Polymorphisms in Genes Involved in Neurodevelopment May Be Associated with Altered Brain Morphology in Schizophrenia: Preliminary Evidence
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Hyperactivated Wnt-Β-Catenin Signaling in the Absence of Sfrp1 and Sfrp5 Disrupts Trophoblast Differentiation Through Repressio
Bao et al. BMC Biology (2020) 18:151 https://doi.org/10.1186/s12915-020-00883-4 RESEARCH ARTICLE Open Access Hyperactivated Wnt-β-catenin signaling in the absence of sFRP1 and sFRP5 disrupts trophoblast differentiation through repression of Ascl2 Haili Bao1,2,3†, Dong Liu2†, Yingchun Xu2, Yang Sun2, Change Mu2, Yongqin Yu2, Chunping Wang2, Qian Han2, Sanmei Liu2, Han Cai1,2, Fan Liu2, Shuangbo Kong1,2, Wenbo Deng1,2, Bin Cao1,2, Haibin Wang1,2*, Qiang Wang3,4* and Jinhua Lu1,2* Abstract Background: Wnt signaling is a critical determinant for the maintenance and differentiation of stem/progenitor cells, including trophoblast stem cells during placental development. Hyperactivation of Wnt signaling has been shown to be associated with human trophoblast diseases. However, little is known about the impact and underlying mechanisms of excessive Wnt signaling during placental trophoblast development. Results: In the present work, we observed that two inhibitors of Wnt signaling, secreted frizzled-related proteins 1 and 5 (Sfrp1 and Sfrp5), are highly expressed in the extraembryonic trophoblast suggesting possible roles in early placental development. Sfrp1 and Sfrp5 double knockout mice exhibited disturbed trophoblast differentiation in the placental ectoplacental cone (EPC), which contains the precursors of trophoblast giant cells (TGCs) and spongiotrophoblast cells. In addition, we employed mouse models expressing a truncated β-catenin with exon 3 deletion globally and trophoblast-specifically, as well as trophoblast stem cell lines, and unraveled that hyperactivation of canonical Wnt pathway exhausted the trophoblast precursor cells in the EPC, resulting in the overabundance of giant cells at the expense of spongiotrophoblast cells. -

Supplementary Table 1: Adhesion Genes Data Set
Supplementary Table 1: Adhesion genes data set PROBE Entrez Gene ID Celera Gene ID Gene_Symbol Gene_Name 160832 1 hCG201364.3 A1BG alpha-1-B glycoprotein 223658 1 hCG201364.3 A1BG alpha-1-B glycoprotein 212988 102 hCG40040.3 ADAM10 ADAM metallopeptidase domain 10 133411 4185 hCG28232.2 ADAM11 ADAM metallopeptidase domain 11 110695 8038 hCG40937.4 ADAM12 ADAM metallopeptidase domain 12 (meltrin alpha) 195222 8038 hCG40937.4 ADAM12 ADAM metallopeptidase domain 12 (meltrin alpha) 165344 8751 hCG20021.3 ADAM15 ADAM metallopeptidase domain 15 (metargidin) 189065 6868 null ADAM17 ADAM metallopeptidase domain 17 (tumor necrosis factor, alpha, converting enzyme) 108119 8728 hCG15398.4 ADAM19 ADAM metallopeptidase domain 19 (meltrin beta) 117763 8748 hCG20675.3 ADAM20 ADAM metallopeptidase domain 20 126448 8747 hCG1785634.2 ADAM21 ADAM metallopeptidase domain 21 208981 8747 hCG1785634.2|hCG2042897 ADAM21 ADAM metallopeptidase domain 21 180903 53616 hCG17212.4 ADAM22 ADAM metallopeptidase domain 22 177272 8745 hCG1811623.1 ADAM23 ADAM metallopeptidase domain 23 102384 10863 hCG1818505.1 ADAM28 ADAM metallopeptidase domain 28 119968 11086 hCG1786734.2 ADAM29 ADAM metallopeptidase domain 29 205542 11085 hCG1997196.1 ADAM30 ADAM metallopeptidase domain 30 148417 80332 hCG39255.4 ADAM33 ADAM metallopeptidase domain 33 140492 8756 hCG1789002.2 ADAM7 ADAM metallopeptidase domain 7 122603 101 hCG1816947.1 ADAM8 ADAM metallopeptidase domain 8 183965 8754 hCG1996391 ADAM9 ADAM metallopeptidase domain 9 (meltrin gamma) 129974 27299 hCG15447.3 ADAMDEC1 ADAM-like, -

