Genomic Non-Redundancy of the Mir-183/96/182 Cluster and Its
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Distinct E-Cadherin-Based Complexes Regulate Cell Behaviour Through Mirna Processing Or Src and P120 Catenin Activity
ARTICLES Distinct E-cadherin-based complexes regulate cell behaviour through miRNA processing or Src and p120 catenin activity Antonis Kourtidis1, Siu P. Ngok1,5, Pamela Pulimeno2,5, Ryan W. Feathers1, Lomeli R. Carpio1, Tiffany R. Baker1, Jennifer M. Carr1, Irene K. Yan1, Sahra Borges1, Edith A. Perez3, Peter Storz1, John A. Copland1, Tushar Patel1, E. Aubrey Thompson1, Sandra Citi4 and Panos Z. Anastasiadis1,6 E-cadherin and p120 catenin (p120) are essential for epithelial homeostasis, but can also exert pro-tumorigenic activities. Here, we resolve this apparent paradox by identifying two spatially and functionally distinct junctional complexes in non-transformed polarized epithelial cells: one growth suppressing at the apical zonula adherens (ZA), defined by the p120 partner PLEKHA7 and a non-nuclear subset of the core microprocessor components DROSHA and DGCR8, and one growth promoting at basolateral areas of cell–cell contact containing tyrosine-phosphorylated p120 and active Src. Recruitment of DROSHA and DGCR8 to the ZA is PLEKHA7 dependent. The PLEKHA7–microprocessor complex co-precipitates with primary microRNAs (pri-miRNAs) and possesses pri-miRNA processing activity. PLEKHA7 regulates the levels of select miRNAs, in particular processing of miR-30b, to suppress expression of cell transforming markers promoted by the basolateral complex, including SNAI1, MYC and CCND1. Our work identifies a mechanism through which adhesion complexes regulate cellular behaviour and reveals their surprising association with the microprocessor. p120 catenin (p120) was identified as a tyrosine phosphorylation with this, E-cadherin is still expressed in several types of aggressive substrate of the Src oncogene1 and an essential component of the and metastatic cancer18–20. -

A Molecular and Genetic Analysis of Otosclerosis
A molecular and genetic analysis of otosclerosis Joanna Lauren Ziff Submitted for the degree of PhD University College London January 2014 1 Declaration I, Joanna Ziff, confirm that the work presented in this thesis is my own. Where information has been derived from other sources, I confirm that this has been indicated in the thesis. Where work has been conducted by other members of our laboratory, this has been indicated by an appropriate reference. 2 Abstract Otosclerosis is a common form of conductive hearing loss. It is characterised by abnormal bone remodelling within the otic capsule, leading to formation of sclerotic lesions of the temporal bone. Encroachment of these lesions on to the footplate of the stapes in the middle ear leads to stapes fixation and subsequent conductive hearing loss. The hereditary nature of otosclerosis has long been recognised due to its recurrence within families, but its genetic aetiology is yet to be characterised. Although many familial linkage studies and candidate gene association studies to investigate the genetic nature of otosclerosis have been performed in recent years, progress in identifying disease causing genes has been slow. This is largely due to the highly heterogeneous nature of this condition. The research presented in this thesis examines the molecular and genetic basis of otosclerosis using two next generation sequencing technologies; RNA-sequencing and Whole Exome Sequencing. RNA–sequencing has provided human stapes transcriptomes for healthy and diseased stapes, and in combination with pathway analysis has helped identify genes and molecular processes dysregulated in otosclerotic tissue. Whole Exome Sequencing has been employed to investigate rare variants that segregate with otosclerosis in affected families, and has been followed by a variant filtering strategy, which has prioritised genes found to be dysregulated during RNA-sequencing. -

