Primepcr™Assay Validation Report
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Transcriptome Analyses of Rhesus Monkey Pre-Implantation Embryos Reveal A
Downloaded from genome.cshlp.org on September 23, 2021 - Published by Cold Spring Harbor Laboratory Press Transcriptome analyses of rhesus monkey pre-implantation embryos reveal a reduced capacity for DNA double strand break (DSB) repair in primate oocytes and early embryos Xinyi Wang 1,3,4,5*, Denghui Liu 2,4*, Dajian He 1,3,4,5, Shengbao Suo 2,4, Xian Xia 2,4, Xiechao He1,3,6, Jing-Dong J. Han2#, Ping Zheng1,3,6# Running title: reduced DNA DSB repair in monkey early embryos Affiliations: 1 State Key Laboratory of Genetic Resources and Evolution, Kunming Institute of Zoology, Chinese Academy of Sciences, Kunming, Yunnan 650223, China 2 Key Laboratory of Computational Biology, CAS Center for Excellence in Molecular Cell Science, Collaborative Innovation Center for Genetics and Developmental Biology, Chinese Academy of Sciences-Max Planck Partner Institute for Computational Biology, Shanghai Institutes for Biological Sciences, Chinese Academy of Sciences, Shanghai 200031, China 3 Yunnan Key Laboratory of Animal Reproduction, Kunming Institute of Zoology, Chinese Academy of Sciences, Kunming, Yunnan 650223, China 4 University of Chinese Academy of Sciences, Beijing, China 5 Kunming College of Life Science, University of Chinese Academy of Sciences, Kunming, Yunnan 650204, China 6 Primate Research Center, Kunming Institute of Zoology, Chinese Academy of Sciences, Kunming, 650223, China * Xinyi Wang and Denghui Liu contributed equally to this work 1 Downloaded from genome.cshlp.org on September 23, 2021 - Published by Cold Spring Harbor Laboratory Press # Correspondence: Jing-Dong J. Han, Email: [email protected]; Ping Zheng, Email: [email protected] Key words: rhesus monkey, pre-implantation embryo, DNA damage 2 Downloaded from genome.cshlp.org on September 23, 2021 - Published by Cold Spring Harbor Laboratory Press ABSTRACT Pre-implantation embryogenesis encompasses several critical events including genome reprogramming, zygotic genome activation (ZGA) and cell fate commitment. -

A Master Autoantigen-Ome Links Alternative Splicing, Female Predilection, and COVID-19 to Autoimmune Diseases
bioRxiv preprint doi: https://doi.org/10.1101/2021.07.30.454526; this version posted August 4, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY 4.0 International license. A Master Autoantigen-ome Links Alternative Splicing, Female Predilection, and COVID-19 to Autoimmune Diseases Julia Y. Wang1*, Michael W. Roehrl1, Victor B. Roehrl1, and Michael H. Roehrl2* 1 Curandis, New York, USA 2 Department of Pathology, Memorial Sloan Kettering Cancer Center, New York, USA * Correspondence: [email protected] or [email protected] 1 bioRxiv preprint doi: https://doi.org/10.1101/2021.07.30.454526; this version posted August 4, 2021. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY 4.0 International license. Abstract Chronic and debilitating autoimmune sequelae pose a grave concern for the post-COVID-19 pandemic era. Based on our discovery that the glycosaminoglycan dermatan sulfate (DS) displays peculiar affinity to apoptotic cells and autoantigens (autoAgs) and that DS-autoAg complexes cooperatively stimulate autoreactive B1 cell responses, we compiled a database of 751 candidate autoAgs from six human cell types. At least 657 of these have been found to be affected by SARS-CoV-2 infection based on currently available multi-omic COVID data, and at least 400 are confirmed targets of autoantibodies in a wide array of autoimmune diseases and cancer. -

