Molecular Genetics Lab Requisition
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

SETD1B-Associated Neurodevelopmental Disorder
Genotype- phenotype correlations J Med Genet: first published as 10.1136/jmedgenet-2019-106756 on 16 June 2020. Downloaded from Original research SETD1B- associated neurodevelopmental disorder Alexandra Roston ,1 Dan Evans,2 Harinder Gill,1,3 Margaret McKinnon,1 Bertrand Isidor,4 Benjamin Cogné ,4,5 Jill Mwenifumbo,1,6 Clara van Karnebeek,7,8 Jianghong An,9 Steven J M Jones,9 Matthew Farrer,2 Michelle Demos,10 Mary Connolly,10 William T Gibson ,1 CAUSES Study, EPGEN Study ► Additional material is ABSTRACT same gene allows causal inference for those geno- published online only. To view Background Dysfunction of histone methyltransferases type–phenotype correlations. This is especially true please visit the journal online (http:// dx. doi. org/ 10. 1136/ and chromatin modifiers has been implicated in complex when perturbation of the general biological process jmedgenet- 2019- 106756). neurodevelopmental syndromes and cancers. SETD1B has, in similar contexts, been shown reproducibly encodes a lysine- specific methyltransferase that assists to result in similar phenotypes. For numbered affiliations see in transcriptional activation of genes by depositing Individual case reports have appeared in the end of article. H3K4 methyl marks. Previous reports of patients with literature describing neurodevelopmental delays rare variants in SETD1B describe a distinctive phenotype associated with rare coding variants in SETD1B, Correspondence to Dr Alexandra Roston, that includes seizures, global developmental delay and and with hemizygosity for SETD1B due to CNVs. Department of Medical intellectual disability. Here we show that pathogenic variants within Genetics, The University of Methods Two of the patients described herein were SETD1B contribute to a conserved phenotype that British Columbia, Vancouver, BC identified via genome-wide and exome-wide testing, includes intellectual disability and childhood- onset, V6T 1Z4, Canada; with microarray and research- based exome, through treatment- refractory seizures. -

Transcriptome-Wide Profiling of Cerebral Cavernous Malformations
www.nature.com/scientificreports OPEN Transcriptome-wide Profling of Cerebral Cavernous Malformations Patients Reveal Important Long noncoding RNA molecular signatures Santhilal Subhash 2,8, Norman Kalmbach3, Florian Wegner4, Susanne Petri4, Torsten Glomb5, Oliver Dittrich-Breiholz5, Caiquan Huang1, Kiran Kumar Bali6, Wolfram S. Kunz7, Amir Samii1, Helmut Bertalanfy1, Chandrasekhar Kanduri2* & Souvik Kar1,8* Cerebral cavernous malformations (CCMs) are low-fow vascular malformations in the brain associated with recurrent hemorrhage and seizures. The current treatment of CCMs relies solely on surgical intervention. Henceforth, alternative non-invasive therapies are urgently needed to help prevent subsequent hemorrhagic episodes. Long non-coding RNAs (lncRNAs) belong to the class of non-coding RNAs and are known to regulate gene transcription and involved in chromatin remodeling via various mechanism. Despite accumulating evidence demonstrating the role of lncRNAs in cerebrovascular disorders, their identifcation in CCMs pathology remains unknown. The objective of the current study was to identify lncRNAs associated with CCMs pathogenesis using patient cohorts having 10 CCM patients and 4 controls from brain. Executing next generation sequencing, we performed