Genetic Modifiers and Rare Mendelian Disease
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Relevant Sources for Orphan Disease Prevalence Data
16 December 2014 1 EMA/452415/2012 Rev. 1 Human Medicines Research and Development Support This document was valid from 16 December 2014 to 9 January 2018. It is no longer valid. Relevant sources for orphan disease prevalence data Sponsors applying for orphan designation for a medicine under Article 3(1) a paragraph 1 of Regulation (EC) No 141/2000 on orphan medicinal products are requested tovalid provide authoritative references to demonstrate that the condition, for which the medicine is intended, does not affect more than 5 in 10,000 people in the EU at the time the application is made. Possible sources include relevant scientific literature and databases. After more than 10 years of the implementation of the Orphan Regulation the Agency has accumulated a considerable amount of data on sources of prevalence of rare diseases from the applications for orphan designation. In most cases those sources are publicly available but not easily accessible. The Agency has decided to make the information collected so far publicly available. This will decrease the administrative burden for applicants for orphan designation and thus encourage the development of medicines for rare diseases. The information is provided in the table below will be updated regularly. The information provided herewith does not replace the obligation for the sponsor under the legislation to establish the prevalence (see Regulation (EC) No 141/2000 and the Guideline on the format and content of applications for designation as orphan medicinal product, ENTR/6283/00). Sponsors are still obliged to submit an original, up-to-date prevalence calculation supported by data with their orphan designation application. -

Transformations of Lamarckism Vienna Series in Theoretical Biology Gerd B
Transformations of Lamarckism Vienna Series in Theoretical Biology Gerd B. M ü ller, G ü nter P. Wagner, and Werner Callebaut, editors The Evolution of Cognition , edited by Cecilia Heyes and Ludwig Huber, 2000 Origination of Organismal Form: Beyond the Gene in Development and Evolutionary Biology , edited by Gerd B. M ü ller and Stuart A. Newman, 2003 Environment, Development, and Evolution: Toward a Synthesis , edited by Brian K. Hall, Roy D. Pearson, and Gerd B. M ü ller, 2004 Evolution of Communication Systems: A Comparative Approach , edited by D. Kimbrough Oller and Ulrike Griebel, 2004 Modularity: Understanding the Development and Evolution of Natural Complex Systems , edited by Werner Callebaut and Diego Rasskin-Gutman, 2005 Compositional Evolution: The Impact of Sex, Symbiosis, and Modularity on the Gradualist Framework of Evolution , by Richard A. Watson, 2006 Biological Emergences: Evolution by Natural Experiment , by Robert G. B. Reid, 2007 Modeling Biology: Structure, Behaviors, Evolution , edited by Manfred D. Laubichler and Gerd B. M ü ller, 2007 Evolution of Communicative Flexibility: Complexity, Creativity, and Adaptability in Human and Animal Communication , edited by Kimbrough D. Oller and Ulrike Griebel, 2008 Functions in Biological and Artifi cial Worlds: Comparative Philosophical Perspectives , edited by Ulrich Krohs and Peter Kroes, 2009 Cognitive Biology: Evolutionary and Developmental Perspectives on Mind, Brain, and Behavior , edited by Luca Tommasi, Mary A. Peterson, and Lynn Nadel, 2009 Innovation in Cultural Systems: Contributions from Evolutionary Anthropology , edited by Michael J. O ’ Brien and Stephen J. Shennan, 2010 The Major Transitions in Evolution Revisited , edited by Brett Calcott and Kim Sterelny, 2011 Transformations of Lamarckism: From Subtle Fluids to Molecular Biology , edited by Snait B. -
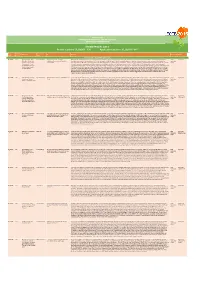
DATA Poster Numbers: P Da001 - 130 Application Posters: P Da001 - 041
POSTER LIST ORDERED ALPHABETICALLY BY POSTER TITLE GROUPED BY THEME/TRACK THEME/TRACK: DATA Poster numbers: P_Da001 - 130 Application posters: P_Da001 - 041 Poster EasyChair Presenting Author list Title Abstract Theme/track Topics number number author APPLICATION POSTERS WITHIN DATA THEME P_Da001 773 Benoît Carrères, Anne Benoît Carrères A systems approach to explore Microalgae are promising platforms for sustainable biofuel production. They produce triacyl-glycerides (TAG) which are easily converted into biofuel. When exposed to nitrogen limitation, Data/ Application Klok, Maria Suarez Diez, triacylglycerol production in Neochloris Neochloris oleoabundans accumulates up to 40% of its dry weight in TAG. However, a feasible production requires a decrease of production costs, which can be partially reached by Application Lenny de Jaeger, Mark oleoabundans increasing TAG yield.We built a constraint-based model describing primary metabolism of N. oleoabundans. It was grown in combinations of light absorption and nitrate supply rates and the poster Sturme, Packo Lamers, parameters needed for modeling of metabolism were measured. Fluxes were then calculated by flux balance analysis. cDNA samples of 16 experimental conditions were sequenced, Rene' Wijffels, Vitor Dos assembled and functionally annotated. Relative expression changes and relative flux changes for all reactions in the model were compared.The model predicts a maximum TAG yield on light Santos, Peter Schaap of 1.07g (mol photons)-1, more than 3 times current yield under optimal conditions. Furthermore, from optimization scenarios we concluded that increasing light efficiency has much higher and Dirk Martens potential to increase TAG yield than blocking entire pathways.Certain reaction expression patterns suggested an interdependence of the response to nitrogen and light supply. -

