Ligand-Dependent Conformations and Dynamics of the Serotonin 5-HT2A Receptor Determine Its Activation and Membrane-Driven Oligomerization Properties
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Peropsin, a Novel Visual Pigment-Like Protein Located in the Apical Microvilli of the Retinal Pigment Epithelium
Proc. Natl. Acad. Sci. USA Vol. 94, pp. 9893–9898, September 1997 Neurobiology Peropsin, a novel visual pigment-like protein located in the apical microvilli of the retinal pigment epithelium HUI SUN*, DEBRA J. GILBERT†,NEAL G. COPELAND†,NANCY A. JENKINS†, AND JEREMY NATHANS*‡§¶i *Department of Molecular Biology and Genetics, §Department of Neuroscience, ¶Department of Ophthalmology, ‡Howard Hughes Medical Institute, Johns Hopkins University School of Medicine, Baltimore, MD 21205; and †Mammalian Genetics Laboratory, Advanced BioScience Laboratories Basic Research Program, National Cancer Institute–Frederick Cancer Research and Development Center, Frederick, MD 21702 Contributed by Jeremy Nathans, June 19, 1997 ABSTRACT A visual pigment-like protein, referred to as bovine RPE binds to all-trans but not 11-cis retinal and absorbs peropsin, has been identified by large-scale sequencing of both visible and ultraviolet light (8, 9). The sequences of cDNAs derived from human ocular tissues. The corresponding retinochrome and RGR opsin form a distinct and highly mRNA was found only in the eye, where it is localized to the divergent branch within the visual pigment family (6, 10). retinal pigment epithelium (RPE). Peropsin immunoreactiv- Whether retinochrome and RGR act as signal-transducing ity, visualized by light and electron microscopy, localizes the light receptors, participate in the visual cycle as retinal isomer- protein to the apical face of the RPE, and most prominently ases, or function in both capacities, is not known. to the microvilli that surround the photoreceptor outer seg- In the vertebrate eye, the RPE lies adjacent to the photo- ments. These observations suggest that peropsin may play a receptor cells and performs a number of functions critical for role in RPE physiology either by detecting light directly or by the viability and activity of the retina (11). -

A Network of Molecular Switches Controls the Activation of the Two-Component Response Regulator Ntrc
ARTICLE Received 8 Aug 2014 | Accepted 26 Apr 2015 | Published 15 Jun 2015 DOI: 10.1038/ncomms8283 A network of molecular switches controls the activation of the two-component response regulator NtrC Dan K. Vanatta1, Diwakar Shukla1,2,3, Morgan Lawrenz1 & Vijay S. Pande1,2 Recent successes in simulating protein structure and folding dynamics have demonstrated the power of molecular dynamics to predict the long timescale behaviour of proteins. Here, we extend and improve these methods to predict molecular switches that characterize conformational change pathways between the active and inactive state of nitrogen regulatory protein C (NtrC). By employing unbiased Markov state model-based molecular dynamics simulations, we construct a dynamic picture of the activation pathways of this key bacterial signalling protein that is consistent with experimental observations and predicts new mutants that could be used for validation of the mechanism. Moreover, these results suggest a novel mechanistic paradigm for conformational switching. 1 Department of Chemistry, Stanford University, Stanford, California 94305, USA. 2 SIMBIOS NIH Center for Biomedical Computation, Stanford University, Stanford, California 94305,USA. 3 Department of Chemical and Bimolecular Engineering, University of Illinois at Urbana-Champaign, Urbana, Illionios 61801, USA. Correspondence and requests for materials should be addressed to V.S.P. (email: [email protected]). NATURE COMMUNICATIONS | 6:7283 | DOI: 10.1038/ncomms8283 | www.nature.com/naturecommunications 1 & 2015 Macmillan Publishers Limited. All rights reserved. ARTICLE NATURE COMMUNICATIONS | DOI: 10.1038/ncomms8283 roteins involved in cellular signalling change their con- Results formation in response to changes in environment (input Simulations reveal molecular switches for NtrC activation. Psignal), such as ligand binding or chemical modification, MSMs for NtrC were built using snapshots along the simulated to control downstream cellular processes (output signal)1. -
Spring 2013 Lecture 23
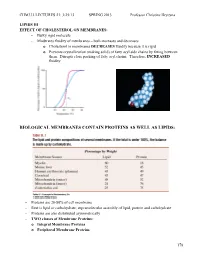
CHM333 LECTURES 23: 3/25/13 SPRING 2013 Professor Christine Hrycyna LIPIDS III EFFECT OF CHOLESTEROL ON MEMBRANES: - Bulky rigid molecule - Moderates fluidity of membranes – both increases and decreases o Cholesterol in membranes DECREASES fluidity because it is rigid o Prevents crystallization (making solid) of fatty acyl side chains by fitting between them. Disrupts close packing of fatty acyl chains. Therefore, INCREASED fluidity BIOLOGICAL MEMBRANES CONTAIN PROTEINS AS WELL AS LIPIDS: - Proteins are 20-80% of cell membrane - Rest is lipid or carbohydrate; supramolecular assembly of lipid, protein and carbohydrate - Proteins are also distributed asymmetrically - TWO classes of Membrane Proteins: o Integral Membrane Proteins o Peripheral Membrane Proteins 178 CHM333 LECTURES 23: 3/25/13 SPRING 2013 Professor Christine Hrycyna - INTEGRAL MEMBRANE PROTEINS o Located WITHIN the lipid bilayer o Usually span the bilayer one or more times – called transmembrane (TM) proteins o Hydrophobic amino acids interact with fatty acid chains in the hydrophobic core of the membrane o Can be removed from the membrane with detergents like SDS – need to disrupt the hydrophobic interactions § Membrane Disruption Animation: o http://www.youtube.com/watch?v=AHT37pvcjc0 o Function: § Transporters – moving molecules into or out of cells or cell membranes § Receptors – transmitting signals from outside of the cell to the inside - β Barrel Integral Membrane Proteins § Barrel-shaped membrane protein that is made up of antiparallel β-strands with hydrophilic (interior) and hydrophobic (facing lipid tails). § So far found only in outer membranes of Gram-negative bacteria, cell wall of Gram-positive bacteria, and outer membranes of mitochondria and chloroplasts. 179 CHM333 LECTURES 23: 3/25/13 SPRING 2013 Professor Christine Hrycyna - α-Helical Membrane Proteins - Can cross the membrane once or many times and have multiple transmembrane segments. -

