Copy Number Changes Can Be a Predictor for Hemoglobin Reduction After S-1 Monotherapy in Gastric Cancer
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-
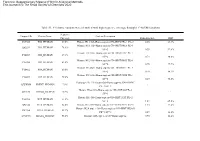
Table S1. 49 Histone Variants Were Identified with High Sequence Coverage Through LC-MS/MS Analysis Electronic Supplementary
Electronic Supplementary Material (ESI) for Analytical Methods. This journal is © The Royal Society of Chemistry 2020 Table S1. 49 histone variants were identified with high sequence coverage through LC-MS/MS analysis Sequence Uniprot IDs Protein Name Protein Description Coverage Ratio E2+/E2- RSD P07305 H10_HUMAN 67.5% Histone H1.0 OS=Homo sapiens GN=H1F0 PE=1 SV=3 4.85 23.3% Histone H1.1 OS=Homo sapiens GN=HIST1H1A PE=1 Q02539 H11_HUMAN 74.4% SV=3 0.35 92.6% Histone H1.2 OS=Homo sapiens GN=HIST1H1C PE=1 P16403 H12_HUMAN 67.1% SV=2 0.73 80.6% Histone H1.3 OS=Homo sapiens GN=HIST1H1D PE=1 P16402 H13_HUMAN 63.8% SV=2 0.75 77.7% Histone H1.4 OS=Homo sapiens GN=HIST1H1E PE=1 P10412 H14_HUMAN 69.0% SV=2 0.70 80.3% Histone H1.5 OS=Homo sapiens GN=HIST1H1B PE=1 P16401 H15_HUMAN 79.6% SV=3 0.29 98.3% Testis-specific H1 histone OS=Homo sapiens GN=H1FNT Q75WM6 H1FNT_HUMAN 7.8% \ \ PE=2 SV=3 Histone H1oo OS=Homo sapiens GN=H1FOO PE=2 Q8IZA3 H1FOO_HUMAN 5.2% \ \ SV=1 Histone H1t OS=Homo sapiens GN=HIST1H1T PE=2 P22492 H1T_HUMAN 31.4% SV=4 1.42 65.0% Q92522 H1X_HUMAN 82.6% Histone H1x OS=Homo sapiens GN=H1FX PE=1 SV=1 1.15 33.2% Histone H2A type 1 OS=Homo sapiens GN=HIST1H2AG P0C0S8 H2A1_HUMAN 99.2% PE=1 SV=2 0.57 26.8% Q96QV6 H2A1A_HUMAN 58.0% Histone H2A type 1-A OS=Homo sapiens 0.90 11.2% GN=HIST1H2AA PE=1 SV=3 Histone H2A type 1-B/E OS=Homo sapiens P04908 H2A1B_HUMAN 99.2% GN=HIST1H2AB PE=1 SV=2 0.92 30.2% Histone H2A type 1-C OS=Homo sapiens Q93077 H2A1C_HUMAN 100.0% GN=HIST1H2AC PE=1 SV=3 0.76 27.6% Histone H2A type 1-D OS=Homo sapiens P20671 -

A Cell Line P53 Mutation Type UM
A Cell line p53 mutation Type UM-SCC 1 wt UM-SCC5 Exon 5, 157 GTC --> TTC Missense mutation by transversion (Valine --> Phenylalanine UM-SCC6 wt UM-SCC9 wt UM-SCC11A wt UM-SCC11B Exon 7, 242 TGC --> TCC Missense mutation by transversion (Cysteine --> Serine) UM-SCC22A Exon 6, 220 TAT --> TGT Missense mutation by transition (Tyrosine --> Cysteine) UM-SCC22B Exon 6, 220 TAT --> TGT Missense mutation by transition (Tyrosine --> Cysteine) UM-SCC38 Exon 5, 132 AAG --> AAT Missense mutation by transversion (Lysine --> Asparagine) UM-SCC46 Exon 8, 278 CCT --> CGT Missense mutation by transversion (Proline --> Alanine) B 1 Supplementary Methods Cell Lines and Cell Culture A panel of ten established HNSCC cell lines from the University of Michigan series (UM-SCC) was obtained from Dr. T. E. Carey at the University of Michigan, Ann Arbor, MI. The UM-SCC cell lines were derived from eight patients with SCC of the upper aerodigestive tract (supplemental Table 1). Patient age at tumor diagnosis ranged from 37 to 72 years. The cell lines selected were obtained from patients with stage I-IV tumors, distributed among oral, pharyngeal and laryngeal sites. All the patients had aggressive disease, with early recurrence and death within two years of therapy. Cell lines established from single isolates of a patient specimen are designated by a numeric designation, and where isolates from two time points or anatomical sites were obtained, the designation includes an alphabetical suffix (i.e., "A" or "B"). The cell lines were maintained in Eagle's minimal essential media supplemented with 10% fetal bovine serum and penicillin/streptomycin. -

Herpes Simplex Virus Blocks Host Transcription Termination Via the Bimodal Activities of ICP27
