B4GALT2 Antibody Cat
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

B4GALT2 Rabbit Pab
Leader in Biomolecular Solutions for Life Science B4GALT2 Rabbit pAb Catalog No.: A17573 Basic Information Background Catalog No. This gene is one of seven beta-1,4-galactosyltransferase (beta4GalT) genes. They A17573 encode type II membrane-bound glycoproteins that appear to have exclusive specificity for the donor substrate UDP-galactose; all transfer galactose in a beta1,4 linkage to Observed MW similar acceptor sugars: GlcNAc, Glc, and Xyl. Each beta4GalT has a distinct function in 42kDa the biosynthesis of different glycoconjugates and saccharide structures. As type II membrane proteins, they have an N-terminal hydrophobic signal sequence that directs Calculated MW the protein to the Golgi apparatus and which then remains uncleaved to function as a transmembrane anchor. By sequence similarity, the beta4GalTs form four groups: Category beta4GalT1 and beta4GalT2, beta4GalT3 and beta4GalT4, beta4GalT5 and beta4GalT6, and beta4GalT7. The enzyme encoded by this gene synthesizes N-acetyllactosamine in Primary antibody glycolipids and glycoproteins. Its substrate specificity is affected by alpha-lactalbumin but it is not expressed in lactating mammary tissue. Three transcript variants encoding Applications two different isoforms have been found for this gene. [provided by RefSeq, Jul 2011] WB,IHC Cross-Reactivity Human, Mouse, Rat Recommended Dilutions Immunogen Information WB 1:500 - 1:2000 Gene ID Swiss Prot 8704 O60909 IHC 1:100 - 1:200 Immunogen Recombinant fusion protein containing a sequence corresponding to amino acids 155-271 of human B4GALT2 (NP_085076.2). Synonyms B4Gal-T2;B4Gal-T3;beta4Gal-T2;B4GALT2 Contact Product Information 400-999-6126 Source Isotype Purification Rabbit IgG Affinity purification [email protected] www.abclonal.com.cn Storage Store at -20℃. -

B4GALT3 (B4GALT2) (NM 030587) Human Tagged ORF Clone Product Data
OriGene Technologies, Inc. 9620 Medical Center Drive, Ste 200 Rockville, MD 20850, US Phone: +1-888-267-4436 [email protected] EU: [email protected] CN: [email protected] Product datasheet for RC233988 B4GALT3 (B4GALT2) (NM_030587) Human Tagged ORF Clone Product data: Product Type: Expression Plasmids Product Name: B4GALT3 (B4GALT2) (NM_030587) Human Tagged ORF Clone Tag: Myc-DDK Symbol: B4GALT2 Synonyms: B4Gal-T2; B4Gal-T3; beta4Gal-T2 Vector: pCMV6-Entry (PS100001) E. coli Selection: Kanamycin (25 ug/mL) Cell Selection: Neomycin ORF Nucleotide >RC233988 representing NM_030587 Sequence: Red=Cloning site Blue=ORF Green=Tags(s) TTTTGTAATACGACTCACTATAGGGCGGCCGGGAATTCGTCGACTGGATCCGGTACCGAGGAGATCTGCC GCCGCGATCGCC ATGGCTGTGGAAGTCCAGGAGCAGTGGCCTTGTTTGCCAGCAGCCGGATGCCCGGGCCCACTGGGCGGGC CAGTGGCCGCCTGCGGGATGAGCAGACTGCTGGGGGGGACGCTGGAGCGCGTCTGCAAGGCTGTGCTCCT TCTCTGCCTGCTGCACTTCCTCGTGGCCGTCATCCTCTACTTTGACGTCTACGCCCAGCACCTGGCCTTC TTCAGCCGCTTCAGTGCCCGAGGCCCTGCCCATGCCCTCCACCCAGCTGCTAGCAGCAGCAGCAGCAGCA GCAACTGCTCCCGGCCCAACGCCACCGCCTCTAGCTCCGGGCTCCCTGAGGTCCCCAGTGCCCTGCCCGG TCCCACGGCTCCCACGCTGCCACCCTGTCCTGACTCGCCACCTGGTCTTGTGGGCAGACTGCTGATCGAG TTCACCTCACCCATGCCCCTGGAGCGGGTGCAGAGGGAGAACCCAGGCGTGCTCATGGGCGGCCGATACA CACCGCCCGACTGCACCCCAGCCCAGACGGTGGCGGTCATCATCCCCTTTAGACACCGGGAACACCACCT GCGCTACTGGCTCCACTATCTACACCCCATCTTGAGGCGGCAGCGGCTGCGCTACGGCGTCTATGTCATC AACCAGCATGGTGAGGACACCTTCAACCGGGCCAAGCTGCTTAACGTGGGCTTCCTAGAGGCGCTGAAGG AGGATGCCGCCTATGACTGCTTCATCTTCAGCGATGTGGACCTGGTCCCCATGGATGACCGCAACCTATA CCGCTGCGGCGACCAACCCCGCCACTTTGCCATTGCCATGGACAAGTTTGGCTTCCGGCTTCCCTATGCT -