List of Genes Associated with Sudden Cardiac Death (Scdgseta) Gene
List of genes associated with sudden cardiac death (SCDgseta) mRNA expression in normal human heart Entrez_I Gene symbol Gene name Uniprot ID Uniprot name fromb D GTEx BioGPS SAGE c d e ATP-binding cassette subfamily B ABCB1 P08183 MDR1_HUMAN 5243 √ √ member 1 ATP-binding cassette subfamily C ABCC9 O60706 ABCC9_HUMAN 10060 √ √ member 9 ACE Angiotensin I–converting enzyme P12821 ACE_HUMAN 1636 √ √ ACE2 Angiotensin I–converting enzyme 2 Q9BYF1 ACE2_HUMAN 59272 √ √ Acetylcholinesterase (Cartwright ACHE P22303 ACES_HUMAN 43 √ √ blood group) ACTC1 Actin, alpha, cardiac muscle 1 P68032 ACTC_HUMAN 70 √ √ ACTN2 Actinin alpha 2 P35609 ACTN2_HUMAN 88 √ √ √ ACTN4 Actinin alpha 4 O43707 ACTN4_HUMAN 81 √ √ √ ADRA2B Adrenoceptor alpha 2B P18089 ADA2B_HUMAN 151 √ √ AGT Angiotensinogen P01019 ANGT_HUMAN 183 √ √ √ AGTR1 Angiotensin II receptor type 1 P30556 AGTR1_HUMAN 185 √ √ AGTR2 Angiotensin II receptor type 2 P50052 AGTR2_HUMAN 186 √ √ AKAP9 A-kinase anchoring protein 9 Q99996 AKAP9_HUMAN 10142 √ √ √ ANK2/ANKB/ANKYRI Ankyrin 2 Q01484 ANK2_HUMAN 287 √ √ √ N B ANKRD1 Ankyrin repeat domain 1 Q15327 ANKR1_HUMAN 27063 √ √ √ ANKRD9 Ankyrin repeat domain 9 Q96BM1 ANKR9_HUMAN 122416 √ √ ARHGAP24 Rho GTPase–activating protein 24 Q8N264 RHG24_HUMAN 83478 √ √ ATPase Na+/K+–transporting ATP1B1 P05026 AT1B1_HUMAN 481 √ √ √ subunit beta 1 ATPase sarcoplasmic/endoplasmic ATP2A2 P16615 AT2A2_HUMAN 488 √ √ √ reticulum Ca2+ transporting 2 AZIN1 Antizyme inhibitor 1 O14977 AZIN1_HUMAN 51582 √ √ √ UDP-GlcNAc: betaGal B3GNT7 beta-1,3-N-acetylglucosaminyltransfe Q8NFL0 -