H1 Linker Histones Silence Repetitive Elements by Promoting Both Histone H3K9 Methylation and Chromatin Compaction
H1 linker histones silence repetitive elements by promoting both histone H3K9 methylation and chromatin compaction Sean E. Healtona,1,2, Hugo D. Pintoa,1, Laxmi N. Mishraa, Gregory A. Hamiltona,b, Justin C. Wheata, Kalina Swist-Rosowskac, Nicholas Shukeirc, Yali Doud, Ulrich Steidla, Thomas Jenuweinc, Matthew J. Gamblea,b, and Arthur I. Skoultchia,2 aDepartment of Cell Biology, Albert Einstein College of Medicine, Bronx, NY 10461; bDepartment of Molecular Pharmacology, Albert Einstein College of Medicine, Bronx, NY 10461; cMax Planck Institute of Immunobiology and Epigenetics, Stübeweg 51, Freiburg D-79108, Germany; and dDepartment of Pathology, University of Michigan, Ann Arbor, MI 48109 Edited by Robert G. Roeder, Rockefeller University, New York, NY, and approved May 1, 2020 (received for review December 15, 2019) Nearly 50% of mouse and human genomes are composed of repetitive mechanisms of this regulation have not been fully explored. To sequences. Transcription of these sequences is tightly controlled during further investigate the roles of H1 in epigenetic regulation, we have development to prevent genomic instability, inappropriate gene used CRISPR-Cas9–mediated genome editing to inactivate addi- activation and other maladaptive processes. Here, we demonstrate tional H1 genes in the H1 TKO ES cells and thereby deplete the H1 an integral role for H1 linker histones in silencing repetitive elements in content to even lower levels. mouse embryonic stem cells. Strong H1 depletion causes a profound Nearly 50% of the mouse and human genomes consist of re- de-repression of several classes of repetitive sequences, including major petitive sequences, including tandem repeats, such as satellite se- satellite, LINE-1, and ERV. -

Practice Online Genome Databases Kaynak
ONLINE GENOME DATABASES AÜTF D1M1, 2019 www.ensemble.org Ensemble’s latest humAn genome “assembly” İnsAn genomundA ArAmA penceresi GRCh38.p13 stAtistics HumAn chromosomes GRCh38.p12 statistics İnsan karyotip görüntüsü Click for further informAtion Diğer kromozomlArı seçin Protein coding genes Non coding genes (RNA genes) Pseudogenes İnsan 1. kromozom görüntüsü ve bilgileri Synteny regions for compArison between orgAnisms’ chromosomes Synteny regions compAred Select orgAnism Select chromosome HumAn chromosome 2 Syntenic regions on chimps chromosomes 2A, 2B www.genenames.org HGNC (Human Genome Organization Gene Nomenclature Committee) Inquiry for gene names and symbols Gen isim ve sembollerini sorgulama örnek: histon genleri Histone genes listing www.genenames.org Histon genleri Given below enlArged Histone cluster 1 H1 gene family member A Symbol: HIST1H1A Kromozom lokusu: 6p22.2 Click for detAils HIST1H1A Click to go to Ensemble link Click sequence HIST1H1A HIST1H1A gene sequence is reAched by clicking the gene row Chromosome location in between red bars The below window in red rectAngle shows the region in detAil with neighboring sequneces/genes Red thin line is enlArged to show the gene of interest in detAil. HIST1H1A geni haritası Gene page TrAnscripts, exons, cDNA and proteins can be reAched HIST1H1A bilgi sayfası 43 orthologues HIST1H1A orthologues İlgilendiğiniz türlerle ilgili seçim yapabilir Scroll down to see other orgAnisms HIST1H1A human – primate orthologues See orgAnisms in a phylogenetic tree Align DNA/protein sequences (choose -

IGF1R Deficiency Attenuates Acute Inflammatory Response in A
www.nature.com/scientificreports OPEN IGF1R deficiency attenuates acute inflammatory response in a bleomycin-induced lung injury Received: 7 November 2016 Accepted: 17 May 2017 mouse model Published: xx xx xxxx Sergio Piñeiro-Hermida1, Icíar P. López1, Elvira Alfaro-Arnedo1, Raquel Torrens1, María Iñiguez2, Lydia Alvarez-Erviti3, Carlos Ruíz-Martínez4 & José G. Pichel 1 IGF1R (Insulin-like Growth Factor 1 Receptor) is a tyrosine kinase with pleiotropic cellular functions. IGF activity maintains human lung homeostasis and is implicated in pulmonary diseases such as cancer, ARDS, COPD, asthma and fibrosis. Here we report that lung transcriptome analysis in mice with a postnatally-induced Igf1r gene deletion showed differentially expressed genes with potentially protective roles related to epigenetics, redox and oxidative stress. After bleomycin-induced lung injury, IGF1R-deficient mice demonstrated improved survival within a week. Three days post injury, IGF1R- deficient lungs displayed changes in expression of IGF system-related genes and reduced vascular fragility and permeability. Mutant lungs presented reduced inflamed area, down-regulation of pro- inflammatory markers and up-regulation of resolution indicators. Decreased inflammatory cell presence in BALF was reflected in diminished lung infiltration mainly affecting neutrophils, also corroborated by reduced neutrophil numbers in bone marrow, as well as reduced lymphocyte and alveolar macrophage counts. Additionally, increased SFTPC expression together with hindered HIF1A expression and augmented levels of Gpx8 indicate that IGF1R deficiency protects against alveolar damage. These findings identify IGF1R as an important player in murine acute lung inflammation, suggesting that targeting IGF1R may counteract the inflammatory component of many lung diseases. Inflammation is a relevant component of many lung diseases including ARDS, COPD, asthma, cancer, fibrosis and pneumonia1–5. -

