Acetyl-Coa Flux from the Cytosol to the ER Regulates Engagement And
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Analysis of Trans Esnps Infers Regulatory Network Architecture
Analysis of trans eSNPs infers regulatory network architecture Anat Kreimer Submitted in partial fulfillment of the requirements for the degree of Doctor of Philosophy in the Graduate School of Arts and Sciences COLUMBIA UNIVERSITY 2014 © 2014 Anat Kreimer All rights reserved ABSTRACT Analysis of trans eSNPs infers regulatory network architecture Anat Kreimer eSNPs are genetic variants associated with transcript expression levels. The characteristics of such variants highlight their importance and present a unique opportunity for studying gene regulation. eSNPs affect most genes and their cell type specificity can shed light on different processes that are activated in each cell. They can identify functional variants by connecting SNPs that are implicated in disease to a molecular mechanism. Examining eSNPs that are associated with distal genes can provide insights regarding the inference of regulatory networks but also presents challenges due to the high statistical burden of multiple testing. Such association studies allow: simultaneous investigation of many gene expression phenotypes without assuming any prior knowledge and identification of unknown regulators of gene expression while uncovering directionality. This thesis will focus on such distal eSNPs to map regulatory interactions between different loci and expose the architecture of the regulatory network defined by such interactions. We develop novel computational approaches and apply them to genetics-genomics data in human. We go beyond pairwise interactions to define network motifs, including regulatory modules and bi-fan structures, showing them to be prevalent in real data and exposing distinct attributes of such arrangements. We project eSNP associations onto a protein-protein interaction network to expose topological properties of eSNPs and their targets and highlight different modes of distal regulation. -

Seq2pathway Vignette
seq2pathway Vignette Bin Wang, Xinan Holly Yang, Arjun Kinstlick May 19, 2021 Contents 1 Abstract 1 2 Package Installation 2 3 runseq2pathway 2 4 Two main functions 3 4.1 seq2gene . .3 4.1.1 seq2gene flowchart . .3 4.1.2 runseq2gene inputs/parameters . .5 4.1.3 runseq2gene outputs . .8 4.2 gene2pathway . 10 4.2.1 gene2pathway flowchart . 11 4.2.2 gene2pathway test inputs/parameters . 11 4.2.3 gene2pathway test outputs . 12 5 Examples 13 5.1 ChIP-seq data analysis . 13 5.1.1 Map ChIP-seq enriched peaks to genes using runseq2gene .................... 13 5.1.2 Discover enriched GO terms using gene2pathway_test with gene scores . 15 5.1.3 Discover enriched GO terms using Fisher's Exact test without gene scores . 17 5.1.4 Add description for genes . 20 5.2 RNA-seq data analysis . 20 6 R environment session 23 1 Abstract Seq2pathway is a novel computational tool to analyze functional gene-sets (including signaling pathways) using variable next-generation sequencing data[1]. Integral to this tool are the \seq2gene" and \gene2pathway" components in series that infer a quantitative pathway-level profile for each sample. The seq2gene function assigns phenotype-associated significance of genomic regions to gene-level scores, where the significance could be p-values of SNPs or point mutations, protein-binding affinity, or transcriptional expression level. The seq2gene function has the feasibility to assign non-exon regions to a range of neighboring genes besides the nearest one, thus facilitating the study of functional non-coding elements[2]. Then the gene2pathway summarizes gene-level measurements to pathway-level scores, comparing the quantity of significance for gene members within a pathway with those outside a pathway. -