Neurology Genetics
An Official Journal of the American Academy of Neurology Neurology.org/ng • Online ISSN: 2376-7839 Volume 3, Number 4, August 2017 Genetics Functionally pathogenic ExACtly zero or once: Clinical and experimental EARS2 variants in vitro A clinically helpful guide studies of a novel P525R may not manifest a to assessing genetic FUS mutation in amyotrophic phenotype in vivo variants in mild epilepsies lateral sclerosis Table of Contents Neurology.org/ng Online ISSN: 2376-7839 Volume 3, Number 4, August 2017 THE HELIX e171 Autopsy case of the C12orf65 mutation in a patient e175 What does phenotype have to do with it? with signs of mitochondrial dysfunction S.M. Pulst H. Nishihara, M. Omoto, M. Takao, Y. Higuchi, M. Koga, M. Kawai, H. Kawano, E. Ikeda, H. Takashima, and T. Kanda EDITORIAL e173 This variant alters protein function, but is it pathogenic? M. Pandolfo e174 Prevalence of spinocerebellar ataxia 36 in a US Companion article, e162 population J.M. Valera, T. Diaz, L.E. Petty, B. Quintáns, Z. Yáñez, ARTICLES E. Boerwinkle, D. Muzny, D. Akhmedov, R. Berdeaux, e162 Functionally pathogenic EARS2 variants in vitro may M.J. Sobrido, R. Gibbs, J.R. Lupski, D.H. Geschwind, not manifest a phenotype in vivo S. Perlman, J.E. Below, and B.L. Fogel N. McNeill, A. Nasca, A. Reyes, B. Lemoine, B. Cantarel, A. Vanderver, R. Schiffmann, and D. Ghezzi Editorial, e173 e170 Loss-of-function variants of SCN8A in intellectual disability without seizures e163 ExACtly zero or once: A clinically helpful guide to J.L. Wagnon, B.S. Barker, M. -

Potential Impact of Mir-137 and Its Targets in Schizophrenia
Georgia State University ScholarWorks @ Georgia State University Psychology Faculty Publications Department of Psychology 4-2013 Potential Impact of miR-137 and Its Targets in Schizophrenia Carrie Wright University of New Mexico, [email protected] Jessica Turner Georgia State University, [email protected] Vince D. Calhoun University of New Mexico, [email protected] Nora I. Perrone-Bizzozero University of New Mexico, [email protected] Follow this and additional works at: https://scholarworks.gsu.edu/psych_facpub Part of the Psychology Commons Recommended Citation Wright C, Turner JA, Calhoun VD and Perrone-Bizzozero N (2013) Potential impact of miR-137 and its tar- gets in schizophrenia. Front. Genet. 4:58. doi: http://dx.doi.org/10.3389/fgene.2013.00058 This Article is brought to you for free and open access by the Department of Psychology at ScholarWorks @ Georgia State University. It has been accepted for inclusion in Psychology Faculty Publications by an authorized administrator of ScholarWorks @ Georgia State University. For more information, please contact [email protected]. HYPOTHESIS AND THEORY ARTICLE published: 26 April 2013 doi: 10.3389/fgene.2013.00058 Potential impact of miR-137 and its targets in schizophrenia Carrie Wright 1, Jessica A.Turner 2,3*,Vince D. Calhoun2,3 and Nora Perrone-Bizzozero1* 1 Department of Neurosciences, Health Sciences Center, University of New Mexico, Albuquerque, NM, USA 2 The Mind Research Network, Albuquerque, NM, USA 3 Psychology Department, University of New Mexico, Albuquerque, NM, USA Edited by: The significant impact of microRNAs (miRNAs) on disease pathology is becoming increas- Francis J. McMahon, National ingly evident.These small non-coding RNAs have the ability to post-transcriptionally silence Institute of Mental Health, USA the expression of thousands of genes. -

