MMP14 in Sarcoma: a Regulator of Tumor Microenvironment Communication in Connective Tissues
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

A Single-Cell Transcriptional Atlas Identifies Extensive Heterogeneity in the Cellular Composition of Tendons
bioRxiv preprint doi: https://doi.org/10.1101/801266; this version posted October 10, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. A single-cell transcriptional atlas identifies extensive heterogeneity in the cellular composition of tendons Jacob B Swanson1, Andrea J De Micheli2, Nathaniel P Disser1, Leandro M Martinez1, Nicholas R Walker1,3, Benjamin D Cosgrove2, Christopher L Mendias1,3,* 1Hospital for Special Surgery, New York, NY, USA 2Meining School of Biomedical Engineering, Cornell University, Ithaca, NY, USA 3Department of Physiology and Biophysics, Weill Cornell Medical College, New York, NY, USA *Corresponding Author Christopher Mendias, PhD Hospital for Special Surgery 535 E 70th St New York, NY 10021 USA +1 212-606-1785 [email protected] Keywords: tenocyte; tendon fibroblast; pericyte; single-cell RNA sequencing bioRxiv preprint doi: https://doi.org/10.1101/801266; this version posted October 10, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Abstract Tendon is a dense, hypocellular connective tissue that transmits forces between muscles and bones. Cellular heterogeneity is increasingly recognized as an important factor in the biological basis of tissue homeostasis and disease, but little is known about the diversity of cells that populate tendon. Our objective was to explore the heterogeneity of cells in mouse Achilles tendons using single-cell RNA sequencing. We identified 13 unique cell types in tendons, including 4 previously undescribed populations of fibroblasts. -

Discovery and Optimization of Selective Inhibitors of Meprin Α (Part II)
pharmaceuticals Article Discovery and Optimization of Selective Inhibitors of Meprin α (Part II) Chao Wang 1,2, Juan Diez 3, Hajeung Park 1, Christoph Becker-Pauly 4 , Gregg B. Fields 5 , Timothy P. Spicer 1,6, Louis D. Scampavia 1,6, Dmitriy Minond 2,7 and Thomas D. Bannister 1,2,* 1 Department of Molecular Medicine, Scripps Research, Jupiter, FL 33458, USA; [email protected] (C.W.); [email protected] (H.P.); [email protected] (T.P.S.); [email protected] (L.D.S.) 2 Department of Chemistry, Scripps Research, Jupiter, FL 33458, USA; [email protected] 3 Rumbaugh-Goodwin Institute for Cancer Research, Nova Southeastern University, 3321 College Avenue, CCR r.605, Fort Lauderdale, FL 33314, USA; [email protected] 4 The Scripps Research Molecular Screening Center, Scripps Research, Jupiter, FL 33458, USA; [email protected] 5 Unit for Degradomics of the Protease Web, Institute of Biochemistry, University of Kiel, Rudolf-Höber-Str.1, 24118 Kiel, Germany; fi[email protected] 6 Department of Chemistry & Biochemistry and I-HEALTH, Florida Atlantic University, 5353 Parkside Drive, Jupiter, FL 33458, USA 7 Dr. Kiran C. Patel College of Allopathic Medicine, Nova Southeastern University, 3301 College Avenue, Fort Lauderdale, FL 33314, USA * Correspondence: [email protected] Abstract: Meprin α is a zinc metalloproteinase (metzincin) that has been implicated in multiple diseases, including fibrosis and cancers. It has proven difficult to find small molecules that are capable Citation: Wang, C.; Diez, J.; Park, H.; of selectively inhibiting meprin α, or its close relative meprin β, over numerous other metzincins Becker-Pauly, C.; Fields, G.B.; Spicer, which, if inhibited, would elicit unwanted effects. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Transmembrane/Cytoplasmic, Rather Than Catalytic, Domains of Mmp14
RESEARCH ARTICLE 343 Development 140, 343-352 (2013) doi:10.1242/dev.084236 © 2013. Published by The Company of Biologists Ltd Transmembrane/cytoplasmic, rather than catalytic, domains of Mmp14 signal to MAPK activation and mammary branching morphogenesis via binding to integrin 1 Hidetoshi Mori1,*, Alvin T. Lo1, Jamie L. Inman1, Jordi Alcaraz1,2, Cyrus M. Ghajar1, Joni D. Mott1, Celeste M. Nelson1,3, Connie S. Chen1, Hui Zhang1, Jamie L. Bascom1, Motoharu Seiki4 and Mina J. Bissell1,* SUMMARY Epithelial cell invasion through the extracellular matrix (ECM) is a crucial step in branching morphogenesis. The mechanisms by which the mammary epithelium integrates cues from the ECM with intracellular signaling in order to coordinate invasion through the stroma to make the mammary tree are poorly understood. Because the cell membrane-bound matrix metalloproteinase Mmp14 is known to play a key role in cancer cell invasion, we hypothesized that it could also be centrally involved in integrating signals for mammary epithelial cells (MECs) to navigate the collagen 1 (CL-1)-rich stroma of the mammary gland. Expression studies in nulliparous mice that carry a NLS-lacZ transgene downstream of the Mmp14 promoter revealed that Mmp14 is expressed in MECs at the tips of the branches. Using both mammary organoids and 3D organotypic cultures, we show that MMP activity is necessary for invasion through dense CL-1 (3 mg/ml) gels, but dispensable for MEC branching in sparse CL-1 (1 mg/ml) gels. Surprisingly, however, Mmp14 without its catalytic activity was still necessary for branching. Silencing Mmp14 prevented cell invasion through CL-1 and disrupted branching altogether; it also reduced integrin 1 (Itgb1) levels and attenuated MAPK signaling, disrupting Itgb1- dependent invasion/branching within CL-1 gels. -

