Membrane Protein Structure Determination and Characterisation by Solution and Solid-State NMR
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

A Network of Molecular Switches Controls the Activation of the Two-Component Response Regulator Ntrc
ARTICLE Received 8 Aug 2014 | Accepted 26 Apr 2015 | Published 15 Jun 2015 DOI: 10.1038/ncomms8283 A network of molecular switches controls the activation of the two-component response regulator NtrC Dan K. Vanatta1, Diwakar Shukla1,2,3, Morgan Lawrenz1 & Vijay S. Pande1,2 Recent successes in simulating protein structure and folding dynamics have demonstrated the power of molecular dynamics to predict the long timescale behaviour of proteins. Here, we extend and improve these methods to predict molecular switches that characterize conformational change pathways between the active and inactive state of nitrogen regulatory protein C (NtrC). By employing unbiased Markov state model-based molecular dynamics simulations, we construct a dynamic picture of the activation pathways of this key bacterial signalling protein that is consistent with experimental observations and predicts new mutants that could be used for validation of the mechanism. Moreover, these results suggest a novel mechanistic paradigm for conformational switching. 1 Department of Chemistry, Stanford University, Stanford, California 94305, USA. 2 SIMBIOS NIH Center for Biomedical Computation, Stanford University, Stanford, California 94305,USA. 3 Department of Chemical and Bimolecular Engineering, University of Illinois at Urbana-Champaign, Urbana, Illionios 61801, USA. Correspondence and requests for materials should be addressed to V.S.P. (email: [email protected]). NATURE COMMUNICATIONS | 6:7283 | DOI: 10.1038/ncomms8283 | www.nature.com/naturecommunications 1 & 2015 Macmillan Publishers Limited. All rights reserved. ARTICLE NATURE COMMUNICATIONS | DOI: 10.1038/ncomms8283 roteins involved in cellular signalling change their con- Results formation in response to changes in environment (input Simulations reveal molecular switches for NtrC activation. Psignal), such as ligand binding or chemical modification, MSMs for NtrC were built using snapshots along the simulated to control downstream cellular processes (output signal)1. -

Membrane Protein Stabilization Strategies for Structural and Functional Studies
membranes Review Membrane Protein Stabilization Strategies for Structural and Functional Studies Ekaitz Errasti-Murugarren 1,2,*, Paola Bartoccioni 1,2 and Manuel Palacín 1,2,3,* 1 Laboratory of Amino acid Transporters and Disease, Institute for Research in Biomedicine (IRB Barcelona), The Barcelona Institute of Science and Technology (BIST), Baldiri Reixac 10, 08028 Barcelona, Spain; [email protected] 2 CIBERER (Centro Español en Red de Biomedicina de Enfermedades Raras), 28029 Barcelona, Spain 3 Department of Biochemistry and Molecular Biomedicine, Universitat de Barcelona, 08028 Barcelona, Spain * Correspondence: [email protected] (E.E.-M.); [email protected] (M.P.) Abstract: Accounting for nearly two-thirds of known druggable targets, membrane proteins are highly relevant for cell physiology and pharmacology. In this regard, the structural determination of pharmacologically relevant targets would facilitate the intelligent design of new drugs. The structural biology of membrane proteins is a field experiencing significant growth as a result of the development of new strategies for structure determination. However, membrane protein preparation for structural studies continues to be a limiting step in many cases due to the inherent instability of these molecules in non-native membrane environments. This review describes the approaches that have been developed to improve membrane protein stability. Membrane protein mutagenesis, detergent selection, lipid membrane mimics, antibodies, and ligands are described in this review as approaches to facilitate the production of purified and stable membrane proteins of interest for structural and functional studies. Keywords: membrane proteins; stability; mutagenesis; detergent; lipid; antibody; nanobody; ligand Citation: Errasti-Murugarren, E.; Bartoccioni, P.; Palacín, M. Membrane Protein Stabilization Strategies for Structural and Functional Studies. -
Spring 2013 Lecture 23
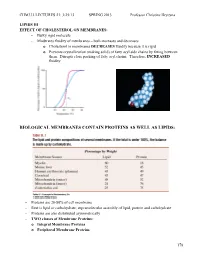
CHM333 LECTURES 23: 3/25/13 SPRING 2013 Professor Christine Hrycyna LIPIDS III EFFECT OF CHOLESTEROL ON MEMBRANES: - Bulky rigid molecule - Moderates fluidity of membranes – both increases and decreases o Cholesterol in membranes DECREASES fluidity because it is rigid o Prevents crystallization (making solid) of fatty acyl side chains by fitting between them. Disrupts close packing of fatty acyl chains. Therefore, INCREASED fluidity BIOLOGICAL MEMBRANES CONTAIN PROTEINS AS WELL AS LIPIDS: - Proteins are 20-80% of cell membrane - Rest is lipid or carbohydrate; supramolecular assembly of lipid, protein and carbohydrate - Proteins are also distributed asymmetrically - TWO classes of Membrane Proteins: o Integral Membrane Proteins o Peripheral Membrane Proteins 178 CHM333 LECTURES 23: 3/25/13 SPRING 2013 Professor Christine Hrycyna - INTEGRAL MEMBRANE PROTEINS o Located WITHIN the lipid bilayer o Usually span the bilayer one or more times – called transmembrane (TM) proteins o Hydrophobic amino acids interact with fatty acid chains in the hydrophobic core of the membrane o Can be removed from the membrane with detergents like SDS – need to disrupt the hydrophobic interactions § Membrane Disruption Animation: o http://www.youtube.com/watch?v=AHT37pvcjc0 o Function: § Transporters – moving molecules into or out of cells or cell membranes § Receptors – transmitting signals from outside of the cell to the inside - β Barrel Integral Membrane Proteins § Barrel-shaped membrane protein that is made up of antiparallel β-strands with hydrophilic (interior) and hydrophobic (facing lipid tails). § So far found only in outer membranes of Gram-negative bacteria, cell wall of Gram-positive bacteria, and outer membranes of mitochondria and chloroplasts. 179 CHM333 LECTURES 23: 3/25/13 SPRING 2013 Professor Christine Hrycyna - α-Helical Membrane Proteins - Can cross the membrane once or many times and have multiple transmembrane segments. -

