UNIT – I - Fundamentals of Genomics and Proteomics– SBI1309
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Developing Back Reflectance Absorbance As a Useful Technique
Design of a Simple Cryogenic System for Ultraviolet-Visible Absorption Spectroscopy with a Back-reflectance Fiber Optic Probe Andrew Vinyard, Kaj Hansen, Ross Byrd, Douglas A Stuart * and John Hansen * Department of Chemistry, University of West Georgia *Corresponding Authors, email: [email protected] and [email protected] Abstract We report a convenient and inexpensive technique for the rapid acquisition of absorption spectra from small samples at cryogenic temperatures using a home built cryostat with novel collection optics. A cylindrical copper block was constructed with a coaxial bore to hold a 4.00 mm diameter EPR tube and mounted on a copper feed in thermal contact with liquid nitrogen. A 6.35 mm diameter hole was bored into the side of the cylinder so a fiber optic cable bundle could be positioned orthogonally to the EPR tube. The light passing through the sample is reflected off of the opposing surfaces of the EPR tube and surrounding copper, back through the sample. The emergent light is then collected by the fiber optic bundle, and analyzed by a dispersive spectrometer. Absorption spectra for KMnO4 were measured between 400 nm and 700 nm. Absorption intensity at 506 nm, 525 nm, 545 nm and 567 nm was found to be proportional to concentration, displaying Beer’s law like behavior. The EPR tube had an internal diameter of 3.2 mm; the double pass of the probe beam through the sample affords a central path length of about 6.4 mm. Comparing these measurements with those recorded on a conventional tabletop spectrometer using a cuvette with a 10.00 mm path length, we consistently found a ratio between intensities of 0.58 rather than the anticipated 0.64. -

A Network of Molecular Switches Controls the Activation of the Two-Component Response Regulator Ntrc
ARTICLE Received 8 Aug 2014 | Accepted 26 Apr 2015 | Published 15 Jun 2015 DOI: 10.1038/ncomms8283 A network of molecular switches controls the activation of the two-component response regulator NtrC Dan K. Vanatta1, Diwakar Shukla1,2,3, Morgan Lawrenz1 & Vijay S. Pande1,2 Recent successes in simulating protein structure and folding dynamics have demonstrated the power of molecular dynamics to predict the long timescale behaviour of proteins. Here, we extend and improve these methods to predict molecular switches that characterize conformational change pathways between the active and inactive state of nitrogen regulatory protein C (NtrC). By employing unbiased Markov state model-based molecular dynamics simulations, we construct a dynamic picture of the activation pathways of this key bacterial signalling protein that is consistent with experimental observations and predicts new mutants that could be used for validation of the mechanism. Moreover, these results suggest a novel mechanistic paradigm for conformational switching. 1 Department of Chemistry, Stanford University, Stanford, California 94305, USA. 2 SIMBIOS NIH Center for Biomedical Computation, Stanford University, Stanford, California 94305,USA. 3 Department of Chemical and Bimolecular Engineering, University of Illinois at Urbana-Champaign, Urbana, Illionios 61801, USA. Correspondence and requests for materials should be addressed to V.S.P. (email: [email protected]). NATURE COMMUNICATIONS | 6:7283 | DOI: 10.1038/ncomms8283 | www.nature.com/naturecommunications 1 & 2015 Macmillan Publishers Limited. All rights reserved. ARTICLE NATURE COMMUNICATIONS | DOI: 10.1038/ncomms8283 roteins involved in cellular signalling change their con- Results formation in response to changes in environment (input Simulations reveal molecular switches for NtrC activation. Psignal), such as ligand binding or chemical modification, MSMs for NtrC were built using snapshots along the simulated to control downstream cellular processes (output signal)1. -
Spring 2013 Lecture 23
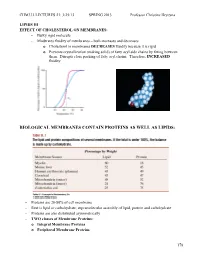
CHM333 LECTURES 23: 3/25/13 SPRING 2013 Professor Christine Hrycyna LIPIDS III EFFECT OF CHOLESTEROL ON MEMBRANES: - Bulky rigid molecule - Moderates fluidity of membranes – both increases and decreases o Cholesterol in membranes DECREASES fluidity because it is rigid o Prevents crystallization (making solid) of fatty acyl side chains by fitting between them. Disrupts close packing of fatty acyl chains. Therefore, INCREASED fluidity BIOLOGICAL MEMBRANES CONTAIN PROTEINS AS WELL AS LIPIDS: - Proteins are 20-80% of cell membrane - Rest is lipid or carbohydrate; supramolecular assembly of lipid, protein and carbohydrate - Proteins are also distributed asymmetrically - TWO classes of Membrane Proteins: o Integral Membrane Proteins o Peripheral Membrane Proteins 178 CHM333 LECTURES 23: 3/25/13 SPRING 2013 Professor Christine Hrycyna - INTEGRAL MEMBRANE PROTEINS o Located WITHIN the lipid bilayer o Usually span the bilayer one or more times – called transmembrane (TM) proteins o Hydrophobic amino acids interact with fatty acid chains in the hydrophobic core of the membrane o Can be removed from the membrane with detergents like SDS – need to disrupt the hydrophobic interactions § Membrane Disruption Animation: o http://www.youtube.com/watch?v=AHT37pvcjc0 o Function: § Transporters – moving molecules into or out of cells or cell membranes § Receptors – transmitting signals from outside of the cell to the inside - β Barrel Integral Membrane Proteins § Barrel-shaped membrane protein that is made up of antiparallel β-strands with hydrophilic (interior) and hydrophobic (facing lipid tails). § So far found only in outer membranes of Gram-negative bacteria, cell wall of Gram-positive bacteria, and outer membranes of mitochondria and chloroplasts. 179 CHM333 LECTURES 23: 3/25/13 SPRING 2013 Professor Christine Hrycyna - α-Helical Membrane Proteins - Can cross the membrane once or many times and have multiple transmembrane segments. -