The Capacity of Long-Term in Vitro Proliferation of Acute Myeloid
The Capacity of Long-Term in Vitro Proliferation of Acute Myeloid Leukemia Cells Supported Only by Exogenous Cytokines Is Associated with a Patient Subset with Adverse Outcome Annette K. Brenner, Elise Aasebø, Maria Hernandez-Valladares, Frode Selheim, Frode Berven, Ida-Sofie Grønningsæter, Sushma Bartaula-Brevik and Øystein Bruserud Supplementary Material S2 of S31 Table S1. Detailed information about the 68 AML patients included in the study. # of blasts Viability Proliferation Cytokine Viable cells Change in ID Gender Age Etiology FAB Cytogenetics Mutations CD34 Colonies (109/L) (%) 48 h (cpm) secretion (106) 5 weeks phenotype 1 M 42 de novo 241 M2 normal Flt3 pos 31.0 3848 low 0.24 7 yes 2 M 82 MF 12.4 M2 t(9;22) wt pos 81.6 74,686 low 1.43 969 yes 3 F 49 CML/relapse 149 M2 complex n.d. pos 26.2 3472 low 0.08 n.d. no 4 M 33 de novo 62.0 M2 normal wt pos 67.5 6206 low 0.08 6.5 no 5 M 71 relapse 91.0 M4 normal NPM1 pos 63.5 21,331 low 0.17 n.d. yes 6 M 83 de novo 109 M1 n.d. wt pos 19.1 8764 low 1.65 693 no 7 F 77 MDS 26.4 M1 normal wt pos 89.4 53,799 high 3.43 2746 no 8 M 46 de novo 26.9 M1 normal NPM1 n.d. n.d. 3472 low 1.56 n.d. no 9 M 68 MF 50.8 M4 normal D835 pos 69.4 1640 low 0.08 n.d. -

