CAACCTGGTGCTcGAgTTCgccGACACCAATCTAGAAGCACTCATCA LF-3' TGAGGCCATCTAGGCCCACAGAGATCCGCTTTAATGC RF-5' CATGGCCTGAGTGGCCAGGGCTAGTACGACCTCG -

The Landscape of Genomic Imprinting Across Diverse Adult Human Tissues
Downloaded from genome.cshlp.org on September 30, 2021 - Published by Cold Spring Harbor Laboratory Press Research The landscape of genomic imprinting across diverse adult human tissues Yael Baran,1 Meena Subramaniam,2 Anne Biton,2 Taru Tukiainen,3,4 Emily K. Tsang,5,6 Manuel A. Rivas,7 Matti Pirinen,8 Maria Gutierrez-Arcelus,9 Kevin S. Smith,5,10 Kim R. Kukurba,5,10 Rui Zhang,10 Celeste Eng,2 Dara G. Torgerson,2 Cydney Urbanek,11 the GTEx Consortium, Jin Billy Li,10 Jose R. Rodriguez-Santana,12 Esteban G. Burchard,2,13 Max A. Seibold,11,14,15 Daniel G. MacArthur,3,4,16 Stephen B. Montgomery,5,10 Noah A. Zaitlen,2,19 and Tuuli Lappalainen17,18,19 1The Blavatnik School of Computer Science, Tel-Aviv University, Tel Aviv 69978, Israel; 2Department of Medicine, University of California San Francisco, San Francisco, California 94158, USA; 3Analytic and Translational Genetics Unit, Massachusetts General Hospital, Boston, Massachusetts 02114, USA; 4Program in Medical and Population Genetics, Broad Institute of Harvard and MIT, Cambridge, Massachusetts 02142, USA; 5Department of Pathology, Stanford University, Stanford, California 94305, USA; 6Biomedical Informatics Program, Stanford University, Stanford, California 94305, USA; 7Wellcome Trust Center for Human Genetics, Nuffield Department of Clinical Medicine, University of Oxford, Oxford, OX3 7BN, United Kingdom; 8Institute for Molecular Medicine Finland, University of Helsinki, 00014 Helsinki, Finland; 9Department of Genetic Medicine and Development, University of Geneva, 1211 Geneva, Switzerland;