2 in 1 Fluorescence and Absorbance Spectrometer
2 in 1 Fluorescence and Application Fluorescence Absorbance Spectrometer: Note How does it work? Life Sciences Figure 1: Duetta (left) and the inside of the Duetta sample compartment (right) with direction of the light path Introduction Duetta™ is a 2-in-1 fluorescence and absorbance excitation energy and the spectrum at longer wavelengths spectrometer from HORIBA Scientific. There are several is acquired to measure the distribution of intensity and benefits of having two different spectroscopies in one energy (wavelength) of the photons emitted. instrument. For high concentration solutions of interest, both primary and secondary inner-filter effects will affect Fluorescence Excitation Spectrum the fluorescence spectrum measured on a standard Acquiring the intensity of photons emitted at a single fluorometer. Having absorbance and fluorescence emission wavelength and scanning the excitation spectroscopy on the same instrument enables Duetta monochromator to excite the population of molecules in and EzSpec software to apply corrections for inner-filter the sample with different wavelengths. A fluorescence effect and provide more accurate data, for a wider range of excitation spectrum is analogous to an absorbance sample concentrations. Another benefit of the two in one spectrum, but is specific to a single emitting species/ instrument is that absorbance results and fluorescence wavelength as opposed to collecting all absorbing species results contain less error since the sample does not move in a sample or solution. from one instrument to another to get both measurements. Standard fluorescence methods such as Emission %Transmittance Spectrum Spectrum and Excitation Spectrum are available on Duetta, This is a ratio, in terms of percentage, of the intensity of but EzSpec software gives a user the ability to measure transmitted light through an absorbing sample (I) compared the Absorbance Spectrum as a stand-alone method or to the transmitted light through a blank solvent (I0). -