ARTICLE https://doi.org/10.1038/s41467-019-14109-x OPEN Herpes simplex virus blocks host transcription termination via the bimodal activities of ICP27 Xiuye Wang 1, Thomas Hennig2, Adam W. Whisnant 2, Florian Erhard 2, Bhupesh K. Prusty 2, Caroline C. Friedel 3, Elmira Forouzmand4,5, William Hu1, Luke Erber 6, Yue Chen6, Rozanne M. Sandri-Goldin 1*, Lars Dölken 2,7* & Yongsheng Shi1* Infection by viruses, including herpes simplex virus-1 (HSV-1), and cellular stresses cause 1234567890():,; widespread disruption of transcription termination (DoTT) of RNA polymerase II (RNAPII) in host genes. However, the underlying mechanisms remain unclear. Here, we demonstrate that the HSV-1 immediate early protein ICP27 induces DoTT by directly binding to the essential mRNA 3’ processing factor CPSF. It thereby induces the assembly of a dead-end 3’ processing complex, blocking mRNA 3’ cleavage. Remarkably, ICP27 also acts as a sequence- dependent activator of mRNA 3’ processing for viral and a subset of host transcripts. Our results unravel a bimodal activity of ICP27 that plays a key role in HSV-1-induced host shutoff and identify CPSF as an important factor that mediates regulation of transcription termination. These findings have broad implications for understanding the regulation of transcription termination by other viruses, cellular stress and cancer. 1 Department of Microbiology and Molecular Genetics, School of Medicine, University of California, Irvine, Irvine, CA 92697, USA. 2 Institute for Virology and Immunobiology, Julius-Maximilians-University Würzburg, Würzburg, Germany. 3 Institute of Informatics, Ludwig-Maximilians-Universität München, München, Germany. 4 Institute for Genomics and Bioinformatics, University of California, Irvine, Irvine, CA 92697, USA. -

Genome-Wide Screen of Cell-Cycle Regulators in Normal and Tumor Cells
bioRxiv preprint doi: https://doi.org/10.1101/060350; this version posted June 23, 2016. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. Genome-wide screen of cell-cycle regulators in normal and tumor cells identifies a differential response to nucleosome depletion Maria Sokolova1, Mikko Turunen1, Oliver Mortusewicz3, Teemu Kivioja1, Patrick Herr3, Anna Vähärautio1, Mikael Björklund1, Minna Taipale2, Thomas Helleday3 and Jussi Taipale1,2,* 1Genome-Scale Biology Program, P.O. Box 63, FI-00014 University of Helsinki, Finland. 2Science for Life laboratory, Department of Biosciences and Nutrition, Karolinska Institutet, SE- 141 83 Stockholm, Sweden. 3Science for Life laboratory, Division of Translational Medicine and Chemical Biology, Department of Medical Biochemistry and Biophysics, Karolinska Institutet, S-171 21 Stockholm, Sweden To identify cell cycle regulators that enable cancer cells to replicate DNA and divide in an unrestricted manner, we performed a parallel genome-wide RNAi screen in normal and cancer cell lines. In addition to many shared regulators, we found that tumor and normal cells are differentially sensitive to loss of the histone genes transcriptional regulator CASP8AP2. In cancer cells, loss of CASP8AP2 leads to a failure to synthesize sufficient amount of histones in the S-phase of the cell cycle, resulting in slowing of individual replication forks. Despite this, DNA replication fails to arrest, and tumor cells progress in an elongated S-phase that lasts several days, finally resulting in death of most of the affected cells. -

Histone-Related Genes Are Hypermethylated in Lung Cancer