Supplemental Data.Pdf
Supplementary material -Table of content Supplementary Figures (Fig 1- Fig 6) Supplementary Tables (1-13) Lists of genes belonging to distinct biological processes identified by GREAT analyses to be significantly enriched with UBTF1/2-bound genes Supplementary Table 14 List of the common UBTF1/2 bound genes within +/- 2kb of their TSSs in NIH3T3 and HMECs. Supplementary Table 15 List of gene identified by microarray expression analysis to be differentially regulated following UBTF1/2 knockdown by siRNA Supplementary Table 16 List of UBTF1/2 binding regions overlapping with histone genes in NIH3T3 cells Supplementary Table 17 List of UBTF1/2 binding regions overlapping with histone genes in HMEC Supplementary Table 18 Sequences of short interfering RNA oligonucleotides Supplementary Table 19 qPCR primer sequences for qChIP experiments Supplementary Table 20 qPCR primer sequences for reverse transcription-qPCR Supplementary Table 21 Sequences of primers used in CHART-PCR Supplementary Methods Supplementary Fig 1. (A) ChIP-seq analysis of UBTF1/2 and Pol I (POLR1A) binding across mouse rDNA. UBTF1/2 is enriched at the enhancer and promoter regions and along the entire transcribed portions of rDNA with little if any enrichment in the intergenic spacer (IGS), which separates the rDNA repeats. This enrichment coincides with the distribution of the largest subunit of Pol I (POLR1A) across the rDNA. All sequencing reads were mapped to the published complete sequence of the mouse rDNA repeat (Gene bank accession number: BK000964). The graph represents the frequency of ribosomal sequences enriched in UBTF1/2 and Pol I-ChIPed DNA expressed as fold change over those of input genomic DNA. -

Towards Understanding the Regulation of Histone H1 Somatic Subtypes with Omics
bioRxiv preprint doi: https://doi.org/10.1101/2020.09.30.320572; this version posted October 2, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Towards understanding the regulation of histone H1 somatic subtypes with OMICs. Authors: Inma Ponte1, Marta Andrés1, Albert Jordan2, Alicia Roque1*. 1Biochemistry and Molecular Biology Department, Bioscience Faculty, Autonomous University of Barcelona, Spain. 2Molecular Biology Institute of Barcelona (IBMB-CSIC), Barcelona, Spain. *corresponding author Keywords: co-regulation; transcription factors; compensatory effects; subtype functional differentiation; chromatin compartments; topologically associated domains (TADs) 1 bioRxiv preprint doi: https://doi.org/10.1101/2020.09.30.320572; this version posted October 2, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Abstract Background: Histone H1 is involved in the regulation of chromatin higher-order structure and compaction. In humans, histone H1 is a multigene family with seven subtypes differentially expressed in somatic cells. Which are the regulatory mechanisms that determine the variability of the H1 complement is a long-standing biological question regarding histone H1. We have used a new approach based on the integration of OMICs data to address this question. Results: We have examined the 3D-chromatin structure, the binding of transcription factors (TFs), and the expression of somatic H1 genes in human cell lines, using data from public repositories, such as ENCODE. Analysis of Hi-C, ChIP-seq, and RNA-seq data, have shown that transcriptional control has a greater impact on H1 regulation than previously thought. -

