Understanding Protein Dynamics with L1-Regularized Reversible Hidden Markov Models
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

A Network of Molecular Switches Controls the Activation of the Two-Component Response Regulator Ntrc
ARTICLE Received 8 Aug 2014 | Accepted 26 Apr 2015 | Published 15 Jun 2015 DOI: 10.1038/ncomms8283 A network of molecular switches controls the activation of the two-component response regulator NtrC Dan K. Vanatta1, Diwakar Shukla1,2,3, Morgan Lawrenz1 & Vijay S. Pande1,2 Recent successes in simulating protein structure and folding dynamics have demonstrated the power of molecular dynamics to predict the long timescale behaviour of proteins. Here, we extend and improve these methods to predict molecular switches that characterize conformational change pathways between the active and inactive state of nitrogen regulatory protein C (NtrC). By employing unbiased Markov state model-based molecular dynamics simulations, we construct a dynamic picture of the activation pathways of this key bacterial signalling protein that is consistent with experimental observations and predicts new mutants that could be used for validation of the mechanism. Moreover, these results suggest a novel mechanistic paradigm for conformational switching. 1 Department of Chemistry, Stanford University, Stanford, California 94305, USA. 2 SIMBIOS NIH Center for Biomedical Computation, Stanford University, Stanford, California 94305,USA. 3 Department of Chemical and Bimolecular Engineering, University of Illinois at Urbana-Champaign, Urbana, Illionios 61801, USA. Correspondence and requests for materials should be addressed to V.S.P. (email: [email protected]). NATURE COMMUNICATIONS | 6:7283 | DOI: 10.1038/ncomms8283 | www.nature.com/naturecommunications 1 & 2015 Macmillan Publishers Limited. All rights reserved. ARTICLE NATURE COMMUNICATIONS | DOI: 10.1038/ncomms8283 roteins involved in cellular signalling change their con- Results formation in response to changes in environment (input Simulations reveal molecular switches for NtrC activation. Psignal), such as ligand binding or chemical modification, MSMs for NtrC were built using snapshots along the simulated to control downstream cellular processes (output signal)1. -
Spring 2013 Lecture 23
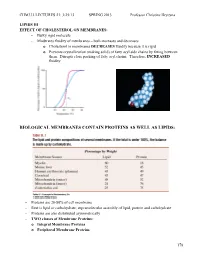
CHM333 LECTURES 23: 3/25/13 SPRING 2013 Professor Christine Hrycyna LIPIDS III EFFECT OF CHOLESTEROL ON MEMBRANES: - Bulky rigid molecule - Moderates fluidity of membranes – both increases and decreases o Cholesterol in membranes DECREASES fluidity because it is rigid o Prevents crystallization (making solid) of fatty acyl side chains by fitting between them. Disrupts close packing of fatty acyl chains. Therefore, INCREASED fluidity BIOLOGICAL MEMBRANES CONTAIN PROTEINS AS WELL AS LIPIDS: - Proteins are 20-80% of cell membrane - Rest is lipid or carbohydrate; supramolecular assembly of lipid, protein and carbohydrate - Proteins are also distributed asymmetrically - TWO classes of Membrane Proteins: o Integral Membrane Proteins o Peripheral Membrane Proteins 178 CHM333 LECTURES 23: 3/25/13 SPRING 2013 Professor Christine Hrycyna - INTEGRAL MEMBRANE PROTEINS o Located WITHIN the lipid bilayer o Usually span the bilayer one or more times – called transmembrane (TM) proteins o Hydrophobic amino acids interact with fatty acid chains in the hydrophobic core of the membrane o Can be removed from the membrane with detergents like SDS – need to disrupt the hydrophobic interactions § Membrane Disruption Animation: o http://www.youtube.com/watch?v=AHT37pvcjc0 o Function: § Transporters – moving molecules into or out of cells or cell membranes § Receptors – transmitting signals from outside of the cell to the inside - β Barrel Integral Membrane Proteins § Barrel-shaped membrane protein that is made up of antiparallel β-strands with hydrophilic (interior) and hydrophobic (facing lipid tails). § So far found only in outer membranes of Gram-negative bacteria, cell wall of Gram-positive bacteria, and outer membranes of mitochondria and chloroplasts. 179 CHM333 LECTURES 23: 3/25/13 SPRING 2013 Professor Christine Hrycyna - α-Helical Membrane Proteins - Can cross the membrane once or many times and have multiple transmembrane segments. -