Multiplexed Engineering Glycosyltransferase Genes in CHO Cells Via Targeted Integration for Producing Antibodies with Diverse Complex‑Type N‑Glycans Ngan T
www.nature.com/scientificreports OPEN Multiplexed engineering glycosyltransferase genes in CHO cells via targeted integration for producing antibodies with diverse complex‑type N‑glycans Ngan T. B. Nguyen, Jianer Lin, Shi Jie Tay, Mariati, Jessna Yeo, Terry Nguyen‑Khuong & Yuansheng Yang* Therapeutic antibodies are decorated with complex‑type N‑glycans that signifcantly afect their biodistribution and bioactivity. The N‑glycan structures on antibodies are incompletely processed in wild‑type CHO cells due to their limited glycosylation capacity. To improve N‑glycan processing, glycosyltransferase genes have been traditionally overexpressed in CHO cells to engineer the cellular N‑glycosylation pathway by using random integration, which is often associated with large clonal variations in gene expression levels. In order to minimize the clonal variations, we used recombinase‑mediated‑cassette‑exchange (RMCE) technology to overexpress a panel of 42 human glycosyltransferase genes to screen their impact on antibody N‑linked glycosylation. The bottlenecks in the N‑glycosylation pathway were identifed and then released by overexpressing single or multiple critical genes. Overexpressing B4GalT1 gene alone in the CHO cells produced antibodies with more than 80% galactosylated bi‑antennary N‑glycans. Combinatorial overexpression of B4GalT1 and ST6Gal1 produced antibodies containing more than 70% sialylated bi‑antennary N‑glycans. In addition, antibodies with various tri‑antennary N‑glycans were obtained for the frst time by overexpressing MGAT5 alone or in combination with B4GalT1 and ST6Gal1. The various N‑glycan structures and the method for producing them in this work provide opportunities to study the glycan structure‑and‑function and develop novel recombinant antibodies for addressing diferent therapeutic applications. -
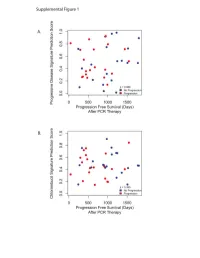
Supplementary Data
Progressive Disease Signature Upregulated probes with progressive disease U133Plus2 ID Gene Symbol Gene Name 239673_at NR3C2 nuclear receptor subfamily 3, group C, member 2 228994_at CCDC24 coiled-coil domain containing 24 1562245_a_at ZNF578 zinc finger protein 578 234224_at PTPRG protein tyrosine phosphatase, receptor type, G 219173_at NA NA 218613_at PSD3 pleckstrin and Sec7 domain containing 3 236167_at TNS3 tensin 3 1562244_at ZNF578 zinc finger protein 578 221909_at RNFT2 ring finger protein, transmembrane 2 1552732_at ABRA actin-binding Rho activating protein 59375_at MYO15B myosin XVB pseudogene 203633_at CPT1A carnitine palmitoyltransferase 1A (liver) 1563120_at NA NA 1560098_at AKR1C2 aldo-keto reductase family 1, member C2 (dihydrodiol dehydrogenase 2; bile acid binding pro 238576_at NA NA 202283_at SERPINF1 serpin peptidase inhibitor, clade F (alpha-2 antiplasmin, pigment epithelium derived factor), m 214248_s_at TRIM2 tripartite motif-containing 2 204766_s_at NUDT1 nudix (nucleoside diphosphate linked moiety X)-type motif 1 242308_at MCOLN3 mucolipin 3 1569154_a_at NA NA 228171_s_at PLEKHG4 pleckstrin homology domain containing, family G (with RhoGef domain) member 4 1552587_at CNBD1 cyclic nucleotide binding domain containing 1 220705_s_at ADAMTS7 ADAM metallopeptidase with thrombospondin type 1 motif, 7 232332_at RP13-347D8.3 KIAA1210 protein 1553618_at TRIM43 tripartite motif-containing 43 209369_at ANXA3 annexin A3 243143_at FAM24A family with sequence similarity 24, member A 234742_at SIRPG signal-regulatory protein gamma -