Published OnlineFirst October 1, 2019; DOI: 10.1158/0008-5472.CAN-19-1019 Cancer Genome and Epigenome Research Histone-Related Genes Are Hypermethylated in Lung Cancer and Hypermethylated HIST1H4F Could Serve as a Pan-Cancer Biomarker Shihua Dong1,Wei Li1, Lin Wang2, Jie Hu3,Yuanlin Song3, Baolong Zhang1, Xiaoguang Ren1, Shimeng Ji3, Jin Li1, Peng Xu1, Ying Liang1, Gang Chen4, Jia-Tao Lou2, and Wenqiang Yu1 Abstract Lung cancer is the leading cause of cancer-related deaths lated in all 17 tumor types from TCGA datasets (n ¼ 7,344), worldwide. Cytologic examination is the current "gold stan- which was further validated in nine different types of cancer dard" for lung cancer diagnosis, however, this has low sensi- (n ¼ 243). These results demonstrate that HIST1H4F can tivity. Here, we identified a typical methylation signature of function as a universal-cancer-only methylation (UCOM) histone genes in lung cancer by whole-genome DNA methyl- marker, which may aid in understanding general tumorigen- ation analysis, which was validated by The Cancer Genome esis and improve screening for early cancer diagnosis. Atlas (TCGA) lung cancer cohort (n ¼ 907) and was further confirmed in 265 bronchoalveolar lavage fluid samples with Significance: These findings identify a new biomarker for specificity and sensitivity of 96.7% and 87.0%, respectively. cancer detection and show that hypermethylation of histone- More importantly, HIST1H4F was universally hypermethy- related genes seems to persist across cancers. Introduction to its low specificity, LDCT is far from satisfactory as a screening tool for clinical application, similar to other currently used cancer Lung cancer is one of the most common malignant tumors and biomarkers, such as carcinoembryonic antigen (CEA), neuron- the leading cause of cancer-related deaths worldwide (1, 2). -

ZNF410 Represses Fetal Globin by Devoted Control of CHD4/Nurd
bioRxiv preprint doi: https://doi.org/10.1101/2020.08.31.272856; this version posted August 31, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. Title ZNF410 represses fetal globin by devoted control of CHD4/NuRD Authors Divya S. Vinjamur1, Qiuming Yao1,2, Mitchel A. Cole1, Connor McGuckin1, Chunyan Ren1, Jing Zeng1, Mir Hossain1, Kevin Luk3, Scot A. Wolfe3, Luca Pinello2, Daniel E. Bauer1,4 1Division of Hematology/Oncology, Boston Children’s Hospital, Department of Pediatric Oncology, Dana-Farber Cancer Institute, Harvard Stem Cell Institute, Broad Institute, Department of Pediatrics, Harvard Medical School, Boston, Massachusetts 02115, USA 2Molecular Pathology Unit, Center for Cancer Research, and Center for Computational and Integrative Biology, Massachusetts General Hospital, Department of Pathology, Harvard Medical School, Boston, Massachusetts 02129, USA 3Department of Molecular, Cell and Cancer Biology, Li Weibo Institute for Rare Diseases Research, University of Massachusetts Medical School, Worcester, Massachusetts 01605, USA 4Correspondence: [email protected] bioRxiv preprint doi: https://doi.org/10.1101/2020.08.31.272856; this version posted August 31, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. Abstract Major effectors of adult-stage fetal globin silencing include the transcription factors (TFs) BCL11A and ZBTB7A/LRF and the NuRD chromatin complex, although each has potential on- target liabilities for rational �-hemoglobinopathy therapeutic inhibition. -

The Human Canonical Core Histone Catalogue David Miguel Susano Pinto*, Andrew Flaus*,†