Site-Specific Tryptophan Labels Reveal Local Microsecond
molecules Article Site-Specific Tryptophan Labels Reveal Local Microsecond–Millisecond Motions of Dihydrofolate Reductase Morgan B. Vaughn 1,*, Chloe Biren 2, Qun Li 2, Ashwin Ragupathi 2 and R. Brian Dyer 2,* 1 Division of Natural Sciences, Oglethorpe University, Atlanta, GA 30319, USA 2 Department of Chemistry, Emory University, Atlanta, GA 30322, USA; [email protected] (C.B.); [email protected] (Q.L.); [email protected] (A.R.) * Correspondence: [email protected] (M.B.V.); [email protected] (R.B.D.); Tel.: +1-404-504-1214 (M.B.V.); +1-404-727-6637 (R.B.D.) Academic Editor: Rafik Karaman Received: 24 July 2020; Accepted: 21 August 2020; Published: 22 August 2020 Abstract: Many enzymes are known to change conformations during their catalytic cycle, but the role of these protein motions is not well understood. Escherichia coli dihydrofolate reductase (DHFR) is a small, flexible enzyme that is often used as a model system for understanding enzyme dynamics. Recently, native tryptophan fluorescence was used as a probe to study micro- to millisecond dynamics of DHFR. Yet, because DHFR has five native tryptophans, the origin of the observed conformational changes could not be assigned to a specific region within the enzyme. Here, we use DHFR mutants, each with a single tryptophan as a probe for temperature jump fluorescence spectroscopy, to further inform our understanding of DHFR dynamics. The equilibrium tryptophan fluorescence of the mutants shows that each tryptophan is in a different environment and that wild-type DHFR fluorescence is not a simple summation of all the individual tryptophan fluorescence signatures due to tryptophan–tryptophan interactions. -

Predicting Protein-Membrane Interfaces of Peripheral Membrane
bioRxiv preprint doi: https://doi.org/10.1101/2021.06.28.450157; this version posted June 29, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. Predicting protein-membrane interfaces of pe- ripheral membrane proteins using ensemble machine learning Alexios Chatzigoulas1,2,* and Zoe Cournia1,* 1Biomedical Research Foundation, Academy of Athens, 4 Soranou Ephessiou, 11527 Athens, Greece, 2Depart- ment of Informatics and Telecommunications, National and Kapodistrian University of Athens, 15784 Athens, Greece *To whom correspondence should be addressed. Abstract Motivation: Abnormal protein-membrane attachment is involved in deregulated cellular pathways and in disease. Therefore, the possibility to modulate protein-membrane interactions represents a new promising therapeutic strategy for peripheral membrane proteins that have been considered so far undruggable. A major obstacle in this drug design strategy is that the membrane binding domains of peripheral membrane proteins are usually not known. The development of fast and efficient algorithms predicting the protein-membrane interface would shed light into the accessibility of membrane-protein interfaces by drug-like molecules. Results: Herein, we describe an ensemble machine learning methodology and algorithm for predicting membrane-penetrating residues. We utilize available experimental data in the literature for training 21 machine learning classifiers and a voting classifier. Evaluation of the ensemble classifier accuracy pro- duced a macro-averaged F1 score = 0.92 and an MCC = 0.84 for predicting correctly membrane-pen- etrating residues on unknown proteins of an independent test set. -