Suramin Inhibits Osteoarthritic Cartilage Degradation by Increasing Extracellular Levels
Molecular Pharmacology Fast Forward. Published on August 10, 2017 as DOI: 10.1124/mol.117.109397 This article has not been copyedited and formatted. The final version may differ from this version. MOL #109397 Suramin inhibits osteoarthritic cartilage degradation by increasing extracellular levels of chondroprotective tissue inhibitor of metalloproteinases 3 (TIMP-3). Anastasios Chanalaris, Christine Doherty, Brian D. Marsden, Gabriel Bambridge, Stephen P. Wren, Hideaki Nagase, Linda Troeberg Arthritis Research UK Centre for Osteoarthritis Pathogenesis, Kennedy Institute of Downloaded from Rheumatology, University of Oxford, Roosevelt Drive, Headington, Oxford OX3 7FY, UK (A.C., C.D., G.B., H.N., L.T.); Alzheimer’s Research UK Oxford Drug Discovery Institute, University of Oxford, Oxford, OX3 7FZ, UK (S.P.W.); Structural Genomics Consortium, molpharm.aspetjournals.org University of Oxford, Old Road Campus Research Building, Old Road Campus, Roosevelt Drive, Headington, Oxford, OX3 7DQ (BDM). at ASPET Journals on September 29, 2021 1 Molecular Pharmacology Fast Forward. Published on August 10, 2017 as DOI: 10.1124/mol.117.109397 This article has not been copyedited and formatted. The final version may differ from this version. MOL #109397 Running title: Repurposing suramin to inhibit osteoarthritic cartilage loss. Corresponding author: Linda Troeberg Address: Kennedy Institute of Rheumatology, University of Oxford, Roosevelt Drive, Headington, Oxford OX3 7FY, UK Phone number: +44 (0)1865 612600 E-mail: [email protected] Downloaded -

A Flexible Microfluidic System for Single-Cell Transcriptome Profiling
www.nature.com/scientificreports OPEN A fexible microfuidic system for single‑cell transcriptome profling elucidates phased transcriptional regulators of cell cycle Karen Davey1,7, Daniel Wong2,7, Filip Konopacki2, Eugene Kwa1, Tony Ly3, Heike Fiegler2 & Christopher R. Sibley 1,4,5,6* Single cell transcriptome profling has emerged as a breakthrough technology for the high‑resolution understanding of complex cellular systems. Here we report a fexible, cost‑efective and user‑ friendly droplet‑based microfuidics system, called the Nadia Instrument, that can allow 3′ mRNA capture of ~ 50,000 single cells or individual nuclei in a single run. The precise pressure‑based system demonstrates highly reproducible droplet size, low doublet rates and high mRNA capture efciencies that compare favorably in the feld. Moreover, when combined with the Nadia Innovate, the system can be transformed into an adaptable setup that enables use of diferent bufers and barcoded bead confgurations to facilitate diverse applications. Finally, by 3′ mRNA profling asynchronous human and mouse cells at diferent phases of the cell cycle, we demonstrate the system’s ability to readily distinguish distinct cell populations and infer underlying transcriptional regulatory networks. Notably this provided supportive evidence for multiple transcription factors that had little or no known link to the cell cycle (e.g. DRAP1, ZKSCAN1 and CEBPZ). In summary, the Nadia platform represents a promising and fexible technology for future transcriptomic studies, and other related applications, at cell resolution. Single cell transcriptome profling has recently emerged as a breakthrough technology for understanding how cellular heterogeneity contributes to complex biological systems. Indeed, cultured cells, microorganisms, biopsies, blood and other tissues can be rapidly profled for quantifcation of gene expression at cell resolution. -