Molecular Characterisation of Renal Cell Carcinoma and Related Disorders
MOLECULAR CHARACTERISATION OF RENAL CELL CARCINOMA AND RELATED DISORDERS by MARIAM JAFRI A thesis submitted to the University of Birmingham for the degree of DOCTOR OF PHILOSOPHY School of Clinical and Experimental Medicine College of Medical and Dentistry September 2015 University of Birmingham Research Archive e-theses repository This unpublished thesis/dissertation is copyright of the author and/or third parties. The intellectual property rights of the author or third parties in respect of this work are as defined by The Copyright Designs and Patents Act 1988 or as modified by any successor legislation. Any use made of information contained in this thesis/dissertation must be in accordance with that legislation and must be properly acknowledged. Further distribution or reproduction in any format is prohibited without the permission of the copyright holder. ABSTRACT Over the last two decades genetic advances have provided novel insights into the molecular basis of familial and sporadic cancers and provided the basis for the development of novel therapeutic approaches. For example, the identification of the gene for von Hippel Lindau disease provided seminal insights into its role in most clear cell renal carcinomas (RCC) and led to new treatments for RCC. In this thesis I investigated three related genetic aspects of neoplasia. Firstly, I analyzed the results of genetic testing for inherited phaeochromocytoma and investigated how clinical features could be used to stratify patients and improve the cost effectiveness of genetic testing. Secondly, I sought to identify novel causes of inherited neoplasia. Through exome sequencing of familial RCC kindreds, CDKN2B was identified as a novel familial RCC gene. -
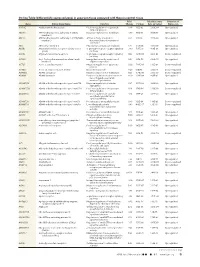
Differentially Expressed Genes in Aneurysm Tissue Compared With
On-line Table: Differentially expressed genes in aneurysm tissue compared with those in control tissue Fold False Discovery Direction of Gene Entrez Gene Name Function Change P Value Rate (q Value) Expression AADAC Arylacetamide deacetylase Positive regulation of triglyceride 4.46 1.33E-05 2.60E-04 Up-regulated catabolic process ABCA6 ATP-binding cassette, subfamily A (ABC1), Integral component of membrane 3.79 9.15E-14 8.88E-12 Up-regulated member 6 ABCC3 ATP-binding cassette, subfamily C (CFTR/MRP), ATPase activity, coupled to 6.63 1.21E-10 7.33E-09 Up-regulated member 3 transmembrane movement of substances ABI3 ABI family, member 3 Peptidyl-tyrosine phosphorylation 6.47 2.47E-05 4.56E-04 Up-regulated ACKR1 Atypical chemokine receptor 1 (Duffy blood G-protein–coupled receptor signaling 3.80 7.95E-10 4.18E-08 Up-regulated group) pathway ACKR2 Atypical chemokine receptor 2 G-protein–coupled receptor signaling 0.42 3.29E-04 4.41E-03 Down-regulated pathway ACSM1 Acyl-CoA synthetase medium-chain family Energy derivation by oxidation of 9.87 1.70E-08 6.52E-07 Up-regulated member 1 organic compounds ACTC1 Actin, ␣, cardiac muscle 1 Negative regulation of apoptotic 0.30 7.96E-06 1.65E-04 Down-regulated process ACTG2 Actin, ␥2, smooth muscle, enteric Blood microparticle 0.29 1.61E-16 2.36E-14 Down-regulated ADAM33 ADAM domain 33 Integral component of membrane 0.23 9.74E-09 3.95E-07 Down-regulated ADAM8 ADAM domain 8 Positive regulation of tumor necrosis 4.69 2.93E-04 4.01E-03 Up-regulated factor (ligand) superfamily member 11 production ADAMTS18 -