Snapshot: ER-Associated Protein Degradation Pathways Shinichi Kawaguchi and Davis T.W
SnapShot: ER-Associated Protein Degradation Pathways Shinichi Kawaguchi and Davis T.W. Ng Temasek Life Sciences Laboratory and Department of Biological Sciences, National University of Singapore, Singapore 117604 1230 Cell 129, June 15, 2007 ©2007 Elsevier Inc. DOI 10.1016/j.cell.2007.06.005 See online version for legend and references. SnapShot: ER-Associated Protein Degradation Pathways Shinichi Kawaguchi and Davis T.W. Ng Temasek Life Sciences Laboratory and Department of Biological Sciences, National University of Singapore, Singapore 117604 Arrows specify the routes of individual pathways. All pathways culminate in substrate degradation by the 26S proteasome. (Left) ERAD Pathways in Budding Yeast (A) Newly synthesized secretory and membrane proteins enter the ER through the Sec61 protein-conducting channel complex unfolded. Hsp70-related molecular chaperones (Kar2p) bind to nascent polypeptides in the ER lumen and to the cytosolic domains of membrane proteins (Hsp70, B). These factors assist in substrate folding and also assist in their disposal if they fail to fold. Mannose residues on misfolded glycoproteins are trimmed by the ER mannosidase Mns1p (E). Mannose trimming facilitates the recognition of misfolded glycoproteins by luminally oriented lectin factors Htm1p and Yos9p. (C, D, and F) At least two ER membrane-localized E3 ubiquitin ligases organize protein complexes that receive and process misfolded proteins. These complexes define three pathways that recognize lesions in the cytosolic (ERAD-C), transmembrane (ERAD-M), and luminal (ERAD-L) domains of substrates. Both ERAD-M and ERAD-L use the Hrd1 ubiquitin ligase but the luminal factor Yos9p is dispensable for ERAD-M. A Hrd1 complex lacking Yos9p has been observed suggesting dedicated complexes for all three pathways. -