Constitutive Activation of G Protein-Coupled Receptors and Diseases: Insights Into Mechanisms of Activation and Therapeutics
Pharmacology & Therapeutics 120 (2008) 129–148 Contents lists available at ScienceDirect Pharmacology & Therapeutics journal homepage: www.elsevier.com/locate/pharmthera Associate editor: S. Enna Constitutive activation of G protein-coupled receptors and diseases: Insights into mechanisms of activation and therapeutics Ya-Xiong Tao ⁎ Department of Anatomy, Physiology and Pharmacology, 212 Greene Hall, College of Veterinary Medicine, Auburn University, Auburn, AL 36849, USA article info abstract The existence of constitutive activity for G protein-coupled receptors (GPCRs) was first described in 1980s. In Keywords: 1991, the first naturally occurring constitutively active mutations in GPCRs that cause diseases were reported G protein-coupled receptor Disease in rhodopsin. Since then, numerous constitutively active mutations that cause human diseases were reported Constitutively active mutation in several additional receptors. More recently, loss of constitutive activity was postulated to also cause Inverse agonist diseases. Animal models expressing some of these mutants confirmed the roles of these mutations in the Mechanism of activation pathogenesis of the diseases. Detailed functional studies of these naturally occurring mutations, combined Transgenic model with homology modeling using rhodopsin crystal structure as the template, lead to important insights into the mechanism of activation in the absence of crystal structure of GPCRs in active state. Search for inverse Abbreviations: agonists on these receptors will be critical for correcting the diseases cause by activating mutations in GPCRs. ADRP, autosomal dominant retinitis pigmentosa Theoretically, these inverse agonists are better therapeutics than neutral antagonists in treating genetic AgRP, Agouti-related protein AR, adrenergic receptor diseases caused by constitutively activating mutations in GPCRs. CAM, constitutively active mutant © 2008 Elsevier Inc. -

G Protein-Coupled Receptors: What a Difference a ‘Partner’ Makes
Int. J. Mol. Sci. 2014, 15, 1112-1142; doi:10.3390/ijms15011112 OPEN ACCESS International Journal of Molecular Sciences ISSN 1422-0067 www.mdpi.com/journal/ijms Review G Protein-Coupled Receptors: What a Difference a ‘Partner’ Makes Benoît T. Roux 1 and Graeme S. Cottrell 2,* 1 Department of Pharmacy and Pharmacology, University of Bath, Bath BA2 7AY, UK; E-Mail: [email protected] 2 Reading School of Pharmacy, University of Reading, Reading RG6 6UB, UK * Author to whom correspondence should be addressed; E-Mail: [email protected]; Tel.: +44-118-378-7027; Fax: +44-118-378-4703. Received: 4 December 2013; in revised form: 20 December 2013 / Accepted: 8 January 2014 / Published: 16 January 2014 Abstract: G protein-coupled receptors (GPCRs) are important cell signaling mediators, involved in essential physiological processes. GPCRs respond to a wide variety of ligands from light to large macromolecules, including hormones and small peptides. Unfortunately, mutations and dysregulation of GPCRs that induce a loss of function or alter expression can lead to disorders that are sometimes lethal. Therefore, the expression, trafficking, signaling and desensitization of GPCRs must be tightly regulated by different cellular systems to prevent disease. Although there is substantial knowledge regarding the mechanisms that regulate the desensitization and down-regulation of GPCRs, less is known about the mechanisms that regulate the trafficking and cell-surface expression of newly synthesized GPCRs. More recently, there is accumulating evidence that suggests certain GPCRs are able to interact with specific proteins that can completely change their fate and function. These interactions add on another level of regulation and flexibility between different tissue/cell-types. -