Site-Specific Tryptophan Labels Reveal Local Microsecond
molecules Article Site-Specific Tryptophan Labels Reveal Local Microsecond–Millisecond Motions of Dihydrofolate Reductase Morgan B. Vaughn 1,*, Chloe Biren 2, Qun Li 2, Ashwin Ragupathi 2 and R. Brian Dyer 2,* 1 Division of Natural Sciences, Oglethorpe University, Atlanta, GA 30319, USA 2 Department of Chemistry, Emory University, Atlanta, GA 30322, USA; [email protected] (C.B.); [email protected] (Q.L.); [email protected] (A.R.) * Correspondence: [email protected] (M.B.V.); [email protected] (R.B.D.); Tel.: +1-404-504-1214 (M.B.V.); +1-404-727-6637 (R.B.D.) Academic Editor: Rafik Karaman Received: 24 July 2020; Accepted: 21 August 2020; Published: 22 August 2020 Abstract: Many enzymes are known to change conformations during their catalytic cycle, but the role of these protein motions is not well understood. Escherichia coli dihydrofolate reductase (DHFR) is a small, flexible enzyme that is often used as a model system for understanding enzyme dynamics. Recently, native tryptophan fluorescence was used as a probe to study micro- to millisecond dynamics of DHFR. Yet, because DHFR has five native tryptophans, the origin of the observed conformational changes could not be assigned to a specific region within the enzyme. Here, we use DHFR mutants, each with a single tryptophan as a probe for temperature jump fluorescence spectroscopy, to further inform our understanding of DHFR dynamics. The equilibrium tryptophan fluorescence of the mutants shows that each tryptophan is in a different environment and that wild-type DHFR fluorescence is not a simple summation of all the individual tryptophan fluorescence signatures due to tryptophan–tryptophan interactions. -

Predicting Protein-Membrane Interfaces of Peripheral Membrane
bioRxiv preprint doi: https://doi.org/10.1101/2021.06.28.450157; this version posted June 29, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. Predicting protein-membrane interfaces of pe- ripheral membrane proteins using ensemble machine learning Alexios Chatzigoulas1,2,* and Zoe Cournia1,* 1Biomedical Research Foundation, Academy of Athens, 4 Soranou Ephessiou, 11527 Athens, Greece, 2Depart- ment of Informatics and Telecommunications, National and Kapodistrian University of Athens, 15784 Athens, Greece *To whom correspondence should be addressed. Abstract Motivation: Abnormal protein-membrane attachment is involved in deregulated cellular pathways and in disease. Therefore, the possibility to modulate protein-membrane interactions represents a new promising therapeutic strategy for peripheral membrane proteins that have been considered so far undruggable. A major obstacle in this drug design strategy is that the membrane binding domains of peripheral membrane proteins are usually not known. The development of fast and efficient algorithms predicting the protein-membrane interface would shed light into the accessibility of membrane-protein interfaces by drug-like molecules. Results: Herein, we describe an ensemble machine learning methodology and algorithm for predicting membrane-penetrating residues. We utilize available experimental data in the literature for training 21 machine learning classifiers and a voting classifier. Evaluation of the ensemble classifier accuracy pro- duced a macro-averaged F1 score = 0.92 and an MCC = 0.84 for predicting correctly membrane-pen- etrating residues on unknown proteins of an independent test set. -