Nuclear Ubiquitin-Proteasome Pathways in Proteostasis Maintenance
biomolecules Review Nuclear Ubiquitin-Proteasome Pathways in Proteostasis Maintenance Dina Frani´c †, Klara Zubˇci´c † and Mirta Boban * Croatian Institute for Brain Research, School of Medicine, University of Zagreb, 10000 Zagreb, Croatia; [email protected] (D.F.); [email protected] (K.Z.) * Correspondence: [email protected] † Equal contribution. Abstract: Protein homeostasis, or proteostasis, is crucial for the functioning of a cell, as proteins that are mislocalized, present in excessive amounts, or aberrant due to misfolding or other type of damage can be harmful. Proteostasis includes attaining the correct protein structure, localization, and the for- mation of higher order complexes, and well as the appropriate protein concentrations. Consequences of proteostasis imbalance are evident in a range of neurodegenerative diseases characterized by protein misfolding and aggregation, such as Alzheimer’s, Parkinson’s, and amyotrophic lateral sclerosis. To protect the cell from the accumulation of aberrant proteins, a network of protein quality control (PQC) pathways identifies the substrates and direct them towards refolding or elimination via regulated protein degradation. The main pathway for degradation of misfolded proteins is the ubiquitin-proteasome system. PQC pathways have been first described in the cytoplasm and the endoplasmic reticulum, however, accumulating evidence indicates that the nucleus is an important PQC compartment for ubiquitination and proteasomal degradation of not only nuclear, but also cyto- plasmic proteins. In this review, we summarize the nuclear ubiquitin-proteasome pathways involved in proteostasis maintenance in yeast, focusing on inner nuclear membrane-associated degradation (INMAD) and San1-mediated protein quality control. Keywords: proteasome; ubiquitin; nucleus; inner nuclear membrane; yeast; proteostasis; protein quality control; protein misfolding Citation: Frani´c,D.; Zubˇci´c,K.; Boban, M. -

Site-Specific Tryptophan Labels Reveal Local Microsecond
molecules Article Site-Specific Tryptophan Labels Reveal Local Microsecond–Millisecond Motions of Dihydrofolate Reductase Morgan B. Vaughn 1,*, Chloe Biren 2, Qun Li 2, Ashwin Ragupathi 2 and R. Brian Dyer 2,* 1 Division of Natural Sciences, Oglethorpe University, Atlanta, GA 30319, USA 2 Department of Chemistry, Emory University, Atlanta, GA 30322, USA; [email protected] (C.B.); [email protected] (Q.L.); [email protected] (A.R.) * Correspondence: [email protected] (M.B.V.); [email protected] (R.B.D.); Tel.: +1-404-504-1214 (M.B.V.); +1-404-727-6637 (R.B.D.) Academic Editor: Rafik Karaman Received: 24 July 2020; Accepted: 21 August 2020; Published: 22 August 2020 Abstract: Many enzymes are known to change conformations during their catalytic cycle, but the role of these protein motions is not well understood. Escherichia coli dihydrofolate reductase (DHFR) is a small, flexible enzyme that is often used as a model system for understanding enzyme dynamics. Recently, native tryptophan fluorescence was used as a probe to study micro- to millisecond dynamics of DHFR. Yet, because DHFR has five native tryptophans, the origin of the observed conformational changes could not be assigned to a specific region within the enzyme. Here, we use DHFR mutants, each with a single tryptophan as a probe for temperature jump fluorescence spectroscopy, to further inform our understanding of DHFR dynamics. The equilibrium tryptophan fluorescence of the mutants shows that each tryptophan is in a different environment and that wild-type DHFR fluorescence is not a simple summation of all the individual tryptophan fluorescence signatures due to tryptophan–tryptophan interactions. -

Micelle Formation and the Hydrophobic Effect†
6778 J. Phys. Chem. B 2004, 108, 6778-6781 Micelle Formation and the Hydrophobic Effect† Lutz Maibaum,‡ Aaron R. Dinner,§ and David Chandler*,‡ Department of Chemistry, UniVersity of California, Berkeley, California 94720, and Department of Chemistry, UniVersity of Chicago, Chicago, Illinois 60637 ReceiVed: NoVember 14, 2003; In Final Form: January 14, 2004 The tendency of amphiphilic molecules to form micelles in aqueous solution is a consequence of the hydrophobic effect. The fundamental difference between micelle assembly and macroscopic phase separation is the stoichiometric constraint that frustrates the demixing of polar and hydrophobic groups. We present a theory for micelle assembly that combines the account of this constraint with a description of the hydrophobic driving force. The latter arises from the length scale dependence of aqueous solvation. The theoretical predictions for temperature dependence and surfactant chain length dependence of critical micelle concentrations for nonionic surfactants agree favorably with experiment. I. Introduction in accord with experimental observations. An appendix is used to augment the discussion in section II. This paper concerns the formation of micelles, which are the simplest form of amphiphilic assemblies. Our treatment of this II. Theory phenomenon is based upon the length scale dependence of hydrophobic effects.1,2 Namely, the free energy to solvate small A. Law of Mass Action. We consider an aqueous solution hydrophobic molecules scales linearly with solute volume, of neutral amphiphilic molecules (i.e., nonionic surfactants), each whereas that to solvate large hydrophobic species scales linearly of which has a single alkyl chain as its hydrophobic tail. In with surface area. The crossover from one regime to the other general, amphiphiles can form aggregates of various sizes and occurs when the oily species presents a surface in water shapes. -