Dual Proteome-Scale Networks Reveal Cell-Specific Remodeling of the Human Interactome
bioRxiv preprint doi: https://doi.org/10.1101/2020.01.19.905109; this version posted January 19, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Dual Proteome-scale Networks Reveal Cell-specific Remodeling of the Human Interactome Edward L. Huttlin1*, Raphael J. Bruckner1,3, Jose Navarrete-Perea1, Joe R. Cannon1,4, Kurt Baltier1,5, Fana Gebreab1, Melanie P. Gygi1, Alexandra Thornock1, Gabriela Zarraga1,6, Stanley Tam1,7, John Szpyt1, Alexandra Panov1, Hannah Parzen1,8, Sipei Fu1, Arvene Golbazi1, Eila Maenpaa1, Keegan Stricker1, Sanjukta Guha Thakurta1, Ramin Rad1, Joshua Pan2, David P. Nusinow1, Joao A. Paulo1, Devin K. Schweppe1, Laura Pontano Vaites1, J. Wade Harper1*, Steven P. Gygi1*# 1Department of Cell Biology, Harvard Medical School, Boston, MA, 02115, USA. 2Broad Institute, Cambridge, MA, 02142, USA. 3Present address: ICCB-Longwood Screening Facility, Harvard Medical School, Boston, MA, 02115, USA. 4Present address: Merck, West Point, PA, 19486, USA. 5Present address: IQ Proteomics, Cambridge, MA, 02139, USA. 6Present address: Vor Biopharma, Cambridge, MA, 02142, USA. 7Present address: Rubius Therapeutics, Cambridge, MA, 02139, USA. 8Present address: RPS North America, South Kingstown, RI, 02879, USA. *Correspondence: [email protected] (E.L.H.), [email protected] (J.W.H.), [email protected] (S.P.G.) #Lead Contact: [email protected] bioRxiv preprint doi: https://doi.org/10.1101/2020.01.19.905109; this version posted January 19, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. -

Glucose-Induced Changes in Gene Expression in Human Pancreatic Islets: Causes Or Consequences of Chronic Hyperglycemia
Diabetes Volume 66, December 2017 3013 Glucose-Induced Changes in Gene Expression in Human Pancreatic Islets: Causes or Consequences of Chronic Hyperglycemia Emilia Ottosson-Laakso,1 Ulrika Krus,1 Petter Storm,1 Rashmi B. Prasad,1 Nikolay Oskolkov,1 Emma Ahlqvist,1 João Fadista,2 Ola Hansson,1 Leif Groop,1,3 and Petter Vikman1 Diabetes 2017;66:3013–3028 | https://doi.org/10.2337/db17-0311 Dysregulation of gene expression in islets from patients In patients with type 2 diabetes (T2D), islet function de- with type 2 diabetes (T2D) might be causally involved clines progressively. Although the initial pathogenic trigger in the development of hyperglycemia, or it could develop of impaired b-cell function is still unknown, elevated glu- as a consequence of hyperglycemia (i.e., glucotoxicity). cose levels are known to further aggravate b-cell function, a To separate the genes that could be causally involved condition referred to as glucotoxicity, which can stimulate in pathogenesis from those likely to be secondary to hy- apoptosis and lead to reduced b-cell mass (1–5). Prolonged perglycemia, we exposed islets from human donors to exposure to hyperglycemia also can induce endoplasmic re- ISLET STUDIES normal or high glucose concentrations for 24 h and ana- ticulum (ER) stress and production of reactive oxygen spe- fi lyzed gene expression. We compared these ndings with cies (6), which can further impair islet function and thereby gene expression in islets from donors with normal glucose the ability of islets to secrete the insulin needed to meet the tolerance and hyperglycemia (including T2D). The genes increased demands imposed by insulin resistance and obe- whose expression changed in the same direction after sity (7). -