Mir-96 Regulates the Progression of Differentiation in Mammalian Cochlear Inner and Outer Hair Cells
miR-96 regulates the progression of differentiation in mammalian cochlear inner and outer hair cells Stephanie Kuhna,1, Stuart L. Johnsona,1, David N. Furnessb, Jing Chenc, Neil Inghamc, Jennifer M. Hiltonc, Georg Steffesc, Morag A. Lewisc, Valeria Zampinia,d, Carole M. Hackneya, Sergio Masettod, Matthew C. Holleya, Karen P. Steelc, and Walter Marcottia,2 aDepartment of Biomedical Science, University of Sheffield, Sheffield S10 2TN, United Kingdom; bInstitute for Science and Technology in Medicine, Keele University, Keele ST5 5BG, United Kingdom; cWellcome Trust Sanger Institute, Hinxton, Cambridge CB10 1SA, United Kingdom; and dDepartment of Physiology, University of Pavia, 27100 Pavia, Italy Edited by Mary-Claire King, University of Washington, Seattle, WA, and approved December 27, 2010 (received for review November 8, 2010) MicroRNAs (miRNAs) are small noncoding RNAs able to regulate cells up to at least P5 (23) and in the spiral ganglion up to P14 a broad range of protein-coding genes involved in many biological (19). Mutations in miR-96 have been associated with non- processes. miR-96 is a sensory organ-specific miRNA expressed in syndromic progressive hearing loss in humans (15) and mice (23). the mammalian cochlea during development. Mutations in miR-96 We found that hair cell development in diminuendo mice with cause nonsyndromic progressive hearing loss in humans and mice. a Mir96 mutation is arrested at around the day of birth, a time The mouse mutant diminuendo has a single base change in the when IHCs and OHCs still exhibit qualitatively similar bio- seed region of the Mir96 gene leading to widespread changes in physical properties. -

Microrna-1182 and Let-7A Exert Synergistic Inhibition on Invasion
Pan et al. Cancer Cell Int (2021) 21:161 https://doi.org/10.1186/s12935-021-01797-z Cancer Cell International PRIMARY RESEARCH Open Access MicroRNA-1182 and let-7a exert synergistic inhibition on invasion, migration and autophagy of cholangiocarcinoma cells through down-regulation of NUAK1 Xin Pan*, Gang Wang and Baoming Wang Abstract Background: Cholangiocarcinoma (CCA) is the second most common primary liver malignancy worldwide. Several microRNAs (miRNAs) have been implicated as potential tumor suppressors in CCA. This study aims to explore the potential efects of miR-1182 and let-7a on CCA development. Methods: Bioinformatics analysis was conducted to screen diferentially expressed genes in CCA, Western blot analysis detected NUAK1 protein expression and RT-qPCR detected miR-1182, let-7a and NUAK1 expression in CCA tissues and cell lines. Dual luciferase reporter gene assay and RIP were applied to validate the relationship between miR-1182 and NUAK1 as well as between let-7a and NUAK1. Functional experiment was conducted to investigate the role of miR-1182, let-7a and NUAK1 in cell migration, proliferation and autophagy. Then, the CCA cells that received various treatments were implanted to mice to establish animal model, followed by tumor observation and HE staining to evaluate lung metastasis. Results: CCA tissues and cells were observed to have a high expression of NUAK1 and poor expression of miR-1182 and let-7a. NUAK1 was indicated as a target gene of miR-1182 and let-7a. Importantly, upregulation of either miR-1182 or let-7a induced autophagy, and inhibited cell progression and in vivo tumor growth and lung metastasis; moreover, combined treatment of miR-1182 and let-7a overexpression presented with enhanced inhibitory efect on NUAK1 expression and CCA progression, but such synergistic efect could be reversed by overexpression of NUAK1. -