Tutorial on Working with Micelles and Other Model Membranes
Tutorial on Working with Micelles and Model Membranes Chuck Sanders Dept. of Biochemistry, Dept. of Medicine, and Center for Structural Biology Vanderbilt University School of Medicine. http://structbio.vanderbilt.edu/sanders/ March, 2017 There are two general classes of membrane proteins. This presentation is on working with integral MPs, which traditionally could be removed from the membrane only by dissolving the membrane with detergents or organic solvents. Multilamellar Vesicles: onion-like assemblies. Each layer is one bilayer. A thin layer of water separates each bilayer. MLVs are what form when lipid powders are dispersed in water. They form spontaneously. Cryo-EM Micrograph of a Multilamellar Vesicle (K. Mittendorf, C. Sanders, and M. Ohi) Unilamellar Multilamellar Vesicle Vesicle Advances in Anesthesia 32(1):133-147 · 2014 Energy from sonication, physical manipulation (such as extrusion by forcing MLV dispersions through filters with fixed pore sizes), or some other high energy mechanism is required to convert multilayered bilayer assemblies into unilamellar vesicles. If the MLVs contain a membrane protein then you should worry about whether the protein will survive these procedures in folded and functional form. Vesicles can also be prepared by dissolving lipids using detergents and then removing the detergent using BioBeads-SM dialysis, size exclusion chromatography or by diluting the solution to below the detergent’s critical micelle concentration. These are much gentler methods that a membrane protein may well survive with intact structure and function. From: Avanti Polar Lipids Catalog Bilayers can undergo phase transitions at a critical temperature, Tm. Native bilayers are usually in the fluid (liquid crystalline) phase. -

Predicting Protein-Membrane Interfaces of Peripheral Membrane
bioRxiv preprint doi: https://doi.org/10.1101/2021.06.28.450157; this version posted June 29, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. Predicting protein-membrane interfaces of pe- ripheral membrane proteins using ensemble machine learning Alexios Chatzigoulas1,2,* and Zoe Cournia1,* 1Biomedical Research Foundation, Academy of Athens, 4 Soranou Ephessiou, 11527 Athens, Greece, 2Depart- ment of Informatics and Telecommunications, National and Kapodistrian University of Athens, 15784 Athens, Greece *To whom correspondence should be addressed. Abstract Motivation: Abnormal protein-membrane attachment is involved in deregulated cellular pathways and in disease. Therefore, the possibility to modulate protein-membrane interactions represents a new promising therapeutic strategy for peripheral membrane proteins that have been considered so far undruggable. A major obstacle in this drug design strategy is that the membrane binding domains of peripheral membrane proteins are usually not known. The development of fast and efficient algorithms predicting the protein-membrane interface would shed light into the accessibility of membrane-protein interfaces by drug-like molecules. Results: Herein, we describe an ensemble machine learning methodology and algorithm for predicting membrane-penetrating residues. We utilize available experimental data in the literature for training 21 machine learning classifiers and a voting classifier. Evaluation of the ensemble classifier accuracy pro- duced a macro-averaged F1 score = 0.92 and an MCC = 0.84 for predicting correctly membrane-pen- etrating residues on unknown proteins of an independent test set. -