Genetic Determinants Underlying Rare Diseases Identified Using Next-Generation Sequencing Technologies
Western University Scholarship@Western Electronic Thesis and Dissertation Repository 8-2-2018 1:30 PM Genetic determinants underlying rare diseases identified using next-generation sequencing technologies Rosettia Ho The University of Western Ontario Supervisor Hegele, Robert A. The University of Western Ontario Graduate Program in Biochemistry A thesis submitted in partial fulfillment of the equirr ements for the degree in Master of Science © Rosettia Ho 2018 Follow this and additional works at: https://ir.lib.uwo.ca/etd Part of the Medical Genetics Commons Recommended Citation Ho, Rosettia, "Genetic determinants underlying rare diseases identified using next-generation sequencing technologies" (2018). Electronic Thesis and Dissertation Repository. 5497. https://ir.lib.uwo.ca/etd/5497 This Dissertation/Thesis is brought to you for free and open access by Scholarship@Western. It has been accepted for inclusion in Electronic Thesis and Dissertation Repository by an authorized administrator of Scholarship@Western. For more information, please contact [email protected]. Abstract Rare disorders affect less than one in 2000 individuals, placing a huge burden on individuals, families and the health care system. Gene discovery is the starting point in understanding the molecular mechanisms underlying these diseases. The advent of next- generation sequencing has accelerated discovery of disease-causing genetic variants and is showing numerous benefits for research and medicine. I describe the application of next-generation sequencing, namely LipidSeq™ ‒ a targeted resequencing panel for the identification of dyslipidemia-associated variants ‒ and whole-exome sequencing, to identify genetic determinants of several rare diseases. Utilization of next-generation sequencing plus associated bioinformatics led to the discovery of disease-associated variants for 71 patients with lipodystrophy, two with early-onset obesity, and families with brachydactyly, cerebral atrophy, microcephaly-ichthyosis, and widow’s peak syndrome. -

The Epidemiology of Listeriosis in Pregnant
Jeffs et al. BMC Public Health (2020) 20:116 https://doi.org/10.1186/s12889-020-8221-z RESEARCH ARTICLE Open Access The epidemiology of listeriosis in pregnant women and children in New Zealand from 1997 to 2016: an observational study Emma Jeffs1, Jonathan Williman2, Cheryl Brunton2, Joanna Gullam3 and Tony Walls1* Abstract Background: Listeria monocytogenes causes the foodborne infection listeriosis. Pregnant women, infants and immunocompromised children are at increased risk for infection. The aim of this study was to describe the trends in the epidemiology of disease notifications and hospital admissions due to listeriosis in pregnant women aged 15 to 45 years and children aged less than 15 years in New Zealand (NZ) from 1997 to 2016. Methods: In this population-based descriptive study, listeriosis notification and hospitalization rates from 1997 to 2016 were analyzed. Notification data were extracted from the Institute of Environmental Science and Research (ESR) Notifiable Diseases Database (EpiSurv) and hospitalization data were extracted from the National Minimum Dataset (NMDS). Pregnant women aged 15 to 45 years and children less than 15 years of age were included. Subgroup analysis was conducted for age and ethnicity. Outcomes of infection were described. Results: In the 20-year period considered, there were 147 pregnancy-associated cases of listeriosis either notified to ESR (n = 106) and/or coded in the NMDS (n = 99), giving a crude incidence rate of 12.3 (95% CI 10.4, 14.4) per 100, 000 births. In addition, there were 22 cases in children aged 28 days to < 15 years (incidence =0.12, 95% CI 0.08 to 0.19 per 100,000). -

Basic Genetic Terms for Teachers
Student Name: Date: Class Period: Page | 1 Basic Genetic Terms Use the available reference resources to complete the table below. After finding out the definition of each word, rewrite the definition using your own words (middle column), and provide an example of how you may use the word (right column). Genetic Terms Definition in your own words An example Allele Different forms of a gene, which produce Different alleles produce different hair colors—brown, variations in a genetically inherited trait. blond, red, black, etc. Genes Genes are parts of DNA and carry hereditary Genes contain blue‐print for each individual for her or information passed from parents to children. his specific traits. Dominant version (allele) of a gene shows its Dominant When a child inherits dominant brown‐hair gene form specific trait even if only one parent passed (allele) from dad, the child will have brown hair. the gene to the child. When a child inherits recessive blue‐eye gene form Recessive Recessive gene shows its specific trait when (allele) from both mom and dad, the child will have blue both parents pass the gene to the child. eyes. Homozygous Two of the same form of a gene—one from Inheriting the same blue eye gene form from both mom and the other from dad. parents result in a homozygous gene. Heterozygous Two different forms of a gene—one from Inheriting different eye color gene forms from mom mom and the other from dad are different. and dad result in a heterozygous gene. Genotype Internal heredity information that contain Blue eye and brown eye have different genotypes—one genetic code. -