The Oncogenic Role of Mir-155 in Breast Cancer
Published OnlineFirst June 26, 2012; DOI: 10.1158/1055-9965.EPI-12-0173 Cancer Epidemiology, MiniReview Biomarkers & Prevention The Oncogenic Role of miR-155 in Breast Cancer Sam Mattiske, Rachel J. Suetani, Paul M. Neilsen, and David F. Callen Abstract miR-155isanoncogenicmiRNAwithwelldescribedrolesinleukemia.However,additionalrolesof miR-155 in breast cancer progression have recently been described. A thorough literature search was conducted to review all published data to date, examining the role of miR-155 in breast cancer. Data on all validated miR-155 target genes was collated to identify biologic pathways relevant to miR-155 and breast cancer progression. Publications describing the clinical relevance, functional characterization, and regu- lation of expression of miR-155 in the context of breast cancer are reviewed. A total of 147 validated miR- 155 target genes were identified from the literature. Pathway analysis of these genes identified likely roles in apoptosis, differentiation, angiogenesis, proliferation, and epithelial–mesenchymal transition. The large number of validated miR-155 targets presented here provide many avenues of interest as to the clinical potential of miR-155. Further investigation of these target genes will be required to elucidate the specific mechanisms and functions of miR-155 in breast cancer. This is the first review examining the role of miR- 155 in breast cancer progression. The collated data of target genes and biologic pathways of miR-155 identified in this review suggest new avenues of research for this oncogenic miRNA. Cancer Epidemiol Biomarkers Prev; 21(8); 1236–43. Ó2012 AACR. Introduction found to regulate levels of LIN-14 protein (7, 8). Since this miRNAs are small noncoding RNAs that control discovery, there have been over 500 miRNAs described, expression of target genes by either inhibiting protein regulating a wide range of genes and cellular processes, translation or directly targeting mRNA transcripts of although the total predicted number of unique miRNAs target genes for degradation (1). -

Arsenic Trioxide-Mediated Suppression of Mir-182-5P Is Associated with Potent Anti-Oxidant Effects Through Up-Regulation of SESN2
www.impactjournals.com/oncotarget/www.oncotarget.com Oncotarget, 2018,Oncotarget, Vol. 9, (No.Advance 22), Publicationspp: 16028-16042 2018 Research Paper Arsenic trioxide-mediated suppression of miR-182-5p is associated with potent anti-oxidant effects through up-regulation of SESN2 Liang-Ting Lin1,10,*, Shin-Yi Liu2,*, Jyh-Der Leu3,4,*, Chun-Yuan Chang1, Shih-Hwa Chiou5,6,7, Te-Chang Lee7,8 and Yi-Jang Lee1,9 1Department of Biomedical Imaging and Radiological Sciences, National Yang-Ming University, Taipei, Taiwan 2Department of Radiation Oncology, MacKay Memorial Hospital, Taipei, Taiwan 3Division of Radiation Oncology, Taipei City Hospital Ren Ai Branch, Taipei, Taiwan 4Institute of Neuroscience, National Chengchi University, Taipei, Taiwan 5Department of Medical Research and Education, Taipei Veterans General Hospital, Taipei, Taiwan 6Institute of Clinical Medicine, School of Medicine, National Yang-Ming University, Taipei, Taiwan 7Institute of Pharmacology, National Yang-Ming University, Taipei, Taiwan 8Institute of Biomedical Sciences, Academia Sinica, Taipei, Taiwan 9Biophotonics and Molecular Imaging Research Center (BMIRC), National Yang-Ming University, Taipei, Taiwan 10Current address: Department of Health Technology and Informatics, The Hong Kong Polytechnic University, Hong Kong *These authors have contributed equally to this work Correspondence to: Te-Chang Lee, email: [email protected] Yi-Jang Lee, email: [email protected] Keywords: arsenic trioxide; sestrin 2; miR-182; oxidative stress; anti-oxidant effect Received: April 12, 2017 Accepted: February 24, 2018 Published: March 23, 2018 Copyright: Lin et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License 3.0 (CC BY 3.