bioRxiv preprint doi: https://doi.org/10.1101/720235; this version posted July 30, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY 4.0 International license. The Human Canonical Core Histone Catalogue David Miguel Susano Pinto*, Andrew Flaus*,† Abstract Core histone proteins H2A, H2B, H3, and H4 are encoded by a large family of genes dis- tributed across the human genome. Canonical core histones contribute the majority of proteins to bulk chromatin packaging, and are encoded in 4 clusters by 65 coding genes comprising 17 for H2A, 18 for H2B, 15 for H3, and 15 for H4, along with at least 17 total pseudogenes. The canonical core histone genes display coding variation that gives rise to 11 H2A, 15 H2B, 4 H3, and 2 H4 unique protein isoforms. Although histone proteins are highly conserved overall, these isoforms represent a surprising and seldom recognised variation with amino acid identity as low as 77 % between canonical histone proteins of the same type. The gene sequence and protein isoform diversity also exceeds com- monly used subtype designations such as H2A.1 and H3.1, and exists in parallel with the well-known specialisation of variant histone proteins. RNA sequencing of histone transcripts shows evidence for differential expression of histone genes but the functional significance of this variation has not yet been investigated. To assist understanding of the implications of histone gene and protein diversity we have catalogued the entire human canonical core histone gene and protein complement. -

Supplemental Information
Supplemental Information Chromatin-to-nucleoprotamine transition is controlled by the histone H2B variant TH2B Emilie Montellier1, Fayçal Boussouar1, Sophie Rousseaux1, Kai Zhang2, Thierry Buchou1, François Fenaille3, Hitoshi Shiota1, Alexandra Debernardi1, Patrick Héry4, Sandrine Curtet1, Mahya Jamshidikia1, Sophie Barral1, Hélène Holota5, Aurélie Bergon5, Fabrice Lopez5, Philippe Guardiola6, Karin Pernet7, Jean Imbert5, Carlo Petosa8, Minjia Tan9,10, Yingming Zhao9,10, Matthieu Gérard4, Saadi Khochbin1* 1 - INSERM, U823; Université Joseph Fourier - Grenoble 1; Institut Albert Bonniot, Grenoble, F-38700 France 2 - State Key Laboratory of Medicinal Chemical Biology & Department of Chemistry, Nankai University, Tianjin 300071, China 3 – Laboratoire d'Etude du Métabolisme des Médicaments, DSV / iBiTec-S / SPI, CEA Saclay, 91191 Gif sur Yvette, Cedex, France 4 - CEA, iBiTec-S, Gif-sur-Yvette, F-91191 France 5 - INSERM UMR_S 1090; TGML/TAGC, Aix-Marseille Université, Marseille, France 6 - INSERM, U892; Centre de Recherche sur le Cancer Nantes Angers et UMR_S 892; Université d’Angers; Plateforme SNP, Transcriptome & Epigénomique; Centre Hospitalier Universitaire d’Angers; Angers, F-49000 France 7 - INSERM U836; Université Joseph Fourier, Grenoble Institute of Neuroscience, Grenoble, F-38700 France, 8 - Institut de Biologie Structurale Jean-Pierre Ebel, UMR 5075 CEA-CNRS-Université Joseph Fourier, 41 Jules Horowitz, 38027 Grenoble cedex 1. 9 - Ben May Department of Cancer Research, The University of Chicago, Chicago, IL 60637, USA 10 - Shanghai Institute of Materia Medica, Chinese Academy of Sciences, 555 Zuchongzhi Road, Shanghai 201203, P.R. China *Correspondance : Saadi Khochbin ([email protected]). 1 SUPPLEMENTAL DATA SUPPLEMENTAL FIGURE LEGENDS Figure S1. Generation of the Th2b tag knock-in and knock-down mouse models. -

HIST1H2BL Polyclonal Antibody Catalog Number PA5-48429 Product Data Sheet