Conformational Proofreading: the Impact of Conformational
Conformational Proofreading: The Impact of Conformational Changes on the Specificity of Molecular Recognition Yonatan Savir1 and Tsvi Tlusty1,* 1Department of Physics of Complex Systems, the Weizmann Institute of Science, Rehovot, Israel, 76100 *Corresponding author. E-mail: [email protected] Abstract To perform recognition, molecules must locate and specifically bind their targets within a noisy biochemical environment with many look-alikes. Molecular recognition processes, especially the induced-fit mechanism, are known to involve conformational changes. This arises a basic question: does molecular recognition gain any advantage by such conformational changes? By introducing a simple statistical-mechanics approach, we study the effect of conformation and flexibility on the quality of recognition processes. Our model relates specificity to the conformation of the participant molecules and thus suggests a possible answer: Optimal specificity is achieved when the ligand is slightly off target, that is a conformational mismatch between the ligand and its main target improves the selectivity of the process. This indicates that deformations upon binding serve as a conformational proofreading mechanism, which may be selected for via evolution. Introduction Practically all biological systems rely on the ability of bio-molecules to specifically recognize each other. Examples are antibodies targeting antigens, regulatory proteins binding DNA and enzymes catalyzing their substrates. These and other molecular recognizers must locate and preferentially interact with their specific targets among a vast variety of molecules that are often structurally similar. This task is further complicated by the inherent noise in the biochemical environment, whose magnitude is comparable with that of the non- covalent binding interactions [1-3]. -

Dimers of Serotonin Receptors: Impact on Ligand Affinity and Signaling
Dimers of serotonin receptors: impact on ligand affinity and signaling Luc Maroteaux, Catherine Béchade, Anne Roumier To cite this version: Luc Maroteaux, Catherine Béchade, Anne Roumier. Dimers of serotonin receptors: impact on ligand affinity and signaling. Biochimie, Elsevier, 2019, 161, pp.23-33. 10.1016/j.biochi.2019.01.009. hal- 01996206 HAL Id: hal-01996206 https://hal.archives-ouvertes.fr/hal-01996206 Submitted on 28 Jan 2019 HAL is a multi-disciplinary open access L’archive ouverte pluridisciplinaire HAL, est archive for the deposit and dissemination of sci- destinée au dépôt et à la diffusion de documents entific research documents, whether they are pub- scientifiques de niveau recherche, publiés ou non, lished or not. The documents may come from émanant des établissements d’enseignement et de teaching and research institutions in France or recherche français ou étrangers, des laboratoires abroad, or from public or private research centers. publics ou privés. Maroteaux et al., 1 Dimers of serotonin receptors: impact on ligand affinity and signaling. Luc Maroteaux1,2,3, Catherine Béchade1,2,3, and Anne Roumier1,2,3 Affiliations: 1: INSERM UMR-S839, S1270, Paris, 75005, France; 2Sorbonne Université, Paris, 75005, France; 3Institut du Fer à Moulin, Paris, 75005, France. Running title: Dimers of serotonin receptors Correspondence should be addressed to: Luc Maroteaux INSERM UMR-S839, S1270, 17 rue du Fer a Moulin Paris, 75005, France; E-mail : [email protected] Abstract Membrane receptors often form complexes with other membrane proteins that directly interact with different effectors of the signal transduction machinery. G-protein-coupled receptors (GPCRs) were for long time considered as single pharmacological entities. -

Blood Platelet Adenosine Receptors As Potential Targets for Anti-Platelet Therapy
International Journal of Molecular Sciences Review Blood Platelet Adenosine Receptors as Potential Targets for Anti-Platelet Therapy Nina Wolska and Marcin Rozalski * Department of Haemostasis and Haemostatic Disorders, Chair of Biomedical Science, Medical University of Lodz, 92-215 Lodz, Poland; [email protected] * Correspondence: [email protected]; Tel.: +48-504-836-536 Received: 30 September 2019; Accepted: 1 November 2019; Published: 3 November 2019 Abstract: Adenosine receptors are a subfamily of highly-conserved G-protein coupled receptors. They are found in the membranes of various human cells and play many physiological functions. Blood platelets express two (A2A and A2B) of the four known adenosine receptor subtypes (A1,A2A, A2B, and A3). Agonization of these receptors results in an enhanced intracellular cAMP and the inhibition of platelet activation and aggregation. Therefore, adenosine receptors A2A and A2B could be targets for anti-platelet therapy, especially under circumstances when classic therapy based on antagonizing the purinergic receptor P2Y12 is insufficient or problematic. Apart from adenosine, there is a group of synthetic, selective, longer-lasting agonists of A2A and A2B receptors reported in the literature. This group includes agonists with good selectivity for A2A or A2B receptors, as well as non-selective compounds that activate more than one type of adenosine receptor. Chemically, most A2A and A2B adenosine receptor agonists are adenosine analogues, with either adenine or ribose substituted by single or multiple foreign substituents. However, a group of non-adenosine derivative agonists has also been described. This review aims to systematically describe known agonists of A2A and A2B receptors and review the available literature data on their effects on platelet function. -