Gene Symbol Category ACAN ECM ADAM10 ECM Remodeling-Related ADAM11 ECM Remodeling-Related ADAM12 ECM Remodeling-Related ADAM15 E
Supplementary Material (ESI) for Integrative Biology This journal is (c) The Royal Society of Chemistry 2010 Gene symbol Category ACAN ECM ADAM10 ECM remodeling-related ADAM11 ECM remodeling-related ADAM12 ECM remodeling-related ADAM15 ECM remodeling-related ADAM17 ECM remodeling-related ADAM18 ECM remodeling-related ADAM19 ECM remodeling-related ADAM2 ECM remodeling-related ADAM20 ECM remodeling-related ADAM21 ECM remodeling-related ADAM22 ECM remodeling-related ADAM23 ECM remodeling-related ADAM28 ECM remodeling-related ADAM29 ECM remodeling-related ADAM3 ECM remodeling-related ADAM30 ECM remodeling-related ADAM5 ECM remodeling-related ADAM7 ECM remodeling-related ADAM8 ECM remodeling-related ADAM9 ECM remodeling-related ADAMTS1 ECM remodeling-related ADAMTS10 ECM remodeling-related ADAMTS12 ECM remodeling-related ADAMTS13 ECM remodeling-related ADAMTS14 ECM remodeling-related ADAMTS15 ECM remodeling-related ADAMTS16 ECM remodeling-related ADAMTS17 ECM remodeling-related ADAMTS18 ECM remodeling-related ADAMTS19 ECM remodeling-related ADAMTS2 ECM remodeling-related ADAMTS20 ECM remodeling-related ADAMTS3 ECM remodeling-related ADAMTS4 ECM remodeling-related ADAMTS5 ECM remodeling-related ADAMTS6 ECM remodeling-related ADAMTS7 ECM remodeling-related ADAMTS8 ECM remodeling-related ADAMTS9 ECM remodeling-related ADAMTSL1 ECM remodeling-related ADAMTSL2 ECM remodeling-related ADAMTSL3 ECM remodeling-related ADAMTSL4 ECM remodeling-related ADAMTSL5 ECM remodeling-related AGRIN ECM ALCAM Cell-cell adhesion ANGPT1 Soluble factors and receptors -
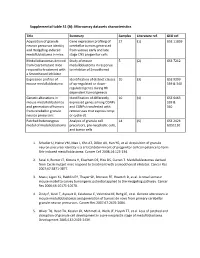
Supplemental Table S1 (A): Microarray Datasets Characteristics
Supplemental table S1 (A): Microarray datasets characteristics Title Summary Samples Literature ref. GEO ref. Acquisition of granule Gene expression profiling of 27 (1) GSE 11859 neuron precursor identity cerebellar tumors generated and Hedgehog‐induced from various early and late medulloblastoma in mice. stage CNS progenitor cells Medulloblastomas derived Study of mouse 5 (2) GSE 7212 from Cxcr6 mutant mice medulloblastoma in response respond to treatment with to inhibitor of Smoothened a Smoothened inhibitor Expression profiles of Identification of distinct classes 10 (3) GSE 9299 mouse medulloblastoma of up‐regulated or down‐ 339 & 340 regulated genes during Hh dependent tumorigenesis Genetic alterations in Identification of differently 10 (4) GSE 6463 mouse medulloblastomas expressed genes among CGNPs 339 & and generation of tumors and CGNPs transfected with 340 from cerebellar granule retroviruses that express nmyc neuron precursors or cyclin‐d1 Patched heterozygous Analysis of granule cell 14 (5) GSE 2426 model of medulloblastoma precursors, pre‐neoplastic cells, GDS1110 and tumor cells 1. Schuller U, Heine VM, Mao J, Kho AT, Dillon AK, Han YG, et al. Acquisition of granule neuron precursor identity is a critical determinant of progenitor cell competence to form Shh‐induced medulloblastoma. Cancer Cell 2008;14:123‐134. 2. Sasai K, Romer JT, Kimura H, Eberhart DE, Rice DS, Curran T. Medulloblastomas derived from Cxcr6 mutant mice respond to treatment with a smoothened inhibitor. Cancer Res 2007;67:3871‐3877. 3. Mao J, Ligon KL, Rakhlin EY, Thayer SP, Bronson RT, Rowitch D, et al. A novel somatic mouse model to survey tumorigenic potential applied to the Hedgehog pathway. Cancer Res 2006;66:10171‐10178. -

Cleavage of Lumican by Membrane-Type Matrix Metalloproteinase-1 Abrogates This Proteoglycan-Mediated Suppression of Tumor Cell Colony Formation in Soft Agar
[CANCER RESEARCH 64, 7058–7064, October 1, 2004] Cleavage of Lumican by Membrane-Type Matrix Metalloproteinase-1 Abrogates This Proteoglycan-Mediated Suppression of Tumor Cell Colony Formation in Soft Agar Yingyi Li,1 Takanori Aoki,1,3 Yuya Mori,1 Munirah Ahmad,1 Hisashi Miyamori,1 Takahisa Takino,1 and Hiroshi Sato1,2 1Department of Molecular Virology and Oncology and 2Center for the Development of Molecular Target Drugs, Cancer Research Institute, Kanazawa University, Kanazawa, Ishikawa, Japan; and 3Daiichi Fine Chemical Co. Ltd., Takaoka, Toyama, Japan ABSTRACT or tumorigenicity in nude mice of rat fibroblast F204 cells trans- formed by K-ras or v-src oncogene (12). The small leucine-rich proteoglycan lumican was identified from a Matrix metalloproteinases (MMPs) are a family of Zn2ϩ-dependent human placenta cDNA library by the expression cloning method as a gene product that interacts with membrane-type matrix metalloproteinase-1 enzymes that are known to cleave extracellular matrix proteins in (MT1-MMP). Coexpression of MT1-MMP with lumican in HEK293T cells normal and pathological conditions (13). Currently, 23 MMP genes reduced the concentration of lumican secreted into culture medium, and have been identified in humans, and they can be subgrouped into this reduction was abolished by addition of the MMP inhibitor BB94. soluble-type and membrane-type MMPs (MT-MMPs; refs. 13, 14). Lumican protein from bovine cornea and recombinant lumican core MMPs are overexpressed in various human malignancies and have protein fused to glutathione S-transferase was shown to be cleaved at been thought to contribute to tumor invasion and metastasis by de- multiple sites by recombinant MT1-MMP. -