Conformational Proofreading: the Impact of Conformational
Conformational Proofreading: The Impact of Conformational Changes on the Specificity of Molecular Recognition Yonatan Savir1 and Tsvi Tlusty1,* 1Department of Physics of Complex Systems, the Weizmann Institute of Science, Rehovot, Israel, 76100 *Corresponding author. E-mail: [email protected] Abstract To perform recognition, molecules must locate and specifically bind their targets within a noisy biochemical environment with many look-alikes. Molecular recognition processes, especially the induced-fit mechanism, are known to involve conformational changes. This arises a basic question: does molecular recognition gain any advantage by such conformational changes? By introducing a simple statistical-mechanics approach, we study the effect of conformation and flexibility on the quality of recognition processes. Our model relates specificity to the conformation of the participant molecules and thus suggests a possible answer: Optimal specificity is achieved when the ligand is slightly off target, that is a conformational mismatch between the ligand and its main target improves the selectivity of the process. This indicates that deformations upon binding serve as a conformational proofreading mechanism, which may be selected for via evolution. Introduction Practically all biological systems rely on the ability of bio-molecules to specifically recognize each other. Examples are antibodies targeting antigens, regulatory proteins binding DNA and enzymes catalyzing their substrates. These and other molecular recognizers must locate and preferentially interact with their specific targets among a vast variety of molecules that are often structurally similar. This task is further complicated by the inherent noise in the biochemical environment, whose magnitude is comparable with that of the non- covalent binding interactions [1-3]. -
![M.Sc. [Botany] 346 13](https://docslib.b-cdn.net/cover/3507/m-sc-botany-346-13-923507.webp)
M.Sc. [Botany] 346 13
cover page as mentioned below: below: mentioned Youas arepage instructedcover the to updateupdate to the coverinstructed pageare asYou mentioned below: Increase the font size of the Course Name. Name. 1. IncreaseCourse the theof fontsize sizefont ofthe the CourseIncrease 1. Name. use the following as a header in the Cover Page. Page. Cover 2. the usein the followingheader a as as a headerfollowing the inuse the 2. Cover Page. ALAGAPPAUNIVERSITY UNIVERSITYALAGAPPA [Accredited with ’A+’ Grade by NAAC (CGPA:3.64) in the Third Cycle Cycle Third the in (CGPA:3.64) [AccreditedNAAC by withGrade ’A+’’A+’ Gradewith by NAAC[Accredited (CGPA:3.64) in the Third Cycle and Graded as Category–I University by MHRD-UGC] MHRD-UGC] by University and Category–I Graded as as Graded Category–I and University by MHRD-UGC] M.Sc. [Botany] 003 630 – KARAIKUDIKARAIKUDI – 630 003 346 13 EDUCATION DIRECTORATEDISTANCE OF OF DISTANCEDIRECTORATE EDUCATION BIOLOGICAL TECHNIQUES IN BOTANY I - Semester BOTANY IN TECHNIQUES BIOLOGICAL M.Sc. [Botany] 346 13 cover page as mentioned below: below: mentioned Youas arepage instructedcover the to updateupdate to the coverinstructed pageare asYou mentioned below: Increase the font size of the Course Name. Name. 1. IncreaseCourse the theof fontsize sizefont ofthe the CourseIncrease 1. Name. use the following as a header in the Cover Page. Page. Cover 2. the usein the followingheader a as as a headerfollowing the inuse the 2. Cover Page. ALAGAPPAUNIVERSITY UNIVERSITYALAGAPPA [Accredited with ’A+’ Grade by NAAC (CGPA:3.64) in the Third Cycle Cycle Third the in (CGPA:3.64) [AccreditedNAAC by withGrade ’A+’’A+’ Gradewith by NAAC[Accredited (CGPA:3.64) in the Third Cycle and Graded as Category–I University by MHRD-UGC] MHRD-UGC] by University and Category–I Graded as as Graded Category–I and University by MHRD-UGC] M.Sc. -

Cell Signaling: G-Protein Coupled Receptors and the Β
Cell Signaling: G-Protein Coupled Receptors and the β2-Adrenergic Receptor Dianna Amasino, Axel Glaubitz, Susan Huang, Joy Li, Hsien-Yu Shih, Junyao Song, Xiao Zhu Madison West High School, Madison, WI 53726 Advisor: Basudeb Bhattacharyya, University of Wisconsin, Madison, WI 53706 Mentors: Dr. Dave Nelson and Peter Vander Velden, University of Wisconsin, Madison, WI 53706 Introduction Mechanism of the β adrenergic receptor: an example of GPCR signal A conserved structure and mechanism G protein-coupled receptors (GPCRs) are the largest transduction family of integral membrane proteins coded by the human genome. GPCRs are important for signal transduction with the general structural characteristic of a plasma membrane receptor with seven transmembrane segments (Figure 1). One example of a GPCR targeted by pharmaceutical companies is the β2-adrenergic receptor. Adrenergic receptors are found through out the body and are triggered by the hormone epinephrine (also known as adrenaline, hence the name adrenergic). When epinephrine binds to the receptors, it causes a slight conformational change within the receptor. This Cherezov, et al., 2007 change then triggers activation of a G-protein, which induces a response within the cell (for example, muscle contraction). When this signal transduction event Figure 4. Overlay of bovine rhodopsin (cyan) and the functions normally in the body, it helps regulate heart rate human β2-adrenergic receptor (orange) and blood pressure and is important for the “fight or Figure 4 above demonstrates how conserved the structure, flight” response. Beta blockers are medically used to and hence the mechannsm, of signal transduction using bind to adrenergic receptors, manipulating the hormone’s GPCRs really is. -