whole transcriptome sequencing (RNA-seq) analysis and identifed 1,967 lncRNAs and 4,928 protein coding genes (PCGs) to be diferentially expressed in CCMs patients. Among these, we selected top 6 diferentially expressed lncRNAs each having signifcant correlative expression with more than 100 diferentially expressed PCGs. The diferential expression status of the top lncRNAs, SMIM25 and LBX2-AS1 in CCMs was further confrmed by qRT-PCR analysis. Additionally, gene set enrichment analysis of correlated PCGs revealed critical pathways related to vascular signaling and important biological processes relevant to CCMs pathophysiology. -

Investigation of the Underlying Hub Genes and Molexular Pathogensis in Gastric Cancer by Integrated Bioinformatic Analyses
bioRxiv preprint doi: https://doi.org/10.1101/2020.12.20.423656; this version posted December 22, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Investigation of the underlying hub genes and molexular pathogensis in gastric cancer by integrated bioinformatic analyses Basavaraj Vastrad1, Chanabasayya Vastrad*2 1. Department of Biochemistry, Basaveshwar College of Pharmacy, Gadag, Karnataka 582103, India. 2. Biostatistics and Bioinformatics, Chanabasava Nilaya, Bharthinagar, Dharwad 580001, Karanataka, India. * Chanabasayya Vastrad [email protected] Ph: +919480073398 Chanabasava Nilaya, Bharthinagar, Dharwad 580001 , Karanataka, India bioRxiv preprint doi: https://doi.org/10.1101/2020.12.20.423656; this version posted December 22, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Abstract The high mortality rate of gastric cancer (GC) is in part due to the absence of initial disclosure of its biomarkers. The recognition of important genes associated in GC is therefore recommended to advance clinical prognosis, diagnosis and and treatment outcomes. The current investigation used the microarray dataset GSE113255 RNA seq data from the Gene Expression Omnibus database to diagnose differentially expressed genes (DEGs). Pathway and gene ontology enrichment analyses were performed, and a proteinprotein interaction network, modules, target genes - miRNA regulatory network and target genes - TF regulatory network were constructed and analyzed. Finally, validation of hub genes was performed. The 1008 DEGs identified consisted of 505 up regulated genes and 503 down regulated genes. -

Supplementary Table S4. FGA Co-Expressed Gene List in LUAD
Supplementary Table S4. FGA co-expressed gene list in LUAD tumors Symbol R Locus Description FGG 0.919 4q28 fibrinogen gamma chain FGL1 0.635 8p22 fibrinogen-like 1 SLC7A2 0.536 8p22 solute carrier family 7 (cationic amino acid transporter, y+ system), member 2 DUSP4 0.521 8p12-p11 dual specificity phosphatase 4 HAL 0.51 12q22-q24.1histidine ammonia-lyase PDE4D 0.499 5q12 phosphodiesterase 4D, cAMP-specific FURIN 0.497 15q26.1 furin (paired basic amino acid cleaving enzyme) CPS1 0.49 2q35 carbamoyl-phosphate synthase 1, mitochondrial TESC 0.478 12q24.22 tescalcin INHA 0.465 2q35 inhibin, alpha S100P 0.461 4p16 S100 calcium binding protein P VPS37A 0.447 8p22 vacuolar protein sorting 37 homolog A (S. cerevisiae) SLC16A14 0.447 2q36.3 solute carrier family 16, member 14 PPARGC1A 0.443 4p15.1 peroxisome proliferator-activated receptor gamma, coactivator 1 alpha SIK1 0.435 21q22.3 salt-inducible kinase 1 IRS2 0.434 13q34 insulin receptor substrate 2 RND1 0.433 12q12 Rho family GTPase 1 HGD 0.433 3q13.33 homogentisate 1,2-dioxygenase PTP4A1 0.432 6q12 protein tyrosine phosphatase type IVA, member 1 C8orf4 0.428 8p11.2 chromosome 8 open reading frame 4 DDC 0.427 7p12.2 dopa decarboxylase (aromatic L-amino acid decarboxylase) TACC2 0.427 10q26 transforming, acidic coiled-coil containing protein 2 MUC13 0.422 3q21.2 mucin 13, cell surface associated C5 0.412 9q33-q34 complement component 5 NR4A2 0.412 2q22-q23 nuclear receptor subfamily 4, group A, member 2 EYS 0.411 6q12 eyes shut homolog (Drosophila) GPX2 0.406 14q24.1 glutathione peroxidase -