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited. -

Mutations Suggests Both Loss-Of-Function and Gain-Of-Function Effects Morag A
© 2021. Published by The Company of Biologists Ltd | Disease Models & Mechanisms (2021) 14, dmm047225. doi:10.1242/dmm.047225 RESEARCH ARTICLE Hearing impairment due to Mir183/96/182 mutations suggests both loss-of-function and gain-of-function effects Morag A. Lewis1,2,*, Francesca Di Domenico1, Neil J. Ingham1,2, Haydn M. Prosser2 and Karen P. Steel1,2 ABSTRACT birth. However, this is not the cause of the hearing loss; even before The microRNA miR-96 is important for hearing, as point mutations in the onset of normal hearing, homozygote hair cells fail to mature humans and mice result in dominant progressive hearing loss. Mir96 both morphologically and physiologically, remaining in their is expressed in sensory cells along with Mir182 and Mir183, but the immature state, and heterozygote hair cells show a developmental roles of these closely-linked microRNAs are as yet unknown. Here, delay. miR-96 is thus thought to be responsible for coordinating we analyse mice carrying null alleles of Mir182, and of Mir183 and hair cell maturation (Chen et al., 2014; Kuhn et al., 2011). Mir96 together to investigate their roles in hearing. We found that Overexpression of the three miRNAs also results in cochlear defects Mir183/96 heterozygous mice had normal hearing and homozygotes and hearing loss (Weston et al., 2018). The complete loss of all were completely deaf with abnormal hair cell stereocilia bundles and mature miRNAs from the inner ear results in early developmental reduced numbers of inner hair cell synapses at 4 weeks of age. defects including a severely truncated cochlear duct (Friedman Mir182 knockout mice developed normal hearing then exhibited et al., 2009; Soukup et al., 2009). -

Timo Lassmann, Yoshiko Maida 2, Yasuhiro Tomaru, Mami Yasukawa
Int. J. Mol. Sci. 2015, 16, 1192-1208; doi:10.3390/ijms16011192 OPEN ACCESS International Journal of Molecular Sciences ISSN 1422-0067 www.mdpi.com/journal/ijms Article Telomerase Reverse Transcriptase Regulates microRNAs Timo Lassmann 1,†, Yoshiko Maida 2,†, Yasuhiro Tomaru 1, Mami Yasukawa 2, Yoshinari Ando 1, Miki Kojima 1, Vivi Kasim 2, Christophe Simon 1, Carsten O. Daub 1, Piero Carninci 1, Yoshihide Hayashizaki 1,* and Kenkichi Masutomi 2,3,* 1 RIKEN Omics Science Center, RIKEN Yokohama Institute, 1-7-22 Suehiro-cho, Tsurumi-ku, Yokohama 230-0045, Japan; E-Mails: [email protected] (T.L.); [email protected] (Y.T.); [email protected] (Y.A.); [email protected] (M.K.); [email protected] (C.S.); [email protected] (C.O.D.); [email protected] (P.C.) 2 Cancer Stem Cell Project, National Cancer Center Research Institute, 5-1-1 Tsukiji, Chuo-ku, Tokyo 104-0045, Japan; E-Mails: [email protected] (Y.M.); [email protected] (M.Y.); [email protected] (V.K.) 3 Precursory Research for Embryonic Science and Technology (PREST), Japan Science and Technology Agency, 4-1-8 Honcho Kawaguchi, Saitama 332-0012, Japan † These authors contributed equally to this work. * Authors to whom correspondence should be addressed; E-Mails: [email protected] (Y.H.); [email protected] (K.M.); Tel.: +81-45-503-9248 (Y.H.); +81-3-3547-5173 (K.M.); Fax: +81-45-503-9216 (Y.H.); +81-3-3547-5123 (K.M.). Academic Editor: Martin Pichler Received: 22 November 2014 / Accepted: 26 December 2014 / Published: 6 January 2015 Abstract: MicroRNAs are small non-coding RNAs that inhibit the translation of target mRNAs. -