Membrane Protein Structure Determination and Characterisation by Solution and Solid-State NMR
biology Review Membrane Protein Structure Determination and Characterisation by Solution and Solid-State NMR Vivien Yeh , Alice Goode and Boyan B. Bonev * School of Life Sciences, University of Nottingham, Nottingham NG7 2UH, UK; [email protected] (V.Y.); [email protected] (A.G.) * Correspondence: [email protected] Received: 21 October 2020; Accepted: 11 November 2020; Published: 12 November 2020 Simple Summary: Cells, life’s smallest units, are defined within the enclosure of thin, continuous membranes, which confine the molecular machinery required for the life and replication of cells. Crucially, membranes of cells establish and actively maintain distinctly different environments inside cells, including electrical and solute gradients vital to normal cellular functions. Membrane proteins are in charge of transport, electrical polarisation, signalling, membrane remodelling and other important functions. As such, membrane proteins are key drug targets and understanding their structure and function is essential to drug development and cellular control. Membrane proteins have physical characteristics that make such studies very challenging. Nuclear magnetic resonance is one advanced tool that enables structural studies of membrane proteins and their interactions at the atomic level of detail. We discuss the applications of NMR in solution and solid state to membrane protein studies alongside new developments in signal and sensitivity enhancement through dynamic nuclear polarisation. Abstract: Biological membranes define the interface of life and its basic unit, the cell. Membrane proteins play key roles in membrane functions, yet their structure and mechanisms remain poorly understood. Breakthroughs in crystallography and electron microscopy have invigorated structural analysis while failing to characterise key functional interactions with lipids, small molecules and membrane modulators, as well as their conformational polymorphism and dynamics. -

Controlling Entropy to Tune the Functions of Intrinsically Disordered
Available online at www.sciencedirect.com ScienceDirect Controlling entropy to tune the functions of intrinsically disordered regions 1 1 Tilman Flock , Robert J Weatheritt , Natasha S Latysheva and M Madan Babu Intrinsically disordered regions (IDRs) are fundamental units of extensive functional regions within polypeptide seg- protein function and regulation. Despite their inability to form a ments that do not form stable tertiary structure. These unique stable tertiary structure in isolation, many IDRs adopt a regions, referred to as intrinsically disordered regions defined conformation upon binding and achieve their function (IDRs), exist in over 35% of human proteins [2,3] func- through their interactions with other biomolecules. However, this tioning in all major cellular processes with significant requirement for IDR functionality seems to be at odds with the enrichment in signalling and regulation [4,5 ]. Although high entropic cost they must incur upon binding an interaction IDRs are capable of performing functions comparable to partner. How is this seeming paradox resolved? While increasing structured regions, they vary significantly in their amino the enthalpy of binding is one approach to compensate for this acid composition [6] and their biophysical properties entropic cost, growing evidence suggests that inherent features [7 ]. Together this suggests that IDRs are independent of IDRs, for instance repeating linear motifs, minimise the units of protein function, capable of achieving biological entropic cost of binding. Moreover, this control of entropic cost function in a manner that is distinct from that of struc- can be carefully modulated by a range of regulatory tured regions [1 ,7 ]. mechanisms, such as alternative splicing and post-translational modifications, which enable allosteric communication and Energy landscape of intrinsically disordered rheostat-like tuning of IDR function. -

Biomodel Engineering for Multiscale Systems Biology
BioModel Engineering for Multiscale Systems Biology Monika Heinera, David Gilbertb aDepartment of Computer Science, Brandenburg University of Technology Postbox 10 13 44, 03013 Cottbus, Germany [email protected] bSchool of Information Systems, Computing and Mathematics Brunel University, Uxbridge, Middlesex UB8 3PH, UK [email protected] Abstract We discuss some motivational challenges arising from the need to model and analyse complex biological systems at multiple scales (spatial and temporal), and present a biomodel engineering framework to address some of these issues within the context of multiscale Systems Biology. Our methodology is based on a structured family of Petri net classes which enables the investigation of a given system using various modelling abstractions: qualitative, stochastic, continuous and hybrid, optionally in a spatial context. We illustrate our approach with case studies demonstrating hierarchical flattening, treatment of space, and hierarchical organisation of space. Keywords: BioModel Engineering; multiscale modelling; Systems Biology; Synthetic Biology; biomolecular networks; coloured qualitative/stochastic/continuous/hybrid Petri nets; model checking. 1. Motivation BioModel Engineering. Biology is increasingly becoming an informa- tional science. This revolution has been driven by technological advances which have supported the development of studies at many levels of intra- and intercellular activity. These advances have facilitated the analysis of how the components of a biological -