Polyclonal Antibody to Protocadherin-12 / PCDH12 (N-Term) - Purified
OriGene Technologies, Inc. OriGene Technologies GmbH 9620 Medical Center Drive, Ste 200 Schillerstr. 5 Rockville, MD 20850 32052 Herford UNITED STATES GERMANY Phone: +1-888-267-4436 Phone: +49-5221-34606-0 Fax: +1-301-340-8606 Fax: +49-5221-34606-11 [email protected] [email protected] AP23866PU-N Polyclonal Antibody to Protocadherin-12 / PCDH12 (N-term) - Purified Alternate names: VE-cad-2, VE-cadherin-2, Vascular cadherin-2, Vascular endothelial cadherin-2 Quantity: 50 µg Concentration: 1 mg/ml Background: PCDH12 / VE-Cadherin-2 belongs to the protocadherin gene family, a subfamily of the cadherin superfamily. The encoded protein consists of an extracellular domain containing 6 cadherin repeats, a transmembrane domain and a cytoplasmic tail that differs from those of the classical cadherins. The gene localizes to the region on chromosome 5 where the protocadherin gene clusters reside. Uniprot ID: Q9NPG4 NCBI: NP_057664 GeneID: 51294 Host: Rabbit Immunogen: 17 amino synthetic peptide near the amino terminus of Human PCDH12. Epitope: N-Terminus. Genename: PCDH12 Format: State: Liquid purfied Ig fraction Purification: Immunoaffinity Chromatography Buffer System: PBS containing 0.02% Sodium Azide Applications: ELISA. Immunohistochemistry on Paraffin Sections: 5 µg/ml. Western Blot: 1 - 2 µg/ml. Other applications not tested. Optimal dilutions are dependent on conditions and should be determined by the user. Specificity: This antibody reacts to the N-term of PCDH12. Species Reactivity: Tested: Human. Expected from sequence similarity: Mouse and Rat. Storage: Store undiluted at 2-8°C for one month or (in aliquots) at -20°C for longer. Avoid repeated freezing and thawing. -

Is Expressed in Endothelial, Trophoblast And
Protocadherin 12 (VE-cadherin 2) is expressed in endothelial, trophoblast and mesangial cells Christine Rampon1, Marie-Hélène Prandini1, Stéphanie Bouillot1, Hervé Pointu2, Emmanuelle Tillet1, Ronald Frank3, Muriel Vernet2 and Philippe Huber1. 1Laboratoire Développement et Vieillissement de l’Endothélium CEA-Inserm EMI-0219 and 2Atelier de Transgenèse, Département de Réponse et Dynamique Cellulaires, CEA-Grenoble, 17 rue des Martyrs 38054 Grenoble, France. 3Department of Chemical Biology, German Research Centre for Biotechnology, D-38124 Braunschweig, Germany. Author for correspondence: Philippe Huber, DRDC-DVE, 17 rue des Martyrs CEA-Grenoble 38 054 Grenoble. Tel: 33-438 78 41 18, Fax: 33-438 78 49 64, e-mail: [email protected]. Short title: Protocadherin 12 tissue expression ABSTRACT Protocadherin 12 protein (PCDH12, VE-cadherin 2) is a cell adhesion molecule which has been isolated from endothelial cells. Here, we have used Northern and Western blots, immunohistology and flow cytometry to examine the distribution of PCDH12 in mouse tissue. It is an N-glycosylated protein of 150 kDa mass. In the endothelium, PCDH12 immunoreactivity was variable and dependent upon the vascular bed. In both the embryo and embryonic stem cell differentiation system, signals were localized in vasculogenic rather than angiogenic endothelium. In addition, the protein was strongly expressed in a subset of invasive cells of the placenta, which were identified as glycogen-rich trophoblasts. In adult mice, strong PCDH12 signals were observed in mesangial cells of kidney glomeruli whereas expression was not detected in other types of perivascular cells. As opposed to most protocadherins, PCDH12 is not expressed in early embryonic (day 12.5) and adult brains. -