The Molecular Signature of the Stroma Response in Prostate
RESEARCH ARTICLE The Molecular Signature of the Stroma Response in Prostate Cancer-Induced Osteoblastic Bone Metastasis Highlights Expansion of Hematopoietic and Prostate Epithelial Stem Cell Niches Berna C. O¨ zdemir1., Janine Hensel1., Chiara Secondini1, Antoinette Wetterwald1, Ruth Schwaninger1, Achim Fleischmann2, Wolfgang Raffelsberger3, OPEN ACCESS Olivier Poch4, Mauro Delorenzi5, Ramzi Temanni6, Ian G. Mills7, Gabri van der Citation: O¨ zdemir BC, Hensel J, Secondini C, Pluijm8, George N. Thalmann1, Marco G. Cecchini1* Wetterwald A, Schwaninger R, et al. (2014) The Molecular Signature of the Stroma Response in 1. Urology Research Laboratory, Department of Urology and Department of Clinical Research, University of Prostate Cancer-Induced Osteoblastic Bone Bern, Bern, Switzerland, 2. Institute of Pathology, University of Bern, Bern, Switzerland, 3. Institut Ge´ne´tique Metastasis Highlights Expansion of Hematopoietic Biologie Mole´culaire Cellulaire (IGBMC), Strasbourg, France, 4. ICube UMR7357, University of Strasbourg, and Prostate Epithelial Stem Cell Niches. PLoS Strasbourg, France, 5. Ludwig Center for Cancer Research, Department of Oncology, University of Lausanne ONE 9(12): e114530. doi:10.1371/journal.pone. and Swiss Institute of Bioinformatics (SIB), Lausanne, Switzerland, 6. Biomedical Informatics Division, Sidra 0114530 Medical and Research Center, Doha, Qatar, 7. Prostate Cancer Research Group, Norway Centre for Editor: Adriano Angelucci, University of L9Aquila, Molecular Medicine (NCMM), University of Oslo, Oslo, Norway, -