Lot Number: SH2432915A Website: thermofisher.com Customer Service (US): 1 800 955 6288 ext. 1 Technical Support (US): 1 800 955 6288 ext. 441 thermofisher.com/contactus HIST1H2BL Polyclonal Antibody Catalog Number PA5-48429 Product Data Sheet Details Species Reactivity Size 400 µl Tested species reactivity Human Host / Isotype Rabbit / IgG Tested Applications Dilution * Class Polyclonal Flow Cytometry (Flow) 1:10-1:50 Type Antibody Western Blot (WB) 1:1000 KLH conjugated synthetic peptide between 1-30 amino acids from the * Suggested working dilutions are given as a guide only. It is recommended that the user titrate the product for use in their Immunogen own experiment using appropriate negative and positive controls. N-terminal region of human HIST1H2BL. Conjugate Unconjugated Form Liquid Concentration Lot-Specific Purification Antigen affinity chromatography Storage Buffer PBS Contains 0.09% sodium azide Storage Conditions -20° C, Avoid Freeze/Thaw Cycles Product Specific Information Predicted to react with mouse based on sequence homology. Background/Target Information Core component of nucleosome. Nucleosomes wrap and compact DNA into chromatin, limiting DNA accessibility to the cellular machineries which require DNA as a template. Histones thereby play a central role in transcription regulation, DNA repair, DNA replication and chromosomal stability. DNA accessibility is regulated via a complex set of post-translational modifications of histones, also called histone code, and nucleosome remodeling. For Research Use Only. Not for use in diagnostic procedures. Not for resale without express authorization. For Research Use Only. Not for use in diagnostic procedures. Not for resale without express authorization. Products are warranted to operate or perform substantially in conformance with published Product specifications in effect at the time of sale, as set forth in the Production documentation, specifications and/or accompanying package inserts ("Documentation"). -

Whole-Genome Sequencing Identifies New Genetic Alterations in Meningiomas
www.impactjournals.com/oncotarget/ Oncotarget, 2017, Vol. 8, (No. 10), pp: 17070-17080 Research Paper Whole-genome sequencing identifies new genetic alterations in meningiomas Mei Tang1,*, Heng Wei2,*, Lu Han1,*, Jiaojiao Deng3, Yuelong Wang3, Meijia Yang1, Yani Tang1, Gang Guo1, Liangxue Zhou3, Aiping Tong1 1The State Key Laboratory of Biotherapy and Cancer Center/Collaborative Innovation Center of Biotherapy, West China Hospital, West China Medical School, Sichuan University, Chengdu 610041, China 2College of Life Science, Sichuan University, Chengdu 610064, China 3Department of Neurosurgery, West China Hospital, West China Medical School, Sichuan University, Chengdu 610041, China *These authors have contributed equally to the work Correspondence to: Aiping Tong, email: [email protected] Liangxue Zhou, email: [email protected] Keywords: whole-genome sequencing, meningioma, chromosome instability, copy number alteration, mutation Received: October 24, 2016 Accepted: January 13, 2017 Published: February 03, 2017 ABSTRACT The major known genetic contributor to meningioma formation was NF2, which is disrupted by mutation or loss in about 50% of tumors. Besides NF2, several recurrent driver mutations were recently uncovered through next-generation sequencing. Here, we performed whole-genome sequencing across 7 tumor-normal pairs to identify somatic genetic alterations in meningioma. As a result, Chromatin regulators, including multiple histone members, histone-modifying enzymes and several epigenetic regulators, are the major category among all of the identified copy number variants and single nucleotide variants. Notably, all samples contained copy number variants in histone members. Recurrent chromosomal rearrangements were detected on chromosome 22q, 6p21-p22 and 1q21, and most of the histone copy number variants occurred in these regions. -