Rainbows Basic Symptom Control in Paediatric Palliative Care
Ninth edition, 2013 Basic Symptom Control in Paediatric Palliative Care The Rainbows Children’s Hospice Guidelines www.togetherforshortlives.org.uk Basic Symptom Control in Paediatric Palliative Care The Rainbows Children’s Hospice Guidelines Ninth Edition, 2013 ISBN: 1 898447284 Basic Symptom Control in Paediatric Palliative Care © Dr Satbir Singh Jassal Formulary © The Association for Paediatric Palliative Medicine (APPM), March 2012 The formulary is due for revision at the end of 2014 Author: Dr Satbir Singh Jassal B.Med Sci, B.M., B.S., DRCOG, Dip Pall Med, DFSRH, MRCGP, FRCPCH (Hon), Medical Director Rainbows Children’s Hospice and General Practitioner Production: Myra Johnson and Katrina Kelly, Together for Short Lives Design: Qube Design Associates Ltd Certified by the Information Standard Together for Short Lives is the leading UK charity that speaks for all children with life-threatening and life-limiting conditions and all those who support, love and care for them. When children are unlikely to reach adulthood, we aim to make a lifetime of difference for them and their families. Together for Short Lives 4th Floor, 48-52 Bridge House, Baldwin Street, Bristol BS1 1QB T: 0117 989 7820 [email protected] www.togetherforshortlives.org.uk Together for Short Lives is a registered charity in England and Wales (1144022) and Scotland (SC044139) and a company limited by guarantee (7783702) Disclaimer: Although Together for Short Lives has taken care to ensure that the contents of this document are correct and up to date at the time of publishing, the information contained in the document is intended for general use only. -

Gene Expression During Normal and FSHD Myogenesis Tsumagari Et Al
Gene expression during normal and FSHD myogenesis Tsumagari et al. Tsumagari et al. BMC Medical Genomics 2011, 4:67 http://www.biomedcentral.com/1755-8794/4/67 (27 September 2011) Tsumagari et al. BMC Medical Genomics 2011, 4:67 http://www.biomedcentral.com/1755-8794/4/67 RESEARCHARTICLE Open Access Gene expression during normal and FSHD myogenesis Koji Tsumagari1, Shao-Chi Chang1, Michelle Lacey2,3, Carl Baribault2,3, Sridar V Chittur4, Janet Sowden5, Rabi Tawil5, Gregory E Crawford6 and Melanie Ehrlich1,3* Abstract Background: Facioscapulohumeral muscular dystrophy (FSHD) is a dominant disease linked to contraction of an array of tandem 3.3-kb repeats (D4Z4) at 4q35. Within each repeat unit is a gene, DUX4, that can encode a protein containing two homeodomains. A DUX4 transcript derived from the last repeat unit in a contracted array is associated with pathogenesis but it is unclear how. Methods: Using exon-based microarrays, the expression profiles of myogenic precursor cells were determined. Both undifferentiated myoblasts and myoblasts differentiated to myotubes derived from FSHD patients and controls were studied after immunocytochemical verification of the quality of the cultures. To further our understanding of FSHD and normal myogenesis, the expression profiles obtained were compared to those of 19 non-muscle cell types analyzed by identical methods. Results: Many of the ~17,000 examined genes were differentially expressed (> 2-fold, p < 0.01) in control myoblasts or myotubes vs. non-muscle cells (2185 and 3006, respectively) or in FSHD vs. control myoblasts or myotubes (295 and 797, respectively). Surprisingly, despite the morphologically normal differentiation of FSHD myoblasts to myotubes, most of the disease-related dysregulation was seen as dampening of normal myogenesis- specific expression changes, including in genes for muscle structure, mitochondrial function, stress responses, and signal transduction. -