B4GALT2 Rabbit Pab
Leader in Biomolecular Solutions for Life Science B4GALT2 Rabbit pAb Catalog No.: A17573 Basic Information Background Catalog No. This gene is one of seven beta-1,4-galactosyltransferase (beta4GalT) genes. They A17573 encode type II membrane-bound glycoproteins that appear to have exclusive specificity for the donor substrate UDP-galactose; all transfer galactose in a beta1,4 linkage to Observed MW similar acceptor sugars: GlcNAc, Glc, and Xyl. Each beta4GalT has a distinct function in 42kDa the biosynthesis of different glycoconjugates and saccharide structures. As type II membrane proteins, they have an N-terminal hydrophobic signal sequence that directs Calculated MW the protein to the Golgi apparatus and which then remains uncleaved to function as a transmembrane anchor. By sequence similarity, the beta4GalTs form four groups: Category beta4GalT1 and beta4GalT2, beta4GalT3 and beta4GalT4, beta4GalT5 and beta4GalT6, and beta4GalT7. The enzyme encoded by this gene synthesizes N-acetyllactosamine in Primary antibody glycolipids and glycoproteins. Its substrate specificity is affected by alpha-lactalbumin but it is not expressed in lactating mammary tissue. Three transcript variants encoding Applications two different isoforms have been found for this gene. [provided by RefSeq, Jul 2011] WB,IHC Cross-Reactivity Human, Mouse, Rat Recommended Dilutions Immunogen Information WB 1:500 - 1:2000 Gene ID Swiss Prot 8704 O60909 IHC 1:100 - 1:200 Immunogen Recombinant fusion protein containing a sequence corresponding to amino acids 155-271 of human B4GALT2 (NP_085076.2). Synonyms B4Gal-T2;B4Gal-T3;beta4Gal-T2;B4GALT2 Contact Product Information 400-999-6126 Source Isotype Purification Rabbit IgG Affinity purification [email protected] www.abclonal.com.cn Storage Store at -20℃. -

Murine SEC24D Can Substitute Functionally for SEC24C in Vivo
bioRxiv preprint doi: https://doi.org/10.1101/284398; this version posted March 22, 2018. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Functional Overlap Between Mouse SEC24C and SEC24D Murine SEC24D Can Substitute Functionally for SEC24C in vivo Elizabeth J. Adams1,2, Rami Khoriaty2,3, Anna Kiseleva1, Audrey C. A. Cleuren1,7, Kärt Tomberg1,4, Martijn A. van der Ent3, Peter Gergics4, K. Sue O’Shea5, Thomas L. Saunders3, David Ginsburg1-4,6,7 1From the Life Sciences Institute, 2Program in Cellular and Molecular Biology, 3Department of Internal Medicine, 4Departement of Human Genetics, 5Department of Cell and Developmental Biology, 6Department of Pediatrics and the 7Howard Hughes Medical Institute, University of Michigan, Ann Arbor, MI 48109 ABSTRACT The COPII component SEC24 mediates the recruitment of transmembrane cargoes or cargo adaptors into newly forming COPII vesicles on the ER membrane. Mammalian genomes encode four Sec24 paralogs (Sec24a-d), with two subfamilies based on sequence homology (SEC24A/B and C/D), though little is known about their comparative functions and cargo-specificities. Complete deficiency for Sec24d results in very early embryonic lethality in mice (before the 8 cell stage), with later embryonic lethality (E 7.5) observed in Sec24c null mice. To test the potential overlap in function between SEC24C/D, we employed dual recombinase mediated cassette exchange to generate a Sec24cc-d allele, in which the C-terminal 90% of SEC24C has been replaced by SEC24D coding sequence. In contrast to the embryonic lethality at E7.5 of SEC24C-deficiency, Sec24cc-d/c-d pups survive to term, though dying shortly after birth. -

Análise Integrativa De Perfis Transcricionais De Pacientes Com