PCDH12 Variants
Brain calcifications and PCDH12 variants Gaël Nicolas, MD, PhD ABSTRACT Monica Sanchez- Objective: To assess the potential connection between PCDH12 and brain calcifications in Contreras, MD, PhD a patient carrying a homozygous nonsense variant in PCDH12 and in adult patients with brain Eliana Marisa Ramos, calcifications. PhD Methods: We performed a CT scan in 1 child with a homozygous PCDH12 nonsense variant. We Roberta R. Lemos, PhD screened DNA samples from 53 patients with primary familial brain calcification (PFBC) and 26 Joana Ferreira, PhD patients with brain calcification of unknown cause (BCUC). Denis Moura, BSc Maria J. Sobrido, MD, Results: We identified brain calcifications in subcortical and perithalamic regions in the patient PCDH12 PhD with a homozygous nonsense variant. The calcification pattern was different from what Anne-Claire Richard, BSc has been observed in PFBC and more similar to what is described in in utero infections. In patients Alma Rosa Lopez, BSc with PFBC or BCUC, we found no protein-truncating variant and 3 rare (minor allele frequency , PCDH12 Andrea Legati, PhD 0.001) predicted damaging missense heterozygous variants in 3 unrelated patients, Jean-François Deleuze, albeit with no segregation data available. MD, PhD Conclusions: Brain calcifications should be added to the phenotypic spectrum associated with Anne Boland, PharmD, PCDH12 biallelic loss of function, in the context of severe cerebral developmental abnormalities. PhD A putative role for PCDH12 variants remains to be determined in PFBC. Neurol Genet 2017;3: Olivier Quenez, MSc e166; doi: 10.1212/NXG.0000000000000166 Pierre Krystkowiak, MD, PhD GLOSSARY Pascal Favrole, MD BCUC 5 brain calcification of unknown cause; ExAC 5 Exome Aggregation Consortium; PFBC 5 primary familial brain calcification. -

Transcriptional Profile of Human Anti-Inflamatory Macrophages Under Homeostatic, Activating and Pathological Conditions
UNIVERSIDAD COMPLUTENSE DE MADRID FACULTAD DE CIENCIAS QUÍMICAS Departamento de Bioquímica y Biología Molecular I TESIS DOCTORAL Transcriptional profile of human anti-inflamatory macrophages under homeostatic, activating and pathological conditions Perfil transcripcional de macrófagos antiinflamatorios humanos en condiciones de homeostasis, activación y patológicas MEMORIA PARA OPTAR AL GRADO DE DOCTOR PRESENTADA POR Víctor Delgado Cuevas Directores María Marta Escribese Alonso Ángel Luís Corbí López Madrid, 2017 © Víctor Delgado Cuevas, 2016 Universidad Complutense de Madrid Facultad de Ciencias Químicas Dpto. de Bioquímica y Biología Molecular I TRANSCRIPTIONAL PROFILE OF HUMAN ANTI-INFLAMMATORY MACROPHAGES UNDER HOMEOSTATIC, ACTIVATING AND PATHOLOGICAL CONDITIONS Perfil transcripcional de macrófagos antiinflamatorios humanos en condiciones de homeostasis, activación y patológicas. Víctor Delgado Cuevas Tesis Doctoral Madrid 2016 Universidad Complutense de Madrid Facultad de Ciencias Químicas Dpto. de Bioquímica y Biología Molecular I TRANSCRIPTIONAL PROFILE OF HUMAN ANTI-INFLAMMATORY MACROPHAGES UNDER HOMEOSTATIC, ACTIVATING AND PATHOLOGICAL CONDITIONS Perfil transcripcional de macrófagos antiinflamatorios humanos en condiciones de homeostasis, activación y patológicas. Este trabajo ha sido realizado por Víctor Delgado Cuevas para optar al grado de Doctor en el Centro de Investigaciones Biológicas de Madrid (CSIC), bajo la dirección de la Dra. María Marta Escribese Alonso y el Dr. Ángel Luís Corbí López Fdo. Dra. María Marta Escribese -