Analysis of Proteins by Immunoprecipitation
Laboratory Procedures, PJ Hansen Laboratory - University of Florida Analysis of Proteins by Immunoprecipitation P.J. Hansen1 1Dept. of Animal Sciences, University of Florida Introduction Immunoprecipitation is a procedure by which peptides or proteins that react specifically with an antibody are removed from solution and examined for quantity or physical characteristics (molecular weight, isoelectric point, etc.). As usually practiced, the name of the procedure is a misnomer since removal of the antigen from solution does not depend upon the formation of an insoluble antibody-antigen complex. Rather, antibody-antigen complexes are removed from solution by addition of an insoluble form of an antibody binding protein such as Protein A, Protein G or second antibody (Figure 1). Thus, unlike other techniques based on immunoprecipitation, it is not necessary to determine the optimal antibody dilution that favors spontaneously-occurring immunoprecipitates. Figure 1. Schematic representation of the principle of immunoprecipitation. An antibody added to a mixture of radiolabeled (*) and unlabeled proteins binds specifically to its antigen (A) (left tube). Antibody- antigen complex is absorbed from solution through the addition of an immobilized antibody binding protein such as Protein A-Sepharose beads (middle panel). Upon centrifugation, the antibody-antigen complex is brought down in the pellet (right panel). Subsequent liberation of the antigen can be achieved by boiling the sample in the presence of SDS. Typically, the antigen is made radioactive before the immunoprecipitation procedure, either by culturing cells with radioactive precursor or by labeling the molecule after synthesis has been completed (e.g., by radioiodination to iodinate tyrosine residues or by sodium [3H]borohydride reduction to label carbohydrate). -

1 Supplementary Information in Vivo Mrna Display Enables
Supplementary Information In vivo mRNA display enables Large-scale Proteomics by Next Generation Sequencing P. Oikonomou1,2,3, R. Salatino2, S. Tavazoie1,2,3* 1Department of Biological Sciences, Columbia University, New York City, New York, United States 2Department of Systems Biology, Columbia University, New York City, New York, United States 3Department of Biochemistry and Molecular Biophysics, Columbia University, New York City, New York, United States * Correspondence: P.O.: [email protected]; S.T.: [email protected] Table of Contents Methods ..................................................................................................................................... 4 Plasmid Construction .............................................................................................................. 4 Yeast Strains .......................................................................................................................... 4 In vivo mRNA display Library Generation ............................................................................... 5 Yeast cell culture .................................................................................................................... 5 Excess Coat Protein ............................................................................................................... 6 Non-Specific Functional Controls for in vivo mRNA display .................................................... 6 Whole cell lysate preparation ................................................................................................. -

Using Peptide-Phage Display to Capture Conditional Motif-Based Interactions
Digital Comprehensive Summaries of Uppsala Dissertations from the Faculty of Science and Technology 1716 Using peptide-phage display to capture conditional motif-based interactions GUSTAV SUNDELL ACTA UNIVERSITATIS UPSALIENSIS ISSN 1651-6214 ISBN 978-91-513-0433-5 UPPSALA urn:nbn:se:uu:diva-359434 2018 Dissertation presented at Uppsala University to be publicly examined in B42, BMC, Husargatan 3, Uppsala, Friday, 19 October 2018 at 09:15 for the degree of Doctor of Philosophy. The examination will be conducted in English. Faculty examiner: Doctor Attila Reményi (nstitute of Enzymology, Research Center for Natural Sciences, Hungarian Academy of Sciences, Budapest, Hungary). Abstract Sundell, G. 2018. Using peptide-phage display to capture conditional motif-based interactions. Digital Comprehensive Summaries of Uppsala Dissertations from the Faculty of Science and Technology 1716. 87 pp. Uppsala: Acta Universitatis Upsaliensis. ISBN 978-91-513-0433-5. This thesis explores the world of conditional protein-protein interactions using combinatorial peptide-phage display and proteomic peptide-phage display (ProP-PD). Large parts of proteins in the human proteome do not fold in to well-defined structures instead they are intrinsically disordered. The disordered parts are enriched in linear binding-motifs that participate in protein-protein interaction. These motifs are 3-12 residue long stretches of proteins where post-translational modifications, like protein phosphorylation, can occur changing the binding preference of the motif. Allosteric changes in a protein or domain due to phosphorylation or binding to second messenger molecules like Ca2+ can also lead conditional interactions. Finding phosphorylation regulated motif-based interactions on a proteome-wide scale has been a challenge for the scientific community. -