How Does Protein Zero Assemble Compact Myelin?
Preprints (www.preprints.org) | NOT PEER-REVIEWED | Posted: 13 May 2020 doi:10.20944/preprints202005.0222.v1 Peer-reviewed version available at Cells 2020, 9, 1832; doi:10.3390/cells9081832 Perspective How Does Protein Zero Assemble Compact Myelin? Arne Raasakka 1,* and Petri Kursula 1,2 1 Department of Biomedicine, University of Bergen, Jonas Lies vei 91, NO-5009 Bergen, Norway 2 Faculty of Biochemistry and Molecular Medicine & Biocenter Oulu, University of Oulu, Aapistie 7A, FI-90220 Oulu, Finland; [email protected] * Correspondence: [email protected] Abstract: Myelin protein zero (P0), a type I transmembrane protein, is the most abundant protein in peripheral nervous system (PNS) myelin – the lipid-rich, periodic structure that concentrically encloses long axonal segments. Schwann cells, the myelinating glia of the PNS, express P0 throughout their development until the formation of mature myelin. In the intramyelinic compartment, the immunoglobulin-like domain of P0 bridges apposing membranes together via homophilic adhesion, forming a dense, macroscopic ultrastructure known as the intraperiod line. The C-terminal tail of P0 adheres apposing membranes together in the narrow cytoplasmic compartment of compact myelin, much like myelin basic protein (MBP). In mouse models, the absence of P0, unlike that of MBP or P2, severely disturbs the formation of myelin. Therefore, P0 is the executive molecule of PNS myelin maturation. How and when is P0 trafficked and modified to enable myelin compaction, and how disease mutations that give rise to incurable peripheral neuropathies alter the function of P0, are currently open questions. The potential mechanisms of P0 function in myelination are discussed, providing a foundation for the understanding of mature myelin development and how it derails in peripheral neuropathies. -

Structural Insights Into Membrane Fusion Mediated by Convergent Small Fusogens
cells Review Structural Insights into Membrane Fusion Mediated by Convergent Small Fusogens Yiming Yang * and Nandini Nagarajan Margam Department of Microbiology and Immunology, Dalhousie University, Halifax, NS B3H 4R2, Canada; [email protected] * Correspondence: [email protected] Abstract: From lifeless viral particles to complex multicellular organisms, membrane fusion is inarguably the important fundamental biological phenomena. Sitting at the heart of membrane fusion are protein mediators known as fusogens. Despite the extensive functional and structural characterization of these proteins in recent years, scientists are still grappling with the fundamental mechanisms underlying membrane fusion. From an evolutionary perspective, fusogens follow divergent evolutionary principles in that they are functionally independent and do not share any sequence identity; however, they possess structural similarity, raising the possibility that membrane fusion is mediated by essential motifs ubiquitous to all. In this review, we particularly emphasize structural characteristics of small-molecular-weight fusogens in the hope of uncovering the most fundamental aspects mediating membrane–membrane interactions. By identifying and elucidating fusion-dependent functional domains, this review paves the way for future research exploring novel fusogens in health and disease. Keywords: fusogen; SNARE; FAST; atlastin; spanin; myomaker; myomerger; membrane fusion 1. Introduction Citation: Yang, Y.; Margam, N.N. Structural Insights into Membrane Membrane fusion -

Membrane Proteins Are Associated with the Membrane of a Cell Or Particular Organelle and Are Generally More Problematic to Purify Than Water-Soluble Proteins
Strategies for the Purification of Membrane Proteins Sinéad Marian Smith Department of Clinical Medicine, School of Medicine, Trinity College Dublin, Ireland. Email: [email protected] Abstract Although membrane proteins account for approximately 30 % of the coding regions of all sequenced genomes and play crucial roles in many fundamental cell processes, there are relatively few membranes with known 3D structure. This is likely due to technical challenges associated with membrane protein extraction, solubilization and purification. Membrane proteins are classified based on the level of interaction with membrane lipid bilayers, with peripheral membrane proteins associating non- covalently with the membrane, and integral membrane proteins associating more strongly by means of hydrophobic interactions. Generally speaking, peripheral membrane proteins can be purified by milder techniques than integral membrane proteins, whose extraction require phospholipid bilayer disruption by detergents. Here, important criteria for strategies of membrane protein purification are addressed, with a focus on the initial stages of membrane protein solublilization, where problems are most frequently are encountered. Protocols are outlined for the successful extraction of peripheral membrane proteins, solubilization of integral membrane proteins, and detergent removal which is important not only for retaining native protein stability and biological functions, but also for the efficiency of downstream purification techniques. Key Words: peripheral membrane protein, integral membrane protein, detergent, protein purification, protein solubilization. 1. Introduction Membrane proteins are associated with the membrane of a cell or particular organelle and are generally more problematic to purify than water-soluble proteins. Membrane proteins represent approximately 30 % of the open-reading frames of an organism’s genome (1-4), and play crucial roles in basic cell functions including signal transduction, energy production, nutrient uptake and cell-cell communication.