Single Cell Derived Clonal Analysis of Human Glioblastoma Links
SUPPLEMENTARY INFORMATION: Single cell derived clonal analysis of human glioblastoma links functional and genomic heterogeneity ! Mona Meyer*, Jüri Reimand*, Xiaoyang Lan, Renee Head, Xueming Zhu, Michelle Kushida, Jane Bayani, Jessica C. Pressey, Anath Lionel, Ian D. Clarke, Michael Cusimano, Jeremy Squire, Stephen Scherer, Mark Bernstein, Melanie A. Woodin, Gary D. Bader**, and Peter B. Dirks**! ! * These authors contributed equally to this work.! ** Correspondence: [email protected] or [email protected]! ! Supplementary information - Meyer, Reimand et al. Supplementary methods" 4" Patient samples and fluorescence activated cell sorting (FACS)! 4! Differentiation! 4! Immunocytochemistry and EdU Imaging! 4! Proliferation! 5! Western blotting ! 5! Temozolomide treatment! 5! NCI drug library screen! 6! Orthotopic injections! 6! Immunohistochemistry on tumor sections! 6! Promoter methylation of MGMT! 6! Fluorescence in situ Hybridization (FISH)! 7! SNP6 microarray analysis and genome segmentation! 7! Calling copy number alterations! 8! Mapping altered genome segments to genes! 8! Recurrently altered genes with clonal variability! 9! Global analyses of copy number alterations! 9! Phylogenetic analysis of copy number alterations! 10! Microarray analysis! 10! Gene expression differences of TMZ resistant and sensitive clones of GBM-482! 10! Reverse transcription-PCR analyses! 11! Tumor subtype analysis of TMZ-sensitive and resistant clones! 11! Pathway analysis of gene expression in the TMZ-sensitive clone of GBM-482! 11! Supplementary figures and tables" 13" "2 Supplementary information - Meyer, Reimand et al. Table S1: Individual clones from all patient tumors are tumorigenic. ! 14! Fig. S1: clonal tumorigenicity.! 15! Fig. S2: clonal heterogeneity of EGFR and PTEN expression.! 20! Fig. S3: clonal heterogeneity of proliferation.! 21! Fig. -

The Role of Lamin Associated Domains in Global Chromatin Organization and Nuclear Architecture
THE ROLE OF LAMIN ASSOCIATED DOMAINS IN GLOBAL CHROMATIN ORGANIZATION AND NUCLEAR ARCHITECTURE By Teresa Romeo Luperchio A dissertation submitted to The Johns Hopkins University in conformity with the requirements for the degree of Doctor of Philosophy Baltimore, Maryland March 2016 © 2016 Teresa Romeo Luperchio All Rights Reserved ABSTRACT Nuclear structure and scaffolding have been implicated in expression and regulation of the genome (Elcock and Bridger 2010; Fedorova and Zink 2008; Ferrai et al. 2010; Li and Reinberg 2011; Austin and Bellini 2010). Discrete domains of chromatin exist within the nuclear volume, and are suggested to be organized by patterns of gene activity (Zhao, Bodnar, and Spector 2009). The nuclear periphery, which consists of the inner nuclear membrane and associated proteins, forms a sub- nuclear compartment that is mostly associated with transcriptionally repressed chromatin and low gene expression (Guelen et al. 2008). Previous studies from our lab and others have shown that repositioning genes to the nuclear periphery is sufficient to induce transcriptional repression (K L Reddy et al. 2008; Finlan et al. 2008). In addition, a number of studies have provided evidence that many tissue types, including muscle, brain and blood, use the nuclear periphery as a compartment during development to regulate expression of lineage specific genes (Meister et al. 2010; Szczerbal, Foster, and Bridger 2009; Yao et al. 2011; Kosak et al. 2002; Peric-Hupkes et al. 2010). These large regions of chromatin that come in molecular contact with the nuclear periphery are called Lamin Associated Domains (LADs). The studies described in this dissertation have furthered our understanding of maintenance and establishment of LADs as well as the relationship of LADs with the epigenome and other factors that influence three-dimensional chromatin structure. -