The Immunoassay Guide to Successful Mass Spectrometry
The Immunoassay Guide to Successful Mass Spectrometry Orr Sharpe Robinson Lab SUMS User Meeting October 29, 2013 What is it ? Hey! Look at that! Something is reacting in here! I just wish I knew what it is! anti-phospho-Tyrosine Maybe we should mass spec it! Coffey GP et.al. 2009 JCS 22(3137-44) True or false 1. A big western blot band means I have a LOT of protein 2. One band = 1 protein Big band on Western blot Bands are affected mainly by: Antibody affinity to the antigen Number of available epitopes Remember: After the Ag-Ab interaction, you are amplifying the signal by using an enzyme linked to a secondary antibody. How many proteins are in a band? Human genome: 20,000 genes=100,000 proteins There are about 5000 different proteins, not including PTMs, in a given cell at a single time point. Huge dynamic range 2D-PAGE: about 1000 spots are visible. 1D-PAGE: about 60 -100 bands are visible - So, how many proteins are in my band? Separation is the key! Can you IP your protein of interest? Can you find other way to help with the separation? -Organelle enrichment -PTMs enrichment -Size enrichment Have you optimized your running conditions? Choose the right gel and the right running conditions! Immunoprecipitation, in theory Step 1: Create a complex between a desired protein (Antigen) and an Antibody Step 2: Pull down the complex and remove the unbound proteins Step 3: Elute your antigen and analyze Immunoprecipitation, in real life Flow through Wash Elution M 170kDa 130kDa 100kDa 70kDa 55kDa 40kDa 35kDa 25kDa Lung tissue lysate, IP with patient sera , Coomassie stain Rabinovitch and Robinson labs, unpublished data Optimizing immunoprecipitation You need: A good antibody that can IP The right beads: i. -

Yeast Surface 2-Hybrid to Detect Protein-Protein Interactions Via the Secretory Pathway As a Platform for Antibody Discovery
CORE Metadata, citation and similar papers at core.ac.uk Provided by Nature Precedings Yeast surface 2-hybrid to detect protein-protein interactions via the secretory pathway as a platform for antibody discovery Xuebo Hu1, Sungkwon Kang1, Xiaoyue Chen1, Charles B Shoemaker2, Moonsoo M Jin1 1Department of Biomedical Engineering, Cornell University, Hungerford Hill Road, Ithaca, NY 14853 2Department of Biomedical Sciences, Tufts Cummings School of Veterinary Medicine, 200 Westboro Rd, North Grafton, MA 01536 Correspondence should be addressed to M.M.J. ([email protected]). Abstract: High throughput methods to measure protein-protein interactions will facilitate uncovering pairs of unknown interactions as well as designing new interactions. We have developed a platform to detect protein interactions on the surface of yeast, where one protein (bait) is covalently anchored to the cell wall and the other (prey) is expressed in secretory form. The prey is released either outside of the cells or remains on the cell surface by its binding to the bait. The strength of their interaction is measured by antibody binding to the epitope tag fused to the prey or direct readout of split fluorescence protein complementation. Our novel ‘yeast surface 2-hybrid’ system was found to differentiate 6-log difference in binding affinities between coiled coil associations and to measure specific interactions of antibodies and antigens. We demonstrate the use of YS2H in exploring activation allostery in integrins and isolating heavy chain only antibodies against botulinum neurotoxin. Introduction Protein-protein interactions are essential to virtually every cellular process and their understanding is of great interest in basic science as well as in the development of effective therapeutics. -

WO 2010/129541 Al
(12) INTERNATIONAL APPLICATION PUBLISHED UNDER THE PATENT COOPERATION TREATY (PCT) (19) World Intellectual Property Organization International Bureau (10) International Publication Number (43) International Publication Date 11 November 2010 (11.11.2010) WO 2010/129541 Al (51) International Patent Classification: (81) Designated States (unless otherwise indicated, for every GOlN 33/53 (2006.01) C07K 16/00 (2006.01) kind of national protection available): AE, AG, AL, AM, AO, AT, AU, AZ, BA, BB, BG, BH, BR, BW, BY, BZ, (21) International Application Number: CA, CH, CL, CN, CO, CR, CU, CZ, DE, DK, DM, DO, PCT/US20 10/033537 DZ, EC, EE, EG, ES, FI, GB, GD, GE, GH, GM, GT, (22) International Filing Date: HN, HR, HU, ID, IL, IN, IS, JP, KE, KG, KM, KN, KP, 4 May 2010 (04.05.2010) KR, KZ, LA, LC, LK, LR, LS, LT, LU, LY, MA, MD, ME, MG, MK, MN, MW, MX, MY, MZ, NA, NG, NI, (25) Filing Language: English NO, NZ, OM, PE, PG, PH, PL, PT, RO, RS, RU, SC, SD, (26) Publication Language: English SE, SG, SK, SL, SM, ST, SV, SY, TH, TJ, TM, TN, TR, TT, TZ, UA, UG, US, UZ, VC, VN, ZA, ZM, ZW. (30) Priority Data: 61/175,076 4 May 2009 (04.05.2009) US (84) Designated States (unless otherwise indicated, for every 61/305,991 19 February 2010 (19.02.2010) US kind of regional protection available): ARIPO (BW, GH, GM, KE, LR, LS, MW, MZ, NA, SD, SL, SZ, TZ, UG, (71) Applicant (for all designated States except US): AF- ZM, ZW), Eurasian (AM, AZ, BY, KG, KZ, MD, RU, TJ, FOMIX CORPORATION [US/US]; 688 E. -