MRTF: Basic Biology and Role in Kidney Disease
International Journal of Molecular Sciences Review MRTF: Basic Biology and Role in Kidney Disease Maria Zena Miranda 1, Zsuzsanna Lichner 1, Katalin Szászi 1,2 and András Kapus 1,2,3,* 1 Keenan Research Centre for Biomedical Science of the St. Michael’s Hospital, Toronto, ON M5B 1W8, Canada; [email protected] (M.Z.M.); [email protected] (Z.L.); [email protected] (K.S.) 2 Department of Surgery, University of Toronto, Toronto, ON M5T 1P5, Canada 3 Department of Biochemistry, University of Toronto, Toronto, ON M5S 1A8, Canada * Correspondence: [email protected] Abstract: A lesser known but crucially important downstream effect of Rho family GTPases is the regulation of gene expression. This major role is mediated via the cytoskeleton, the organization of which dictates the nucleocytoplasmic shuttling of a set of transcription factors. Central among these is myocardin-related transcription factor (MRTF), which upon actin polymerization translocates to the nucleus and binds to its cognate partner, serum response factor (SRF). The MRTF/SRF complex then drives a large cohort of genes involved in cytoskeleton remodeling, contractility, extracellular matrix organization and many other processes. Accordingly, MRTF, activated by a variety of mechanical and chemical stimuli, affects a plethora of functions with physiological and pathological relevance. These include cell motility, development, metabolism and thus metastasis formation, inflammatory responses and—predominantly-organ fibrosis. The aim of this review is twofold: to provide an up- to-date summary about the basic biology and regulation of this versatile transcriptional coactivator; and to highlight its principal involvement in the pathobiology of kidney disease. -

Neurodegeneration Induces a Developmental RNA Processing Program by Calpain
bioRxiv preprint doi: https://doi.org/10.1101/2020.07.24.219121; this version posted July 25, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Neurodegeneration induces a developmental RNA processing program by calpain- mediated MBNL2 degradation Authors/Affiliations: Lee-Hsin Wanga,b, Yu-Mei Linb, Chien-Yu Linb, Yijuang Cherna,b,c and Guey-Shin Wanga,b,c,* aTaiwan International Graduate Program in Interdisciplinary Neuroscience, National Yang- Ming University and Academia Sinica, Taipei, Taiwan bInstitute of Biomedical Sciences, Academia Sinica, Taipei 11529, Taiwan cProgram in Molecular Medicine, National Yang-Ming University and Academia Sinica, Taipei, Taiwan *To whom correspondence should be addressed: Guey-Shin Wang, PhD Institute of Biomedical Sciences Academia Sinica 128, Section 2, Academia Rd. Taipei 11529, Taiwan. Tel: (886)-226-523-051 Fax: (886)-227-829-142 Email: [email protected] 1 bioRxiv preprint doi: https://doi.org/10.1101/2020.07.24.219121; this version posted July 25, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Abstract The Muscleblind-like (MBNL) protein family plays an important role in regulating developmental RNA processing transition. Loss of MBNL2 function has been implicated in the neurodegeneration of myotonic dystrophy type 1 (DM1). However, the causal mechanism of neurodegeneration-induced MBNL2 loss of function remains elusive. Here, we show that neurodegenerative conditions including NMDAR-mediated excitotoxicity and dysregulated calcium homeostasis triggered nuclear translocation of calpain-2 resulting in MBNL2 degradation and reversion of MBNL2-regulated RNA processing to developmental patterns. -