Mono-Uridylation of Pre-Microrna As a Key Step in the Biogenesis of Group II Let-7 Micrornas
Mono-Uridylation of Pre-MicroRNA as a Key Step in the Biogenesis of Group II let-7 MicroRNAs Inha Heo,1,2,3 Minju Ha,1,2,3 Jaechul Lim,1,2 Mi-Jeong Yoon,2 Jong-Eun Park,1,2 S. Chul Kwon,1,2 Hyeshik Chang,1,2 and V. Narry Kim1,2,* 1Institute for Basic Science 2School of Biological Sciences Seoul National University, Seoul 151-742, Korea 3These authors contributed equally to this work *Correspondence: [email protected] http://dx.doi.org/10.1016/j.cell.2012.09.022 SUMMARY hairpin (Denli et al., 2004; Gregory et al., 2004; Han et al., 2004, 2006; Landthaler et al., 2004). Like other RNase-III-type RNase III Drosha initiates microRNA (miRNA) matu- endonucleases, Drosha introduces a staggered cut such that ration by cleaving a primary miRNA transcript and the product acquires a characteristic 2 nt overhang at the 30 releasing a pre-miRNA with a 2 nt 30 overhang. Dicer terminus. After cleavage, the pre-miRNA is exported to the recognizes the 2 nt 30 overhang structure to selec- cytoplasm by exportin 5 in a complex with Ran-GTP (Bohnsack tively process pre-miRNAs. Here, we find that, unlike et al., 2004; Lund et al., 2004; Yi et al., 2003). The cytoplasmic prototypic pre-miRNAs (group I), group II pre- RNase III Dicer processes the pre-miRNA further to liberate a small RNA duplex (Bernstein et al., 2001; Grishok et al., miRNAs acquire a shorter (1 nt) 30 overhang from 0 2001; Hutva´ gner et al., 2001; Ketting et al., 2001; Knight and Drosha processing and therefore require a 3 -end Bass, 2001). -
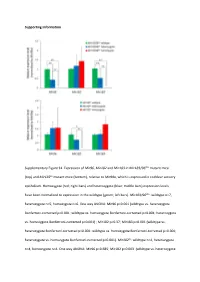
Supporting Information Supplementary Figure S1
Supporting Information Supplementary Figure S1. Expression of Mir96, Mir182 and Mir183 in Mir183/96dko mutant mice (top) and Mir182ko mutant mice (bottom), relative to Mir99a, which is expressed in cochlear sensory epithelium. Homozygote (red; right bars) and heterozygote (blue; middle bars) expression levels have been normalised to expression in the wildtype (green; left bars). Mir183/96dko: wildtype n=7, heterozygote n=5, homozygote n=6. One way ANOVA: Mir96 p<0.001 (wildtype vs. heterozygote Bonferroni‐corrected p<0.001; wildtype vs. homozygote Bonferroni‐corrected p<0.001; heterozygote vs. homozygote Bonferroni‐corrected p=0.001) ; Mir182 p=0.37; Mir183 p<0.001 (wildtype vs. heterozygote Bonferroni‐corrected p=0.001; wildtype vs. homozygote Bonferroni‐corrected p<0.001; heterozygote vs. homozygote Bonferroni‐corrected p<0.001). Mir182ko: wildtype n=4, heterozygote n=4, homozygote n=4. One way ANOVA: Mir96 p=0.685; Mir182 p=0.003 (wildtype vs. heterozygote Bonferroni‐corrected p=0.397; wildtype vs. homozygote Bonferroni‐corrected p=0.003; heterozygote vs. homozygote Bonferroni‐corrected p=0.032); Mir183 p=0.04 (wildtype vs. heterozygote Bonferroni‐corrected p=1.0; wildtype vs. homozygote Bonferroni‐corrected p=0.068; heterozygote vs. homozygote Bonferroni‐corrected p=0.094), Error bars are standard deviation (* = P < 0.05, ** = P ≤ 0.01). Supplementary Figure S2. Individual ABR thresholds of wildtype, heterozygous and homozygous Mir183/96dko mice at all ages tested. Number of mice of each genotype tested at each age is shown on the threshold plot. Supplementary Figure S3. Individual ABR thresholds of wildtype, heterozygous and homozygous Mir182ko mice at all ages tested. -