Oligogenic Inheritance of Congenital Heart Disease Involving a NKX2-5 Modifier
bioRxiv preprint doi: https://doi.org/10.1101/266726; this version posted February 20, 2018. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-ND 4.0 International license. Title: Oligogenic inheritance of congenital heart disease involving a NKX2-5 modifier Short Title: Oligogenic congenital heart disease Authors: Casey A. Gifford1,2, Sanjeev S. Ranade1,2, Ryan Samarakoon1,2, Hazel T. Salunga1,2, T. Yvanka de Soysa1,2, Yu Huang1, Ping Zhou1, Aryé Elfenbein1,2, Stacia K. Wyman1†, Yen Kim Bui1,2, Kimberly R. Cordes Metzler1,2, Philip Ursell3, Kathryn N. Ivey1,2,4§ and Deepak Srivastava1,2,4,5,* Affiliations: 1 Gladstone Institute of Cardiovascular Disease, San Francisco, CA 94158, USA 2 Roddenberry Center for Stem Cell Biology and Medicine at Gladstone, San Francisco, CA 94158, USA Departments of 3 Pathology, 4 Pediatrics, and 5 Biochemistry and Biophysics, University of California, San Francisco, San Francisco, CA 94143, USA † Current address: Innovative Genomics Institute, Berkeley, CA 94704 § Current address: Tenaya Therapeutics, South San Francisco, CA 94080 *Corresponding author: Email: [email protected] 1 bioRxiv preprint doi: https://doi.org/10.1101/266726; this version posted February 20, 2018. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-ND 4.0 International license. Abstract Complex genetic inheritance is thought to underlie many human diseases, yet experimental proof of this model has been elusive. -
Neuropathy in Childhood Mitochondrial Disease, Including Riboflavin Transporter Deficiency: Phenotype, Neurophysiology and Disease-Modifying Therapy in a Recently Described Treatable
COPYRIGHT AND USE OF THIS THESIS This thesis must be used in accordance with the provisions of the Copyright Act 1968. Reproduction of material protected by copyright may be an infringement of copyright and copyright owners may be entitled to take legal action against persons who infringe their copyright. Section 51 (2) of the Copyright Act permits an authorized officer of a university library or archives to provide a copy (by communication or otherwise) of an unpublished thesis kept in the library or archives, to a person who satisfies the authorized officer that he or she requires the reproduction for the purposes of research or study. The Copyright Act grants the creator of a work a number of moral rights, specifically the right of attribution, the right against false attribution and the right of integrity. You may infringe the author’s moral rights if you: - fail to acknowledge the author of this thesis if you quote sections from the work - attribute this thesis to another author - subject this thesis to derogatory treatment which may prejudice the author’s reputation For further information contact the University’s Copyright Service. sydney.edu.au/copyright Neuropathy in childhood mitochondrial disease, including riboflavin transporter deficiency: phenotype, neurophysiology and disease-modifying therapy in a recently described treatable disorder Dr Manoj Peter Menezes A thesis submitted for the degree of Doctor of Philosophy in the faculty of Medicine, The University of Sydney, 2015. Submitted October 2015 1 DECLARATION I hereby declare that this submission is my own work and that, to the best of my knowledge and belief, it contains no material previously published or written by another person or material which to a substantial extent has been accepted for the award of any other degree or diploma of the university or other institute of higher learning, except where due acknowledgement has been made in the text. -

Lesson Plan Mendelian Inheritance
Dolan DNA Learning Center Mendelian Inheritance __________________________________________________________________________________________ Overview • Computer with internet access This 90 minute lesson (two class periods of 45 minutes) is an introduction to Mendel’s Laws of Inheritance for students in Lesson Structure grades 5 through 8. By studying inherited traits in humans such as tasting PTC paper and inherited traits in plants such as Pre-lab (45 minutes) – Day 1 maize, we can understand how traits are passed down through Teacher Prep generations. A discussion of dominant and recessive traits in humans will encourage students to further explore their • Become familiar with Lab Center inheritance as well as their family inheritance. http://www.dnalc.org/labcenter/mendeliangenetics/m endeliangenetics_d.html Learning Outcomes • Print and copy Background Reading from the Students will be able to: Student Lab Notebook on the Lab center. • discuss the contributions of Gregor Mendel and his • Print and copy Student Pre-lab Worksheets from experiments with the garden pea. the Student Lab Notebook on the Lab center. • review the structure of DNA and chromosomes. • Cut paper strips for Sentence Strips activity. • compare a dominant trait to a recessive trait. • Make sure computers with Internet access are • compare a homozygous trait to a heterozygous trait. available. • identify traits in themselves that are either dominant or recessive. Before class • use maize as a model organism to study Mendelian Students will receive the background reading to read for inheritance. homework the night before starting lab. They will write 2 to 3 • demonstrate Mendel’s Law of Dominance and Law questions they have about the background information.