Molecular Diagnostic Requisition
BAYLOR MIRACA GENETICS LABORATORIES SHIP TO: Baylor Miraca Genetics Laboratories 2450 Holcombe, Grand Blvd. -Receiving Dock PHONE: 800-411-GENE | FAX: 713-798-2787 | www.bmgl.com Houston, TX 77021-2024 Phone: 713-798-6555 MOLECULAR DIAGNOSTIC REQUISITION PATIENT INFORMATION SAMPLE INFORMATION NAME: DATE OF COLLECTION: / / LAST NAME FIRST NAME MI MM DD YY HOSPITAL#: ACCESSION#: DATE OF BIRTH: / / GENDER (Please select one): FEMALE MALE MM DD YY SAMPLE TYPE (Please select one): ETHNIC BACKGROUND (Select all that apply): UNKNOWN BLOOD AFRICAN AMERICAN CORD BLOOD ASIAN SKELETAL MUSCLE ASHKENAZIC JEWISH MUSCLE EUROPEAN CAUCASIAN -OR- DNA (Specify Source): HISPANIC NATIVE AMERICAN INDIAN PLACE PATIENT STICKER HERE OTHER JEWISH OTHER (Specify): OTHER (Please specify): REPORTING INFORMATION ADDITIONAL PROFESSIONAL REPORT RECIPIENTS PHYSICIAN: NAME: INSTITUTION: PHONE: FAX: PHONE: FAX: NAME: EMAIL (INTERNATIONAL CLIENT REQUIREMENT): PHONE: FAX: INDICATION FOR STUDY SYMPTOMATIC (Summarize below.): *FAMILIAL MUTATION/VARIANT ANALYSIS: COMPLETE ALL FIELDS BELOW AND ATTACH THE PROBAND'S REPORT. GENE NAME: ASYMPTOMATIC/POSITIVE FAMILY HISTORY: (ATTACH FAMILY HISTORY) MUTATION/UNCLASSIFIED VARIANT: RELATIONSHIP TO PROBAND: THIS INDIVIDUAL IS CURRENTLY: SYMPTOMATIC ASYMPTOMATIC *If family mutation is known, complete the FAMILIAL MUTATION/ VARIANT ANALYSIS section. NAME OF PROBAND: ASYMPTOMATIC/POPULATION SCREENING RELATIONSHIP TO PROBAND: OTHER (Specify clinical findings below): BMGL LAB#: A COPY OF ORIGINAL RESULTS ATTACHED IF PROBAND TESTING WAS PERFORMED AT ANOTHER LAB, CALL TO DISCUSS PRIOR TO SENDING SAMPLE. A POSITIVE CONTROL MAY BE REQUIRED IN SOME CASES. REQUIRED: NEW YORK STATE PHYSICIAN SIGNATURE OF CONSENT I certify that the patient specified above and/or their legal guardian has been informed of the benefits, risks, and limitations of the laboratory test(s) requested. -

High Constitutive Cytokine Release by Primary Human Acute Myeloid Leukemia Cells Is Associated with a Specific Intercellular Communication Phenotype
Supplementary Information High Constitutive Cytokine Release by Primary Human Acute Myeloid Leukemia Cells Is Associated with a Specific Intercellular Communication Phenotype Håkon Reikvam 1,2,*, Elise Aasebø 1, Annette K. Brenner 2, Sushma Bartaula-Brevik 1, Ida Sofie Grønningsæter 2, Rakel Brendsdal Forthun 2, Randi Hovland 3,4 and Øystein Bruserud 1,2 1 Department of Clinical Science, University of Bergen, 5020, Bergen, Norway 2 Department of Medicine, Haukeland University Hospital, 5021, Bergen, Norway 3 Department of Medical Genetics, Haukeland University Hospital, 5021, Bergen, Norway 4 Institute of Biomedicine, University of Bergen, 5020, Bergen, Norway * Correspondence: [email protected]; Tel.: +55-97-50-00 J. Clin. Med. 2019, 8, x 2 of 36 Figure S1. Mutational studies in a cohort of 71 AML patients. The figure shows the number of patients with the various mutations (upper), the number of mutations in for each patient (middle) and the number of main classes with mutation(s) in each patient (lower). 