Peripheral Nerve Single-Cell Analysis Identifies Mesenchymal Ligands That Promote Axonal Growth
Research Article: New Research Development Peripheral Nerve Single-Cell Analysis Identifies Mesenchymal Ligands that Promote Axonal Growth Jeremy S. Toma,1 Konstantina Karamboulas,1,ª Matthew J. Carr,1,2,ª Adelaida Kolaj,1,3 Scott A. Yuzwa,1 Neemat Mahmud,1,3 Mekayla A. Storer,1 David R. Kaplan,1,2,4 and Freda D. Miller1,2,3,4 https://doi.org/10.1523/ENEURO.0066-20.2020 1Program in Neurosciences and Mental Health, Hospital for Sick Children, 555 University Avenue, Toronto, Ontario M5G 1X8, Canada, 2Institute of Medical Sciences University of Toronto, Toronto, Ontario M5G 1A8, Canada, 3Department of Physiology, University of Toronto, Toronto, Ontario M5G 1A8, Canada, and 4Department of Molecular Genetics, University of Toronto, Toronto, Ontario M5G 1A8, Canada Abstract Peripheral nerves provide a supportive growth environment for developing and regenerating axons and are es- sential for maintenance and repair of many non-neural tissues. This capacity has largely been ascribed to paracrine factors secreted by nerve-resident Schwann cells. Here, we used single-cell transcriptional profiling to identify ligands made by different injured rodent nerve cell types and have combined this with cell-surface mass spectrometry to computationally model potential paracrine interactions with peripheral neurons. These analyses show that peripheral nerves make many ligands predicted to act on peripheral and CNS neurons, in- cluding known and previously uncharacterized ligands. While Schwann cells are an important ligand source within injured nerves, more than half of the predicted ligands are made by nerve-resident mesenchymal cells, including the endoneurial cells most closely associated with peripheral axons. At least three of these mesen- chymal ligands, ANGPT1, CCL11, and VEGFC, promote growth when locally applied on sympathetic axons. -
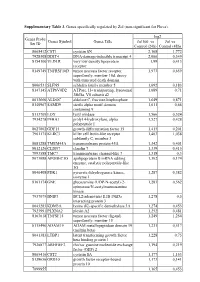
Supplementary Table 3. Genes Specifically Regulated by Zol (Non-Significant for Fluva)
Supplementary Table 3. Genes specifically regulated by Zol (non-significant for Fluva). log2 Genes Probe Genes Symbol Genes Title Zol100 vs Zol vs Set ID Control (24h) Control (48h) 8065412 CST1 cystatin SN 2,168 1,772 7928308 DDIT4 DNA-damage-inducible transcript 4 2,066 0,349 8154100 VLDLR very low density lipoprotein 1,99 0,413 receptor 8149749 TNFRSF10D tumor necrosis factor receptor 1,973 0,659 superfamily, member 10d, decoy with truncated death domain 8006531 SLFN5 schlafen family member 5 1,692 0,183 8147145 ATP6V0D2 ATPase, H+ transporting, lysosomal 1,689 0,71 38kDa, V0 subunit d2 8013660 ALDOC aldolase C, fructose-bisphosphate 1,649 0,871 8140967 SAMD9 sterile alpha motif domain 1,611 0,66 containing 9 8113709 LOX lysyl oxidase 1,566 0,524 7934278 P4HA1 prolyl 4-hydroxylase, alpha 1,527 0,428 polypeptide I 8027002 GDF15 growth differentiation factor 15 1,415 0,201 7961175 KLRC3 killer cell lectin-like receptor 1,403 1,038 subfamily C, member 3 8081288 TMEM45A transmembrane protein 45A 1,342 0,401 8012126 CLDN7 claudin 7 1,339 0,415 7993588 TMC7 transmembrane channel-like 7 1,318 0,3 8073088 APOBEC3G apolipoprotein B mRNA editing 1,302 0,174 enzyme, catalytic polypeptide-like 3G 8046408 PDK1 pyruvate dehydrogenase kinase, 1,287 0,382 isozyme 1 8161174 GNE glucosamine (UDP-N-acetyl)-2- 1,283 0,562 epimerase/N-acetylmannosamine kinase 7937079 BNIP3 BCL2/adenovirus E1B 19kDa 1,278 0,5 interacting protein 3 8043283 KDM3A lysine (K)-specific demethylase 3A 1,274 0,453 7923991 PLXNA2 plexin A2 1,252 0,481 8163618 TNFSF15 tumor necrosis