The Interaction of Selective A1 and A2A Adenosine Receptor Antagonists with Magnesium and Zinc Ions in Mice: Behavioural, Biochemical and Molecular Studies
International Journal of Molecular Sciences Article The Interaction of Selective A1 and A2A Adenosine Receptor Antagonists with Magnesium and Zinc Ions in Mice: Behavioural, Biochemical and Molecular Studies Aleksandra Szopa 1,* , Karolina Bogatko 1, Mariola Herbet 2 , Anna Serefko 1 , Marta Ostrowska 2 , Sylwia Wo´sko 1, Katarzyna Swi´ ˛ader 3, Bernadeta Szewczyk 4, Aleksandra Wla´z 5, Piotr Skałecki 6, Andrzej Wróbel 7 , Sławomir Mandziuk 8, Aleksandra Pochodyła 3, Anna Kudela 2, Jarosław Dudka 2, Maria Radziwo ´n-Zaleska 9, Piotr Wla´z 10 and Ewa Poleszak 1,* 1 Chair and Department of Applied and Social Pharmacy, Laboratory of Preclinical Testing, Medical University of Lublin, 1 Chod´zkiStreet, PL 20–093 Lublin, Poland; [email protected] (K.B.); [email protected] (A.S.); [email protected] (S.W.) 2 Chair and Department of Toxicology, Medical University of Lublin, 8 Chod´zkiStreet, PL 20–093 Lublin, Poland; [email protected] (M.H.); [email protected] (M.O.); [email protected] (A.K.) [email protected] (J.D.) 3 Chair and Department of Applied and Social Pharmacy, Medical University of Lublin, 1 Chod´zkiStreet, PL 20–093 Lublin, Poland; [email protected] (K.S.);´ [email protected] (A.P.) 4 Department of Neurobiology, Polish Academy of Sciences, Maj Institute of Pharmacology, 12 Sm˛etnaStreet, PL 31–343 Kraków, Poland; [email protected] 5 Department of Pathophysiology, Medical University of Lublin, 8 Jaczewskiego Street, PL 20–090 Lublin, Poland; [email protected] Citation: Szopa, A.; Bogatko, K.; 6 Department of Commodity Science and Processing of Raw Animal Materials, University of Life Sciences, Herbet, M.; Serefko, A.; Ostrowska, 13 Akademicka Street, PL 20–950 Lublin, Poland; [email protected] M.; Wo´sko,S.; Swi´ ˛ader, K.; Szewczyk, 7 Second Department of Gynecology, 8 Jaczewskiego Street, PL 20–090 Lublin, Poland; B.; Wla´z,A.; Skałecki, P.; et al. -

Rods Contribute to the Light-Induced Phase Shift of The
Rods contribute to the light-induced phase shift of the retinal clock in mammals Hugo Calligaro, Christine Coutanson, Raymond Najjar, Nadia Mazzaro, Howard Cooper, Nasser Haddjeri, Marie-Paule Felder-Schmittbuhl, Ouria Dkhissi-Benyahya To cite this version: Hugo Calligaro, Christine Coutanson, Raymond Najjar, Nadia Mazzaro, Howard Cooper, et al.. Rods contribute to the light-induced phase shift of the retinal clock in mammals. PLoS Biology, Public Library of Science, 2019, 17 (3), pp.e2006211. 10.1371/journal.pbio.2006211. inserm-02137592 HAL Id: inserm-02137592 https://www.hal.inserm.fr/inserm-02137592 Submitted on 23 May 2019 HAL is a multi-disciplinary open access L’archive ouverte pluridisciplinaire HAL, est archive for the deposit and dissemination of sci- destinée au dépôt et à la diffusion de documents entific research documents, whether they are pub- scientifiques de niveau recherche, publiés ou non, lished or not. The documents may come from émanant des établissements d’enseignement et de teaching and research institutions in France or recherche français ou étrangers, des laboratoires abroad, or from public or private research centers. publics ou privés. RESEARCH ARTICLE Rods contribute to the light-induced phase shift of the retinal clock in mammals Hugo Calligaro1, Christine Coutanson1, Raymond P. Najjar2,3, Nadia Mazzaro4, Howard M. Cooper1, Nasser Haddjeri1, Marie-Paule Felder-Schmittbuhl4, Ouria Dkhissi- Benyahya1* 1 Univ Lyon, Universite Claude Bernard Lyon 1, Inserm, Stem Cell and Brain Research Institute, Bron, France,