Investigation of COVID-19 Comorbidities Reveals Genes and Pathways Coincident with the SARS-Cov-2 Viral Disease
bioRxiv preprint doi: https://doi.org/10.1101/2020.09.21.306720; this version posted September 21, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-ND 4.0 International license. Title: Investigation of COVID-19 comorbidities reveals genes and pathways coincident with the SARS-CoV-2 viral disease. Authors: Mary E. Dolan1*,2, David P. Hill1,2, Gaurab Mukherjee2, Monica S. McAndrews2, Elissa J. Chesler2, Judith A. Blake2 1 These authors contributed equally and should be considered co-first authors * Corresponding author [email protected] 2 The Jackson Laboratory, 600 Main St, Bar Harbor, ME 04609, USA Abstract: The emergence of the SARS-CoV-2 virus and subsequent COVID-19 pandemic initiated intense research into the mechanisms of action for this virus. It was quickly noted that COVID-19 presents more seriously in conjunction with other human disease conditions such as hypertension, diabetes, and lung diseases. We conducted a bioinformatics analysis of COVID-19 comorbidity-associated gene sets, identifying genes and pathways shared among the comorbidities, and evaluated current knowledge about these genes and pathways as related to current information about SARS-CoV-2 infection. We performed our analysis using GeneWeaver (GW), Reactome, and several biomedical ontologies to represent and compare common COVID- 19 comorbidities. Phenotypic analysis of shared genes revealed significant enrichment for immune system phenotypes and for cardiovascular-related phenotypes, which might point to alleles and phenotypes in mouse models that could be evaluated for clues to COVID-19 severity. -

Mir-205: a Potential Biomedicine for Cancer Therapy
cells Review miR-205: A Potential Biomedicine for Cancer Therapy Neeraj Chauhan 1,2 , Anupam Dhasmana 1,2, Meena Jaggi 1,2, Subhash C. Chauhan 1,2 and Murali M. Yallapu 1,2,* 1 Department of Immunology and Microbiology, School of Medicine, University of Texas Rio Grande Valley, McAllen, TX 78504, USA; [email protected] (N.C.); [email protected] (A.D.); [email protected] (M.J.); [email protected] (S.C.C.) 2 South Texas Center of Excellence in Cancer Research, School of Medicine, University of Texas Rio Grande Valley, McAllen, TX 78504, USA * Correspondence: [email protected]; Tel.: +1-(956)-296-1734 Received: 3 June 2020; Accepted: 21 August 2020; Published: 25 August 2020 Abstract: microRNAs (miRNAs) are a class of small non-coding RNAs that regulate the expression of their target mRNAs post transcriptionally. miRNAs are known to regulate not just a gene but the whole gene network (signaling pathways). Accumulating evidence(s) suggests that miRNAs can work either as oncogenes or tumor suppressors, but some miRNAs have a dual nature since they can act as both. miRNA 205 (miR-205) is one such highly conserved miRNA that can act as both, oncomiRNA and tumor suppressor. However, most reports confirm its emerging role as a tumor suppressor in many cancers. This review focuses on the downregulated expression of miR-205 and discusses its dysregulation in breast, prostate, skin, liver, gliomas, pancreatic, colorectal and renal cancers. This review also confers its role in tumor initiation, progression, cell proliferation, epithelial to mesenchymal transition, and tumor metastasis.