Conformational Changes in Ligand Binding Processes
Conformational Changes in Ligand Binding Processes Conformational Changes in Ligand Binding Processes Dissertation for the award of the degree Doctor rerum naturalium of the Georg-August-Universit¨atG¨ottingen within the doctoral program IMPRS-PBCS of the Georg-August University School of Science (GAUSS) Submitted by B´elaVoß from Hamburg G¨ottingen,2014 • Thesis Committee: { Prof. Dr. Helmut Grubm¨uller Department for Theoretical and Computational Biophysics Max-Planck-Institute for Biophysical Chemistry { Prof. Dr. J¨orgEnderlein Third Institute of Physics Faculty of Physics Georg-August-Universit¨atG¨ottingen { Prof. Dr. Marina Benatti Electron Spin Resonance Spectroscopy Group Max-Planck-Institute for Biophysical Chemistry • Examination Board: { Prof. Dr. Helmut Grubm¨uller(supervisor, reviewer) Department for Theoretical and Computational Biophysics Max-Planck-Institute for Biophysical Chemistry { Prof. Dr. J¨orgEnderlein (reviewer) Third Institute of Physics Faculty of Physics Georg-August-Universit¨atG¨ottingen { Prof. Dr. Marina Benatti Electron Spin Resonance Spectroscopy Group Max-Planck-Institute for Biophysical Chemistry { Prof. Dr. Sarah K¨oster CRC Research Group \Nanoscale Imaging of Cellular Dynamics" Faculty of Physics Georg-August-Universit¨atG¨ottingen { Dr. Thomas Burg Biological Micro- and Nanotechnology Group Max-Planck-Institute for Biophysical Chemistry { Prof. Dr. Markus M¨uller Institute for Theoretical Physics Faculty of Physics Georg-August-Universit¨atG¨ottingen Date of the disputation: 2015-01-30 iv Hiermit erkl¨areich, dass ich die vorliegende Arbeit selbstst¨andigverfasst und keine anderen als die angegebenen Quellen und Hilfsmittel verwendet habe. G¨ottingen,den 26.11.2014 B´elaVoß v Contents List of Abbreviations xi I. Opening 1 1. Introduction 3 2. Theoretical Background 5 2.1. -

Induced Fit, Conformational Selection and Independent Dynamic Segments: an Extended View of Binding Events
Opinion Cover story in Trends in Biochemical Sciences 35, 539-546 (2010) Induced fit, conformational selection and independent dynamic segments: an extended view of binding events Peter Csermely1,a, Robin Palotai1 and Ruth Nussinov2,3 1Department of Medical Chemistry, Semmelweis University, P O Box 260., H-1444 Budapest 8, Hungary 2Center for Cancer Research Nanobiology Program, SAIC-Frederick, Inc., NCI-Frederick, Frederick, MD 21702, USA 3Sackler Institute of Molecular Medicine, Department of Human Genetics and Molecular Medicine, Sackler School of Medicine, Tel Aviv University, Tel Aviv 69978, Israel Single molecule and NMR measurements of protein dynamics increasingly uncover the complexity of binding scenarios. Here we describe an extended conformational selection model which embraces a repertoire of selection and adjustment processes. Induced fit can be viewed as a subset of this repertoire, whose contribution is affected by the bond- types stabilizing the interaction and the differences between the interacting partners. We argue that protein segments whose dynamics are distinct from the rest of the protein (‘discrete breathers’) can govern conformational transitions and allosteric propagation that accompany binding processes, and as such may be more sensitive to mutational events. Additionally, we highlight the dynamic complexity of binding scenarios as they relate to events such as aggregation and signalling, and the crowded cellular environment. The induced fit and the original conformational selection models The prevailing view of binding mechanisms has evolved from the early ‘lock-and-key’ hypothesis [1] to the now popular ‘induced fit’ model [2]. According to the induced fit scenario, the interaction between a protein and a rigid binding partner induces a conformational change in the protein.