2 J. Clin. Med. 2019, 8, x; doi: www.mdpi.com/journal/jcm J. Clin. Med. 2019, 8, x 3 of 36 Figure S2. The immunophenotype of primary human AML cells derived from 62 unselected patients. The expression of the eight differentiation markers CD13, CD14, CD15, CD33, CD34, CD45, CD117 and HLA-DR was investigated for 62 of the 71 patients included in our present study. We performed an unsupervised hierarchical cluster analysis and identified four patient main clusters/patient subsets. The mutational profile for each f the 62 patients is also given (middle), no individual mutation of main class of mutations showed any significant association with any of the for differentiation marker clusters (middle). -

Substrate Stiffness-Dependent Regulation of the SRF−Mkl1 Co-Activator Complex Requires the Inner Nuclear Membrane Protein Emerin Margaret K
© 2017. Published by The Company of Biologists Ltd | Journal of Cell Science (2017) 130, 2111-2118 doi:10.1242/jcs.197517 SHORT REPORT Substrate stiffness-dependent regulation of the SRF−Mkl1 co-activator complex requires the inner nuclear membrane protein Emerin Margaret K. Willer and Christopher W. Carroll* ABSTRACT transcription is thought to underlie persistent myofibroblast The complex comprising serum response factor (SRF) and activation (Zhou et al., 2013). Thus, determining how mechanical − megakaryoblastic leukemia 1 protein (Mkl1) promotes myofibroblast cues modulate SRF Mkl1-dependent gene expression is critical to differentiation during wound healing. SRF−Mkl1 is sensitive to the understand wound healing and pathologies caused by altered ECM mechanical properties of the extracellular environment; but how cells mechanics. sense and transduce mechanical cues to modulate SRF−Mkl1- The nuclear lamina is a network of nuclear envelope-associated dependent gene expression is not well understood. Here, we intermediate filaments and their associated proteins. Recently, the demonstrate that the nuclear lamina-associated inner nuclear nuclear lamina proteins Lamins A/C and Emerin were shown to − membrane protein Emerin stimulates SRF−Mkl1-dependent gene promote SRF Mkl1-dependent transcription (Ho et al., 2014). activity in a substrate stiffness-dependent manner. Specifically, Actin-generated forces regulate Lamins A/C and Emerin (Buxboim Emerin was required for Mkl1 nuclear accumulation and maximal et al., 2014; Guilluy et al., 2014; Swift et al., 2013), but whether the − SRF−Mkl1-dependent gene expression in response to serum nuclear lamina contributes to the mechanical control of SRF Mkl1- stimulation of cells grown on stiff substrates but was dispensable on dependent gene expression has not been tested. -
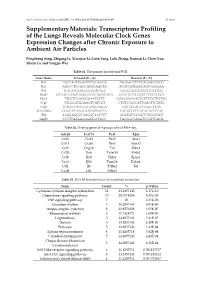
Transcriptome Profiling of the Lungs Reveals Molecular Clock Genes Expression Changes After Chronic Exposure to Ambient Air Particles
Int. J. Environ. Res. Public Health 2017, 14, 0090; doi:10.3390/ijerph14010090 S1 of S6 Supplementary Materials: Transcriptome Profiling of the Lungs Reveals Molecular Clock Genes Expression Changes after Chronic Exposure to Ambient Air Particles Pengcheng Song, Zhigang Li, Xiaoqian Li, Lixin Yang, Lulu Zhang, Nannan Li, Chen Guo, Shuyu Lu and Yongjie Wei Table S1. The primers for real-time PCR. Gene Name Forward (5'—3') Reverse (5'—3') Per1 CAGCAGTGGAGTCTGGAGGA TAGGAGCTCTGAGAAGCGGG Per2 AGCCCTGCAGCATGGAAGTA ACGTCATGAGGAGCCAGGAA Per3 TGTGTTCAAGGGTCCACTGC