Acetyl-Coa Flux from the Cytosol to the ER Regulates Engagement And
www.nature.com/scientificreports OPEN Acetyl‑CoA fux from the cytosol to the ER regulates engagement and quality of the secretory pathway Inca A. Dieterich1,2,3,11, Yusi Cui4,11, Megan M. Braun1,2,3, Alexis J. Lawton5,6, Nicklaus H. Robinson1,2, Jennifer L. Peotter5, Qing Yu4,10, Jason C. Casler7, Benjamin S. Glick7, Anjon Audhya5, John M. Denu5,6, Lingjun Li4* & Luigi Puglielli1,2,8,9* Nε‑lysine acetylation in the ER is an essential component of the quality control machinery. ER acetylation is ensured by a membrane transporter, AT‑1/SLC33A1, which translocates cytosolic acetyl‑ CoA into the ER lumen, and two acetyltransferases, ATase1 and ATase2, which acetylate nascent polypeptides within the ER lumen. Dysfunctional AT‑1, as caused by gene mutation or duplication events, results in severe disease phenotypes. Here, we used two models of AT‑1 dysregulation to investigate dynamics of the secretory pathway: AT‑1 sTg, a model of systemic AT‑1 overexpression, and AT‑1S113R/+, a model of AT‑1 haploinsufciency. The animals displayed reorganization of the ER, ERGIC, and Golgi apparatus. In particular, AT‑1 sTg animals displayed a marked delay in Golgi‑to‑ plasma membrane protein trafcking, signifcant alterations in Golgi‑based N‑glycan modifcation, and a marked expansion of the lysosomal network. Collectively our results indicate that AT‑1 is essential to maintain proper organization and engagement of the secretory pathway. Nε-lysine acetylation in the Endoplasmic Reticulum (ER) has emerged as an essential component of the qual- ity control (QC) machinery that maintains protein homeostasis (proteostasis) within the ER1–6. -

Vast Human-Specific Delay in Cortical Ontogenesis Associated With
Supplementary information Extension of cortical synaptic development distinguishes humans from chimpanzees and macaques Supplementary Methods Sample collection We used prefrontal cortex (PFC) and cerebellar cortex (CBC) samples from postmortem brains of 33 human (aged 0-98 years), 14 chimpanzee (aged 0-44 years) and 44 rhesus macaque individuals (aged 0-28 years) (Table S1). Human samples were obtained from the NICHD Brain and Tissue Bank for Developmental Disorders at the University of Maryland, USA, the Netherlands Brain Bank, Amsterdam, Netherlands and the Chinese Brain Bank Center, Wuhan, China. Informed consent for use of human tissues for research was obtained in writing from all donors or their next of kin. All subjects were defined as normal by forensic pathologists at the corresponding brain bank. All subjects suffered sudden death with no prolonged agonal state. Chimpanzee samples were obtained from the Yerkes Primate Center, GA, USA, the Anthropological Institute & Museum of the University of Zürich-Irchel, Switzerland and the Biomedical Primate Research Centre, Netherlands (eight Western chimpanzees, one Central/Eastern and five of unknown origin). Rhesus macaque samples were obtained from the Suzhou Experimental Animal Center, China. All non-human primates used in this study suffered sudden deaths for reasons other than their participation in this study and without any relation to the tissue used. CBC dissections were made from the cerebellar cortex. PFC dissections were made from the frontal part of the superior frontal gyrus. All samples contained an approximately 2:1 grey matter to white matter volume ratio. RNA microarray hybridization RNA isolation, hybridization to microarrays, and data preprocessing were performed as described previously (Khaitovich et al. -