UNIVERSIDADE DE SÃO PAULO FACULDADE DE MEDICINA DE RIBEIRÃO PRETO PROGRAMA DE PÓS-GRADUAÇÃO EM GENÉTICA ADRIANE FEIJÓ EVANGELISTA Análise integrativa de perfis transcricionais de pacientes com diabetes mellitus tipo 1, tipo 2 e gestacional, comparando-os com manifestações demográficas, clínicas, laboratoriais, fisiopatológicas e terapêuticas Ribeirão Preto – 2012 ADRIANE FEIJÓ EVANGELISTA Análise integrativa de perfis transcricionais de pacientes com diabetes mellitus tipo 1, tipo 2 e gestacional, comparando-os com manifestações demográficas, clínicas, laboratoriais, fisiopatológicas e terapêuticas Tese apresentada à Faculdade de Medicina de Ribeirão Preto da Universidade de São Paulo para obtenção do título de Doutor em Ciências. Área de Concentração: Genética Orientador: Prof. Dr. Eduardo Antonio Donadi Co-orientador: Prof. Dr. Geraldo A. S. Passos Ribeirão Preto – 2012 AUTORIZO A REPRODUÇÃO E DIVULGAÇÃO TOTAL OU PARCIAL DESTE TRABALHO, POR QUALQUER MEIO CONVENCIONAL OU ELETRÔNICO, PARA FINS DE ESTUDO E PESQUISA, DESDE QUE CITADA A FONTE. FICHA CATALOGRÁFICA Evangelista, Adriane Feijó Análise integrativa de perfis transcricionais de pacientes com diabetes mellitus tipo 1, tipo 2 e gestacional, comparando-os com manifestações demográficas, clínicas, laboratoriais, fisiopatológicas e terapêuticas. Ribeirão Preto, 2012 192p. Tese de Doutorado apresentada à Faculdade de Medicina de Ribeirão Preto da Universidade de São Paulo. Área de Concentração: Genética. Orientador: Donadi, Eduardo Antonio Co-orientador: Passos, Geraldo A. 1. Expressão gênica – microarrays 2. Análise bioinformática por module maps 3. Diabetes mellitus tipo 1 4. Diabetes mellitus tipo 2 5. Diabetes mellitus gestacional FOLHA DE APROVAÇÃO ADRIANE FEIJÓ EVANGELISTA Análise integrativa de perfis transcricionais de pacientes com diabetes mellitus tipo 1, tipo 2 e gestacional, comparando-os com manifestações demográficas, clínicas, laboratoriais, fisiopatológicas e terapêuticas. -

Supplementary Table 2 Gene Sets Used in GSEA
Supplementary Table 2 Gene sets used in GSEA Up in RNAi and Sign Confirmed in Inducible Gene Probe Set ID Accession Symbol Gene Title 200660_at NM_005620 S100A11 S100 calcium binding protein A11 (calgizzarin) 200785_s_at NM_002332 LRP1 low density lipoprotein-related protein 1 (alpha-2-macroglobulin receptor) 201325_s_at NM_001423 EMP1 epithelial membrane protein 1 201373_at NM_000445 PLEC1 plectin 1, intermediate filament binding protein 500kDa 201466_s_at NM_002228 JUN v-jun sarcoma virus 17 oncogene homolog (avian) 201952_at AA156721 ALCAM activated leukocyte cell adhesion molecule 202042_at NM_002109 HARS histidyl-tRNA synthetase 202074_s_at NM_021980 OPTN optineurin 202087_s_at NM_001912 CTSL cathepsin L 202588_at NM_000476 AK1 adenylate kinase 1 202609_at NM_004447 EPS8 epidermal growth factor receptor pathway substrate 8 202733_at NM_004199 P4HA2 procollagen-proline, 2-oxoglutarate 4-dioxygenase (proline 4-hydroxylase), alpha polypeptide II 202756_s_at NM_002081 GPC1 glypican 1 202786_at NM_013233 STK39 serine threonine kinase 39 (STE20/SPS1 homolog, yeast) 202859_x_at NM_000584 IL8 interleukin 8 203083_at NM_003247 THBS2 thrombospondin 2 203186_s_at NM_002961 S100A4 S100 calcium binding protein A4 (calcium protein, calvasculin, metastasin, murine placental homolog) 203232_s_at NM_000332 ATXN1 ataxin 1 203233_at NM_000418 IL4R interleukin 4 receptor 203771_s_at AA740186 BLVRA biliverdin reductase A 203821_at NM_001945 HBEGF heparin-binding EGF-like growth factor 203939_at NM_002526 NT5E 5'-nucleotidase, ecto (CD73) 203955_at NM_014811 -
Download Validation Data
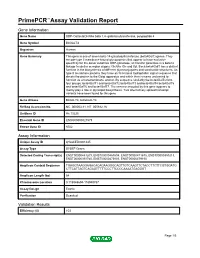