Optical and Electron Paramagnetic Resonance Characterization of Point Defects in Semiconductors
Air Force Institute of Technology AFIT Scholar Theses and Dissertations Student Graduate Works 3-1-2019 Optical and Electron Paramagnetic Resonance Characterization of Point Defects in Semiconductors Elizabeth M. Scherrer Follow this and additional works at: https://scholar.afit.edu/etd Part of the Electromagnetics and Photonics Commons Recommended Citation Scherrer, Elizabeth M., "Optical and Electron Paramagnetic Resonance Characterization of Point Defects in Semiconductors" (2019). Theses and Dissertations. 2463. https://scholar.afit.edu/etd/2463 This Dissertation is brought to you for free and open access by the Student Graduate Works at AFIT Scholar. It has been accepted for inclusion in Theses and Dissertations by an authorized administrator of AFIT Scholar. For more information, please contact [email protected]. OPTICAL AND ELECTRON PARAMAGNETIC RESONANCE CHARACTERIZATION OF POINT DEFECTS IN SEMICONDUCTORS DISSERTATION Elizabeth M. Scherrer, Captain, USAF AFIT-ENP-DS-19-M-091 DEPARTMENT OF THE AIR FORCE AIR UNIVERSITY AIR FORCE INSTITUTE OF TECHNOLOGY Wright-Patterson Air Force Base, Ohio DISTRIBUTION STATEMENT A APPROVED FOR PUBLIC RELEASE; DISTRIBUTION UNLIMITED. The views expressed in this thesis are those of the author and do not reflect the official policy or position of the United States Air Force, Department of Defense, or the United States Government. This material is declared a work of the U.S. Government and is not subject to copyright protection in the United States. AFIT-ENP-DS-19-M-091 OPTICAL AND ELECTRON PARAMAGNETIC RESONANCE CHARACTERIZATION OF POINT DEFECTS IN SEMICONDUCTORS DISSERTATION Presented to the Faculty Department of Engineering Physics Graduate School of Engineering and Management Air Force Institute of Technology Air University Air Education and Training Command In Partial Fulfillment of the Requirements for the Doctor of Philosophy Degree Elizabeth M. -

Development of a Phage Display Library for Discovery of Antigenic Brucella Peptides Jeffrey Williams Iowa State University
Iowa State University Capstones, Theses and Graduate Theses and Dissertations Dissertations 2018 Development of a phage display library for discovery of antigenic Brucella peptides Jeffrey Williams Iowa State University Follow this and additional works at: https://lib.dr.iastate.edu/etd Part of the Microbiology Commons Recommended Citation Williams, Jeffrey, "Development of a phage display library for discovery of antigenic Brucella peptides" (2018). Graduate Theses and Dissertations. 16896. https://lib.dr.iastate.edu/etd/16896 This Thesis is brought to you for free and open access by the Iowa State University Capstones, Theses and Dissertations at Iowa State University Digital Repository. It has been accepted for inclusion in Graduate Theses and Dissertations by an authorized administrator of Iowa State University Digital Repository. For more information, please contact [email protected]. Development of a phage display library for discovery of antigenic Brucella peptides by Jeffrey Williams A thesis submitted to the graduate faculty in partial fulfillment of the requirements for the degree of MASTER OF SCIENCE Major: Microbiology Program of Study Committee: Bryan H. Bellaire, Major Professor Steven Olsen Steven Carlson The student author, whose presentation of the scholarship herein was approved by the program of study committee, is solely responsible for the content of this thesis. The Graduate College will ensure this thesis is globally accessible and will not permit alterations after a degree is conferred. Iowa State University