Decoding Transcriptional Regulation Via a Human Gene Expression Predictor
bioRxiv preprint doi: https://doi.org/10.1101/2020.04.07.029470; this version posted April 8, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. Decoding transcriptional regulation via a human gene expression predictor Yuzhou Wang*1,2, Yu Zhang*1, Jiazhen Gong1, Jianqiang Bao2, Shisong Ma1,3 1. Hefei National Laboratory for Physical Sciences at the Microscale, School of Life Sciences, Division of Life Sciences and Medicine, University of Science and Technology of China, Innovative Academy of Seed Design, Chinese Academy of Sciences, Hefei, China 2. The First Affiliated Hospital of USTC, Division of Life Sciences and Medicine, University of Science and Technology of China, Hefei, China 3. School of Data Science, University of Science and Technology of China, Hefei, China * These authors contribute equally. Correspondence: Shisong Ma ([email protected]) ABSTRACT Transcription factors (TF) regulate cellular activities via controlling gene expression, but a predictive model describing how TFs quantitatively modulate human transcriptomes was lacking. We constructed a universal human gene expression predictor and utilized it to decode transcriptional regulation. Using 1613 TFs’ expression, the predictor reconstituted highly accurate transcriptomes for samples derived from a wide range of tissues and conditions. The predictor’s broad applicability indicated it had recapitulated the quantitative relationships between TFs and target genes ubiquitous across tissues. Significant interacting TF-target gene pairs were then extracted from the predictor and enabled downstream inference of TF regulators for diverse pathways involved in development, immunity, metabolism, and stress response. -

Supplementary Materials
Supplementary Materials 1 Supplementary Figure S1. Expression of BPIFB4 in murine hearts. Representative immunohistochemistry images showing the expression of BPIFB4 in the left ventricle of non-diabetic mice (ND) and diabetic mice (Diab) given vehicle or LAV-BPIFB4. BPIFB4 is shown in green, nuclei are identified by the blue fluorescence of DAPI. Scale bars: 1 mm and 100 μm. 2 Supplementary Table S1 – List of PCR primers. Target gene Primer Sequence NCBI Accession number / Reference Forward AAGTCCCTCACCCTCCCAA Actb [1] Reverse AAGCAATGCTGTCACCTTC Forward TCTAGGCAATGCCGTTCAC Cpt1b [2] Reverse GAGCACATGGGCACCATAC Forward GGAAATGATCAACAAAAAAAGAAGTATTT Acadm (Mcad) [2] Reverse GCCGCCACATCAGA Forward TGATGGTTTGGAGGTTGGGG Acot1 NM_012006.2 Reverse TGAAACTCCATTCCCAGCCC Forward GGTGTCCCGTCTAATGGAGA Hmgcs2 NM_008256.4 Reverse ACACCCAGGATTCACAGAGG Forward CAAGCAGCAACATGGGAAGA Cs [2] Reverse GTCAGGATCAAGAACCGAAGTCT Forward GCCATTGTCAACTGTGCTGA Ucp3 NM_009464.3 Reverse TCCTGAGCCACCATCTTCAG Forward CGTGAGGGCAATGATTTATACCAT Atp5b [2] Reverse TCCTGGTCTCTGAAGTATTCAGCAA Pdk4 Forward CCGCTGTCCATGAAGCA [2] 3 Reverse GCAGAAAAGCAAAGGACGTT Forward AGAGTCCTATGCAGCCCAGA Tomm20 NM_024214.2 Reverse CAAAGCCCCACATCTGTCCT Forward GCCTCAGATCGTCGTAGTGG Drp1 NM_152816.3 Reverse TTCCATGTGGCAGGGTCATT Forward GGGAAGGTGAAGAAGCTTGGA Mfn2 NM_001285920.1 Reverse ACAACTGGAACAGAGGAGAAGTT Forward CGGAAATCATATCCAACCAG [2] Ppargc1a (Pgc1α) Reverse TGAGAACCGCTAGCAAGTTTG Forward AGGCTTGGAAAAATCTGTCTC [2] Tfam Reverse TGCTCTTCCCAAGACTTCATT Forward TGCCCCAGAGCTGTTAATGA Bcl2l1 NM_001289716.1