PrimePCR™Assay Validation Report Gene Information Gene Name UDP-Gal:betaGlcNAc beta 1,4- galactosyltransferase, polypeptide 4 Gene Symbol B4GALT4 Organism Human Gene Summary This gene is one of seven beta-14-galactosyltransferase (beta4GalT) genes. They encode type II membrane-bound glycoproteins that appear to have exclusive specificity for the donor substrate UDP-galactose; all transfer galactose in a beta14 linkage to similar acceptor sugars: GlcNAc Glc and Xyl. Each beta4GalT has a distinct function in the biosynthesis of different glycoconjugates and saccharide structures. As type II membrane proteins they have an N-terminal hydrophobic signal sequence that directs the protein to the Golgi apparatus and which then remains uncleaved to function as a transmembrane anchor. By sequence similarity the beta4GalTs form four groups: beta4GalT1 and beta4GalT2 beta4GalT3 and beta4GalT4 beta4GalT5 and beta4GalT6 and beta4GalT7. The enzyme encoded by this gene appears to mainly play a role in glycolipid biosynthesis. Two alternatively spliced transcript variants have been found for this gene. Gene Aliases B4Gal-T4, beta4Gal-T4 RefSeq Accession No. NC_000003.11, NT_005612.16 UniGene ID Hs.13225 Ensembl Gene ID ENSG00000121578 Entrez Gene ID 8702 Assay Information Unique Assay ID qHsaCED0001445 Assay Type SYBR® Green Detected Coding Transcript(s) ENST00000483209, ENST00000467604, ENST00000471675, ENST00000359213, ENST00000393765, ENST00000475803, ENST00000479150 Amplicon Context Sequence TGAGGTAAGGAGACACAGAAGGGCAGTTGTCAAGTTCTACCTTCTTCGTGGATG CTTCATTAGTCAGAGTTTTTCCCTTCCCCAAAATGAGGGT Amplicon Length (bp) 64 Chromosome Location 3:118948694-118948787 Assay Design Exonic Purification Desalted Validation Results Efficiency (%) 103 Page 1/5 PrimePCR™Assay Validation Report R2 0.9998 cDNA Cq 21.23 cDNA Tm (Celsius) 78 gDNA Cq 23.9 Specificity (%) 100 Information to assist with data interpretation is provided at the end of this report. -

Discovery of the First Genome-Wide Significant Risk Loci for Attention Deficit/Hyperactivity Disorder
ARTICLES https://doi.org/10.1038/s41588-018-0269-7 Discovery of the first genome-wide significant risk loci for attention deficit/hyperactivity disorder Ditte Demontis 1,2,3,69, Raymond K. Walters 4,5,69, Joanna Martin5,6,7, Manuel Mattheisen1,2,3,8,9,10, Thomas D. Als 1,2,3, Esben Agerbo 1,11,12, Gísli Baldursson13, Rich Belliveau5, Jonas Bybjerg-Grauholm 1,14, Marie Bækvad-Hansen1,14, Felecia Cerrato5, Kimberly Chambert5, Claire Churchhouse4,5,15, Ashley Dumont5, Nicholas Eriksson16, Michael Gandal17,18,19,20, Jacqueline I. Goldstein4,5,15, Katrina L. Grasby21, Jakob Grove 1,2,3,22, Olafur O. Gudmundsson13,23,24, Christine S. Hansen 1,14,25, Mads Engel Hauberg1,2,3, Mads V. Hollegaard1,14, Daniel P. Howrigan4,5, Hailiang Huang4,5, Julian B. Maller5,26, Alicia R. Martin4,5,15, Nicholas G. Martin 21, Jennifer Moran5, Jonatan Pallesen1,2,3, Duncan S. Palmer4,5, Carsten Bøcker Pedersen1,11,12, Marianne Giørtz Pedersen1,11,12, Timothy Poterba4,5,15, Jesper Buchhave Poulsen1,14, Stephan Ripke4,5,27, Elise B. Robinson4,28, F. Kyle Satterstrom 4,5,15, Hreinn Stefansson 23, Christine Stevens5, Patrick Turley4,5, G. Bragi Walters 23,24, Hyejung Won 17,18, Margaret J. Wright 29, ADHD Working Group of the Psychiatric Genomics Consortium (PGC)30, Early Lifecourse & Genetic Epidemiology (EAGLE) Consortium30, 23andMe Research Team30, Ole A. Andreassen 31, Philip Asherson32, Christie L. Burton 33, Dorret I. Boomsma34,35, Bru Cormand36,37,38,39, Søren Dalsgaard 11, Barbara Franke40, Joel Gelernter 41,42, Daniel Geschwind 17,18,19, Hakon Hakonarson43, Jan Haavik44,45, Henry R. -