A Novel PLEKHA7 Interactor at Adherens Junctions
Thesis PDZD11: a novel PLEKHA7 interactor at adherens junctions GUERRERA, Diego Abstract PLEKHA7 is a recently identified protein of the AJ that has been involved by genetic and genomic studies in the regulation of miRNA signaling and cardiac contractility, hypertension and glaucoma. However, the molecular mechanisms behind PLEKHA7 involvement in tissue physiology and pathology remain unknown. In my thesis I report novel results which uncover PLEKHA7 functions in epithelial and endothelial cells, through the identification of a novel molecular interactor of PLEKHA7, PDZD11, by yeast two-hybrid screening, mass spectrometry, co-immunoprecipitation and pulldown assays. I dissected the structural basis of their interaction, showing that the WW domain of PLEKHA7 binds to the N-terminal region of PDZD11; this interaction mediates the junctional recruitment of PDZD11, identifying PDZD11 as a novel AJ protein. I provided evidence that PDZD11 forms a complex with nectins at AJ, its PDZ domain binds to the PDZ-binding motif of nectins. PDZD11 stabilizes nectins promoting the early steps of junction assembly. Reference GUERRERA, Diego. PDZD11: a novel PLEKHA7 interactor at adherens junctions. Thèse de doctorat : Univ. Genève, 2016, no. Sc. 4962 URN : urn:nbn:ch:unige-877543 DOI : 10.13097/archive-ouverte/unige:87754 Available at: http://archive-ouverte.unige.ch/unige:87754 Disclaimer: layout of this document may differ from the published version. 1 / 1 UNIVERSITE DE GENÈVE FACULTE DES SCIENCES Section de Biologie Prof. Sandra Citi Département de Biologie Cellulaire PDZD11: a novel PLEKHA7 interactor at adherens junctions THÈSE Présentée à la Faculté des sciences de l’Université de Genève Pour obtenir le grade de Doctor ès science, mention Biologie par DIEGO GUERRERA de Benevento (Italie) Thèse N° 4962 GENÈVE Atelier d'impression Repromail 2016 1 Table of contents RÉSUMÉ .................................................................................................................. -

Language Impairment Resulting from a De Novo Deletion of 7Q32.1-Q33: a Case
bioRxiv preprint doi: https://doi.org/10.1101/047241; this version posted April 6, 2016. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. 1 Language impairment resulting from a de novo deletion of 7q32.1-q33: a case report Mª Salud Jiménez-Romero, Montserrat Barcos-Martínez, Isabel Espejo-Portero, Antonio Benítez-Burraco Corresponding author: Antonio Benítez-Burraco, PhD. Department of Spanish Philology and its Didactics, University of Huelva, Huelva, Spain e-mail: [email protected] Affiliations for the other authors: Mª Salud Jiménez-Romero: Department of Psychology, University of Córdoba, Córdoba, Spain & Maimónides Institute of Biomedical Research, Córdoba, Spain Montserrat Barcos-Martínez: Laboratory of Molecular Genetics, University Hospital “Reina Sofía”, Córdoba, Spain &Maimónides Institute of Biomedical Research, Córdoba, Spain Isabel Espejo-Portero: Laboratory of Molecular Genetics, University Hospital “Reina Sofía”, Córdoba, Spain & Maimónides Institute of Biomedical Research, Córdoba, Spain bioRxiv preprint doi: https://doi.org/10.1101/047241; this version posted April 6, 2016. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. 2 ABSTRACT Chromosome 7 is a hot spot for cognitive disorders involving language deficits. We report on a girl who presents with a cognitive and speech delay, motor problems, hearing loss, and behavioral disturbances, and a de novo deletion within 7q32.1-q33 (chromosome position: chr7:127109685-132492196, hg 18). Several genes involved in brain development and function are located within the deleted region.