Autocrine IFN Signaling Inducing Profibrotic Fibroblast Responses By
Downloaded from http://www.jimmunol.org/ by guest on September 23, 2021 Inducing is online at: average * The Journal of Immunology , 11 of which you can access for free at: 2013; 191:2956-2966; Prepublished online 16 from submission to initial decision 4 weeks from acceptance to publication August 2013; doi: 10.4049/jimmunol.1300376 http://www.jimmunol.org/content/191/6/2956 A Synthetic TLR3 Ligand Mitigates Profibrotic Fibroblast Responses by Autocrine IFN Signaling Feng Fang, Kohtaro Ooka, Xiaoyong Sun, Ruchi Shah, Swati Bhattacharyya, Jun Wei and John Varga J Immunol cites 49 articles Submit online. Every submission reviewed by practicing scientists ? is published twice each month by Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts http://jimmunol.org/subscription Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html http://www.jimmunol.org/content/suppl/2013/08/20/jimmunol.130037 6.DC1 This article http://www.jimmunol.org/content/191/6/2956.full#ref-list-1 Information about subscribing to The JI No Triage! Fast Publication! Rapid Reviews! 30 days* Why • • • Material References Permissions Email Alerts Subscription Supplementary The Journal of Immunology The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2013 by The American Association of Immunologists, Inc. All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. This information is current as of September 23, 2021. The Journal of Immunology A Synthetic TLR3 Ligand Mitigates Profibrotic Fibroblast Responses by Inducing Autocrine IFN Signaling Feng Fang,* Kohtaro Ooka,* Xiaoyong Sun,† Ruchi Shah,* Swati Bhattacharyya,* Jun Wei,* and John Varga* Activation of TLR3 by exogenous microbial ligands or endogenous injury-associated ligands leads to production of type I IFN. -

Coexpression Networks Based on Natural Variation in Human Gene Expression at Baseline and Under Stress
University of Pennsylvania ScholarlyCommons Publicly Accessible Penn Dissertations Fall 2010 Coexpression Networks Based on Natural Variation in Human Gene Expression at Baseline and Under Stress Renuka Nayak University of Pennsylvania, [email protected] Follow this and additional works at: https://repository.upenn.edu/edissertations Part of the Computational Biology Commons, and the Genomics Commons Recommended Citation Nayak, Renuka, "Coexpression Networks Based on Natural Variation in Human Gene Expression at Baseline and Under Stress" (2010). Publicly Accessible Penn Dissertations. 1559. https://repository.upenn.edu/edissertations/1559 This paper is posted at ScholarlyCommons. https://repository.upenn.edu/edissertations/1559 For more information, please contact [email protected]. Coexpression Networks Based on Natural Variation in Human Gene Expression at Baseline and Under Stress Abstract Genes interact in networks to orchestrate cellular processes. Here, we used coexpression networks based on natural variation in gene expression to study the functions and interactions of human genes. We asked how these networks change in response to stress. First, we studied human coexpression networks at baseline. We constructed networks by identifying correlations in expression levels of 8.9 million gene pairs in immortalized B cells from 295 individuals comprising three independent samples. The resulting networks allowed us to infer interactions between biological processes. We used the network to predict the functions of poorly-characterized human genes, and provided some experimental support. Examining genes implicated in disease, we found that IFIH1, a diabetes susceptibility gene, interacts with YES1, which affects glucose transport. Genes predisposing to the same diseases are clustered non-randomly in the network, suggesting that the network may be used to identify candidate genes that influence disease susceptibility. -

Assessing Mitochondrial Theory of Aging on the Transcriptome Level
Supplemental Material MicroRNA, mRNA, and Protein Expression Link Development and Aging in Human and Macaque Brain Supplemental Tables......................................................................................................... 3 Table S1. Sample characteristics (human). ......................................................................................... 3 Table S2. Sample characteristics (macaque). ...................................................................................... 4 Table S3. Proportions of transition genes in mRNA, miRNA and protein datasets. ........................... 5 Table S4. miRNA changes with age in human and mouse.................................................................. 6 Table S5. miRNA-target pairs involved in human cortex development and aging............................. 7 Table S6. miRNA-target pairs overlapping with experimentally verified target sets........................ 10 Table S7. Gene Ontology categories enriched among gene groups. ................................................. 12 Table S8. KEGG pathways enriched among gene groups................................................................. 13 Supplemental Figures ..................................................................................................... 15 Figure S1. Age effect on expression and correlation between datasets............................................. 15 Figure S2. Correlation among genes within gene groups. ................................................................ 17 Figure