GGTGCTGGCAACTTCTTTCG Bmal1 CCAAGAAAGTATGGACACAGACAAA GCATTCTTGATCCTTCCTTGGT Clock TTGCTCCACGGGAATCCTT GGAGGGAAAGTGCTCTGTTGTAG Cry1 CTGGCGTGGAAGTCATCGT CTGTCCGCCATTGAGTTCTATG Cry2 TGTCCCTTCCTGTGTGGAAGA GCTCCCAGCTTGGCTTGA REV-ERBα GGGCACAAGCAACATTACCA CACGTCCCCACACACCTTAC Dbp AAGGAGCGCAAGGCAACTCT AGATGTCAAGCCTGCGCGGT Gapdh CCCTTAAGAGGGATGCTGCC TACGGCCAAATCCGTTCACA Table S2. Overlap genes of 4 groups data of RNA-Seq. Adcy9 Cxcl14 Per1 A2m Arntl Cxcl2 Per2 Alas1 Ccl11 Cxcl6 Per3 Alox15 Ccl2 Gng10 Tnf Ddit4 Ccl20 Ifnk Tnfsf10 Efnb2 Ccl9 Il10 Hif3a Epas1 Cry1 Il1b Trim16 Fabp4 Ctf1 Il6 Prl8a2 Tef Cxcl1 Lifr Prl4a1 Table S3. DAVID bioinformatics for functional annotation. Term Count % p-Value Cytokine-cytokine receptor interaction 12 34.2857143 1.27E-10 Chemokine signaling pathway 10 28.5714286 8.32E-09 TNF signaling pathway 7 20 2.31E-06 Circadian rhythm 5 14.2857143 4.03E-06 Herpes simplex infection 8 22.8571429 1.05E-05 Rheumatoid arthritis 6 17.1428571 1.60E-05 Legionellosis 5 14.2857143 5.41E-05 Malaria 5 14.2857143 6.20E-05 Pertussis 5 14.2857143 1.43E-04 African trypanosomiasis 4 11.4285714 3.62E-04 Circadian entrainment 5 14.2857143 4.28E-04 Chagas disease (American 5 14.2857143 6.21E-04 trypanosomiasis) NOD-like receptor signaling pathway 4 11.4285714 0.001137577 Jak-STAT signaling pathway 5 14.2857143 0.001482411 Inflammatory bowel disease (IBD) 4 11.4285714 0.001752567 Int. -

UC San Diego UC San Diego Electronic Theses and Dissertations
UC San Diego UC San Diego Electronic Theses and Dissertations Title Regulation of gene expression programs by serum response factor and megakaryoblastic leukemia 1/2 in macrophages Permalink https://escholarship.org/uc/item/8cc7d0t0 Author Sullivan, Amy Lynn Publication Date 2009 Peer reviewed|Thesis/dissertation eScholarship.org Powered by the California Digital Library University of California UNIVERSITY OF CALIFORNIA, SAN DIEGO Regulation of Gene Expression Programs by Serum Response Factor and Megakaryoblastic Leukemia 1/2 in Macrophages A dissertation submitted in partial satisfaction of the requirements for the degree Doctor of Philosophy in Biomedical Sciences by Amy Lynn Sullivan Committee in charge: Professor Christopher K. Glass, Chair Professor Stephen M. Hedrick Professor Marc R. Montminy Professor Nicholas J. Webster Professor Joseph L. Witztum 2009 Copyright Amy Lynn Sullivan, 2009 All rights reserved. The Dissertation of Amy Lynn Sullivan is approved, and it is acceptable in quality and form for publication on microfilm and electronically: ______________________________________________________________ ______________________________________________________________ ______________________________________________________________ ______________________________________________________________ ______________________________________________________________ Chair University of California, San Diego 2009 iii DEDICATION To my husband, Shane, for putting up with me through all of the long hours, last minute late nights, and for not letting me quit no matter how many times my projects fell apart. To my son, Tyler, for always making me smile and for making every day an adventure. To my gifted colleagues, for all of the thought-provoking discussions, technical help and moral support through the roller- coaster ride that has been my graduate career. To my family and friends, for all of your love and support. I couldn’t have done it without you! iv EPIGRAPH If at first you don’t succeed, try, try, again.