TET1 Is a Tumor Suppressor of Hematopoietic Malignancy
TET1 is a tumor suppressor of hematopoietic malignancy The MIT Faculty has made this article openly available. Please share how this access benefits you. Your story matters. Citation Cimmino, Luisa et al. “TET1 Is a Tumor Suppressor of Hematopoietic Malignancy.” Nature Immunology 16.6 (2015): 653– 662. As Published http://dx.doi.org/10.1038/ni.3148 Publisher Nature Publishing Group Version Author's final manuscript Citable link http://hdl.handle.net/1721.1/108307 Terms of Use Article is made available in accordance with the publisher's policy and may be subject to US copyright law. Please refer to the publisher's site for terms of use. HHS Public Access Author manuscript Author Manuscript Author ManuscriptNat Immunol Author Manuscript. Author manuscript; Author Manuscript available in PMC 2015 August 21. Published in final edited form as: Nat Immunol. 2015 June ; 16(6): 653–662. doi:10.1038/ni.3148. Tet1 is a tumor suppressor of hematopoietic malignancy Luisa Cimmino1,2,9, Meelad M. Dawlaty3,4,9, Delphine Ndiaye-Lobry1,2, Yoon Sing Yap1,2, Sofia Bakogianni1,2, Yiting Yu5, Sanchari Bhattacharyya5, Rita Shaknovich6, Huimin Geng7, Camille Lobry1,2, Jasper Mullenders1,2, Bryan King1,2, Thomas Trimarchi1,2, Beatriz Aranda-Orgilles1,2, Cynthia Liu1, Steven Shen8, Amit K. Verma5, Rudolf Jaenisch3,4, and Iannis Aifantis1,2 1Howard Hughes Medical Institute and Department of Pathology, NYU School of Medicine, New York, NY, 10016, USA 2NYU Cancer Institute and Helen L. and Martin S. Kimmel Center for Stem Cell Biology, NYU School of Medicine, New -
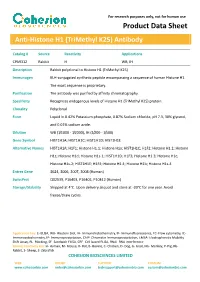
Product Data Sheet
For research purposes only, not for human use Product Data Sheet Anti-Histone H1 (TriMethyl K25) Antibody Catalog # Source Reactivity Applications CPA9312 Rabbit H WB, IH Description Rabbit polyclonal to Histone H1 (TriMethyl K25) Immunogen KLH-conjugated synthetic peptide encompassing a sequence of human Histone H1. The exact sequence is proprietary. Purification The antibody was purified by affinity chromatography. Specificity Recognizes endogenous levels of Histone H1 (TriMethyl K25) protein. Clonality Polyclonal Form Liquid in 0.42% Potassium phosphate, 0.87% Sodium chloride, pH 7.3, 30% glycerol, and 0.01% sodium azide. Dilution WB (1/1000 - 1/2000), IH (1/200 - 1/500) Gene Symbol HIST1H1A; HIST1H1C; HIST1H1D; HIST1H1E Alternative Names HIST1H1A; H1F1; Histone H1.1; Histone H1a; HIST1H1C; H1F2; Histone H1.2; Histone H1c; Histone H1d; Histone H1s-1; HIST1H1D; H1F3; Histone H1.3; Histone H1c; Histone H1s-2; HIST1H1E; H1F4; Histone H1.4; Histone H1b; Histone H1s-4 Entrez Gene 3024, 3006, 3007, 3008 (Human) SwissProt Q02539, P16403, P16402, P10412 (Human) Storage/Stability Shipped at 4°C. Upon delivery aliquot and store at -20°C for one year. Avoid freeze/thaw cycles. Application key: E- ELISA, WB- Western blot, IH- Immunohistochemistry, IF- Immunofluorescence, FC- Flow cytometry, IC- Immunocytochemistry, IP- Immunoprecipitation, ChIP- Chromatin Immunoprecipitation, EMSA- Electrophoretic Mobility Shift Assay, BL- Blocking, SE- Sandwich ELISA, CBE- Cell-based ELISA, RNAi- RNA interference Species reactivity key: H- Human, M- Mouse, R- Rat, B- Bovine, C- Chicken, D- Dog, G- Goat, Mk- Monkey, P- Pig, Rb- Rabbit, S- Sheep, Z- Zebrafish COHESION BIOSCIENCES LIMITED WEB ORDER SUPPORT CUSTOM www.cohesionbio.com [email protected] [email protected] [email protected] For research purposes only, not for human use Product Data Sheet Western blot analysis of Histone H1 (TriMethyl K25) expression in Hela (A) whole cell lysates. -

Linkages Between Changes in the 3D Organization of the Genome and Transcription During Myotube Differentiation in Vitro Malina D
Doynova et al. Skeletal Muscle (2017) 7:5 DOI 10.1186/s13395-017-0122-1 RESEARCH Open Access Linkages between changes in the 3D organization of the genome and transcription during myotube differentiation in vitro Malina D. Doynova1, James F. Markworth1, David Cameron-Smith1, Mark H. Vickers1 and Justin M. O’Sullivan1,2* Abstract Background: The spatial organization of eukaryotic genomes facilitates and reflects the underlying nuclear processes that are occurring in the cell. As such, the spatial organization of a genome represents a window on the genome biology that enables analysis of the nuclear regulatory processes that contribute to mammalian development. Methods: In this study, Hi-C and RNA-seq were used to capture the genome organization and transcriptome in mouse muscle progenitor cells (C2C12 myoblasts) before and after differentiation to myotubes, in the presence or absence of the cytidine analogue AraC. Results: We observed significant local and global developmental changes despite high levels of correlation between the myotubes and myoblast genomes. Notably, the genes that exhibited the greatest variation in transcript levels between the different developmental stages were predominately within the euchromatic compartment. There was significant re-structuring and changes in the expression of replication-dependent histone variants within the HIST1 locus. Finally, treating terminally differentiated myotubes with AraC resulted in additional changes to the transcriptome and 3D genome organization of sets of genes that were all involved in pyroptosis. Conclusions: Collectively, our results provide evidence for muscle cell-specific responses to developmental and environmental stimuli mediated through a chromatin structure mechanism. Keywords: Muscle, Genome organization, Development, Hi-C, Transcriptome Background Muscle differentiation and cell cycle arrest are tightly Skeletal muscle development (myogenesis) is a complex, regulated and highly interdependent. -