B4GALT3 (B4GALT2) (NM 030587) Human Tagged ORF Clone Product Data
OriGene Technologies, Inc. 9620 Medical Center Drive, Ste 200 Rockville, MD 20850, US Phone: +1-888-267-4436 [email protected] EU: [email protected] CN: [email protected] Product datasheet for RC233988 B4GALT3 (B4GALT2) (NM_030587) Human Tagged ORF Clone Product data: Product Type: Expression Plasmids Product Name: B4GALT3 (B4GALT2) (NM_030587) Human Tagged ORF Clone Tag: Myc-DDK Symbol: B4GALT2 Synonyms: B4Gal-T2; B4Gal-T3; beta4Gal-T2 Vector: pCMV6-Entry (PS100001) E. coli Selection: Kanamycin (25 ug/mL) Cell Selection: Neomycin ORF Nucleotide >RC233988 representing NM_030587 Sequence: Red=Cloning site Blue=ORF Green=Tags(s) TTTTGTAATACGACTCACTATAGGGCGGCCGGGAATTCGTCGACTGGATCCGGTACCGAGGAGATCTGCC GCCGCGATCGCC ATGGCTGTGGAAGTCCAGGAGCAGTGGCCTTGTTTGCCAGCAGCCGGATGCCCGGGCCCACTGGGCGGGC CAGTGGCCGCCTGCGGGATGAGCAGACTGCTGGGGGGGACGCTGGAGCGCGTCTGCAAGGCTGTGCTCCT TCTCTGCCTGCTGCACTTCCTCGTGGCCGTCATCCTCTACTTTGACGTCTACGCCCAGCACCTGGCCTTC TTCAGCCGCTTCAGTGCCCGAGGCCCTGCCCATGCCCTCCACCCAGCTGCTAGCAGCAGCAGCAGCAGCA GCAACTGCTCCCGGCCCAACGCCACCGCCTCTAGCTCCGGGCTCCCTGAGGTCCCCAGTGCCCTGCCCGG TCCCACGGCTCCCACGCTGCCACCCTGTCCTGACTCGCCACCTGGTCTTGTGGGCAGACTGCTGATCGAG TTCACCTCACCCATGCCCCTGGAGCGGGTGCAGAGGGAGAACCCAGGCGTGCTCATGGGCGGCCGATACA CACCGCCCGACTGCACCCCAGCCCAGACGGTGGCGGTCATCATCCCCTTTAGACACCGGGAACACCACCT GCGCTACTGGCTCCACTATCTACACCCCATCTTGAGGCGGCAGCGGCTGCGCTACGGCGTCTATGTCATC AACCAGCATGGTGAGGACACCTTCAACCGGGCCAAGCTGCTTAACGTGGGCTTCCTAGAGGCGCTGAAGG AGGATGCCGCCTATGACTGCTTCATCTTCAGCGATGTGGACCTGGTCCCCATGGATGACCGCAACCTATA CCGCTGCGGCGACCAACCCCGCCACTTTGCCATTGCCATGGACAAGTTTGGCTTCCGGCTTCCCTATGCT -

Multiplexed Engineering Glycosyltransferase Genes in CHO Cells Via Targeted Integration for Producing Antibodies with Diverse Complex‑Type N‑Glycans Ngan T
www.nature.com/scientificreports OPEN Multiplexed engineering glycosyltransferase genes in CHO cells via targeted integration for producing antibodies with diverse complex‑type N‑glycans Ngan T. B. Nguyen, Jianer Lin, Shi Jie Tay, Mariati, Jessna Yeo, Terry Nguyen‑Khuong & Yuansheng Yang* Therapeutic antibodies are decorated with complex‑type N‑glycans that signifcantly afect their biodistribution and bioactivity. The N‑glycan structures on antibodies are incompletely processed in wild‑type CHO cells due to their limited glycosylation capacity. To improve N‑glycan processing, glycosyltransferase genes have been traditionally overexpressed in CHO cells to engineer the cellular N‑glycosylation pathway by using random integration, which is often associated with large clonal variations in gene expression levels. In order to minimize the clonal variations, we used recombinase‑mediated‑cassette‑exchange (RMCE) technology to overexpress a panel of 42 human glycosyltransferase genes to screen their impact on antibody N‑linked glycosylation. The bottlenecks in the N‑glycosylation pathway were identifed and then released by overexpressing single or multiple critical genes. Overexpressing B4GalT1 gene alone in the CHO cells produced antibodies with more than 80% galactosylated bi‑antennary N‑glycans. Combinatorial overexpression of B4GalT1 and ST6Gal1 produced antibodies containing more than 70% sialylated bi‑antennary N‑glycans. In addition, antibodies with various tri‑antennary N‑glycans were obtained for the frst time by overexpressing MGAT5 alone or in combination with B4GalT1 and ST6Gal1. The various N‑glycan structures and the method for producing them in this work provide opportunities to study the glycan structure‑and‑function and develop novel recombinant antibodies for addressing diferent therapeutic applications.