Molecular Characterization of Triple Negative Breast Cancer Formaldehyde-Fixed Paraffin-Embedded Samples by Data-Independent
bioRxiv preprint doi: https://doi.org/10.1101/2020.09.21.306654; this version posted September 24, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Molecular characterization of triple negative breast cancer formaldehyde-fixed paraffin-embedded samples by data-independent acquisition proteomics Silvia García-Adrián1¶, Lucía Trilla-Fuertes2¶, Angelo Gámez-Pozo3, Cristina Chiva4,5, Rocío López-Vacas3, Elena López-Camacho2, Guillermo Prado-Vázquez2, Andrea Zapater-Moros2, María I. Lumbreras-Herrera2, David Hardisson6,7,8, Laura Yébenes6, Pilar Zamora7,9,10, Eduard Sabidó4,5, Juan Ángel Fresno Vara3,7*, Enrique Espinosa7,9,10* 1 Medical Oncology Service, Hospital Universitario de Móstoles, Madrid, Spain 2 Biomedica Molecular Medicine SL, Madrid, Spain 3 Molecular Oncology & Pathology Lab, Institute of Medical and Molecular Genetics-INGEMM, La Paz University Hospital-IdiPAZ, Madrid, Spain 4Proteomics Unit, Center for Genomics Regulation, Barcelona Institute of Science and Technology (BIST), Barcelona, Spain 5 Proteomics Unit, Universitat Pompeu Fabra, Barcelona, Spain 6Molecular Pathology and Therapeutic Targets Group, La Paz University Hospital (IdiPAZ) 7 Biomedical Research Networking Center on Oncology-CIBERONC, ISCIII, Madrid, Spain 8 Faculty of Medicine, Universidad Autónoma de Madrid, Madrid, Spain 9Medical Oncology Service, La Paz University Hospital-IdiPAZ, Madrid, Spain 10 Cátedra UAM-Amgen, Universidad Autónoma de Madrid, Madrid, Spain ¶ These authors contributed equally to this work *Corresponding authors: JAFV [email protected]; EE [email protected] bioRxiv preprint doi: https://doi.org/10.1101/2020.09.21.306654; this version posted September 24, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. -

Clinical Implications of Chromatin Accessibility in Human Cancers
www.oncotarget.com Oncotarget, 2020, Vol. 11, (No. 18), pp: 1666-1678 Research Paper Clinical implications of chromatin accessibility in human cancers Yuexin Liu1 1Department of Bioinformatics and Computational Biology, The University of Texas MD Anderson Cancer Center, Houston, Texas, USA Correspondence to: Yuexin Liu, email: [email protected] Keywords: ATAC-seq; chromatin accessibility; TCGA; promoter; survival Received: September 16, 2019 Accepted: April 03, 2020 Published: May 05, 2020 Copyright: Liu. This is an open-access article distributed under the terms of the Creative Commons Attribution License 3.0 (CC BY 3.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited. ABSTRACT Assay for transposase-accessible chromatin using sequencing (ATAC-seq) has not yet been widely used in cancer research. Clinical implications of chromatin accessibility assessed by ATAC-seq profiling in human cancers especially in a large patient cohort is largely unknown. In this study, we analyzed ATAC-seq data in 404 cancer patients from the Cancer Genome Atlas, representing the largest cancer patient cohort with ATAC-seq data, and correlated chromatin accessibility with patient demographics, tumor histology, molecular subtypes, and survival. Our results showed that chromatin accessibility varies from chromosome to chromosome, and is different in different genomic regions along the same chromosome. Chromatin accessibility especially on the X chromosome is strongly dependent on patient sex, but not much on patient age or tumor stage. Striking difference in chromatin accessibility is observed between lung adenocarcinoma and lung squamous cell carcinoma, the two most common histological subgroups in lung cancer.