Tgfβ Suppresses Type 2 Immunity to Cancer Ming Liu1
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-
(12) Patent Application Publication (10) Pub. No.: US 2012/0070450 A1 Ishikawa Et Al
US 20120070450A1 (19) United States (12) Patent Application Publication (10) Pub. No.: US 2012/0070450 A1 Ishikawa et al. (43) Pub. Date: Mar. 22, 2012 (54) LEUKEMA STEM CELLMARKERS Publication Classification (51) Int. Cl. A 6LX 39/395 (2006.01) (75) Inventors: Fumihiko Ishikawa, Kanagawa CI2O I/68 (2006.01) (JP): Osamu Ohara, Kanagawa GOIN 2L/64 (2006.01) (JP); Yoriko Saito, Kanagawa (JP); A6IP35/02 (2006.01) Hiroshi Kitamura, Kanagawa (JP); C40B 30/04 (2006.01) Atsushi Hijikata, Kanagawa (JP); A63L/7088 (2006.01) Hidetoshi Ozawa, Kanagawa (JP); C07K 6/8 (2006.01) Leonard D. Shultz, Bar Harbor, C7H 2L/00 (2006.01) A6II 35/12 (2006.01) ME (US) CI2N 5/078 (2010.01) (52) U.S. Cl. .................. 424/173.1; 424/178.1; 424/93.7: (73) Assignee: RIKEN, Wako-shi (JP) 435/6.14; 435/723; 435/375; 506/9: 514/44 A: 530/389.6; 530/391.7:536/24.5 (57) ABSTRACT (21) Appl. No.: 13/258,993 The invention provides a test method for predicting the initial onset or a recurrence of acute myeloid leukemia (AML) com PCT Fled: prising (1) measuring the expression level of human leukemic (22) Mar. 24, 2010 stem cell (LSC) marker genes in a biological sample collected from a Subject for a transcription product or translation prod uct of the gene as an analyte and (2) comparing the expression (86) PCT NO.: PCT/UP2010/0551.31 level with a reference value; an LSC-targeting therapeutic agent for AML capable of Suppressing the expression of a S371 (c)(1), gene selected from among LSC marker genes or a Substance (2), (4) Date: Dec. -

Role of IL-4 Receptor &Alpha
Role of IL-4 receptor a–positive CD41 T cells in chronic airway hyperresponsiveness Frank Kirstein, PhD, Natalie E. Nieuwenhuizen, PhD,* Jaisubash Jayakumar, PhD, William G. C. Horsnell, PhD, and Frank Brombacher, PhD Cape Town, South Africa, and Berlin, Germany Background: TH2 cells and their cytokines are associated with IL-17–producing T cells and, consequently, increased airway allergic asthma in human subjects and with mouse models of neutrophilia. allergic airway disease. IL-4 signaling through the IL-4 receptor Conclusion: IL-4–responsive T helper cells are dispensable for 1 a (IL-4Ra) chain on CD4 T cells leads to TH2 cell acute OVA-induced airway disease but crucial in maintaining differentiation in vitro, implying that IL-4Ra–responsive CD41 chronic asthmatic pathology. (J Allergy Clin Immunol T cells are critical for the induction of allergic asthma. However, 2016;137:1852-62.) mechanisms regulating acute and chronic allergen-specific T 2 H Key words: responses in vivo remain incompletely understood. TH2 cell, acute allergic airway disease, chronic asthma, Objective: This study defines the requirements for IL-4Ra– cytokine receptors, IL-4, IL-13, gene-deficient mice responsive CD41 T cells and the IL-4Ra ligands IL-4 and IL-13 in the development of allergen-specific TH2 responses during the Allergic asthma is a chronic inflammatory disease of the airways onset and chronic phase of experimental allergic airway disease. characterized by an inappropriate immune response to harmless Methods: Development of acute and chronic ovalbumin environmental antigens. T 2 cells regulate adaptive immune 1 H (OVA)–induced allergic asthma was assessed weekly in CD4 T responses to allergens, and their presence correlates with disease 2 cell–specific IL-4Ra–deficient BALB/c mice (LckcreIL-4Ra /lox) symptoms in human subjects and mice.1 IL-4 plays a crucial role and respective control mice in the presence or absence of IL-4 in the in vitro and in vivo differentiation of TH2 cells, suggesting or IL-13. -
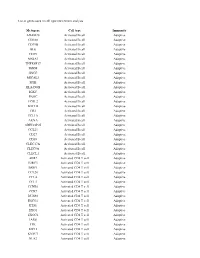
List of Genes Used in Cell Type Enrichment Analysis
List of genes used in cell type enrichment analysis Metagene Cell type Immunity ADAM28 Activated B cell Adaptive CD180 Activated B cell Adaptive CD79B Activated B cell Adaptive BLK Activated B cell Adaptive CD19 Activated B cell Adaptive MS4A1 Activated B cell Adaptive TNFRSF17 Activated B cell Adaptive IGHM Activated B cell Adaptive GNG7 Activated B cell Adaptive MICAL3 Activated B cell Adaptive SPIB Activated B cell Adaptive HLA-DOB Activated B cell Adaptive IGKC Activated B cell Adaptive PNOC Activated B cell Adaptive FCRL2 Activated B cell Adaptive BACH2 Activated B cell Adaptive CR2 Activated B cell Adaptive TCL1A Activated B cell Adaptive AKNA Activated B cell Adaptive ARHGAP25 Activated B cell Adaptive CCL21 Activated B cell Adaptive CD27 Activated B cell Adaptive CD38 Activated B cell Adaptive CLEC17A Activated B cell Adaptive CLEC9A Activated B cell Adaptive CLECL1 Activated B cell Adaptive AIM2 Activated CD4 T cell Adaptive BIRC3 Activated CD4 T cell Adaptive BRIP1 Activated CD4 T cell Adaptive CCL20 Activated CD4 T cell Adaptive CCL4 Activated CD4 T cell Adaptive CCL5 Activated CD4 T cell Adaptive CCNB1 Activated CD4 T cell Adaptive CCR7 Activated CD4 T cell Adaptive DUSP2 Activated CD4 T cell Adaptive ESCO2 Activated CD4 T cell Adaptive ETS1 Activated CD4 T cell Adaptive EXO1 Activated CD4 T cell Adaptive EXOC6 Activated CD4 T cell Adaptive IARS Activated CD4 T cell Adaptive ITK Activated CD4 T cell Adaptive KIF11 Activated CD4 T cell Adaptive KNTC1 Activated CD4 T cell Adaptive NUF2 Activated CD4 T cell Adaptive PRC1 Activated -

CD4 T-Cell Cytokines Synergize to Induce Proliferation of Malignant and Nonmalignant Innate Intraepithelial Lymphocytes
CD4 T-cell cytokines synergize to induce proliferation of malignant and nonmalignant innate intraepithelial lymphocytes Yvonne M. C. Kooy-Winkelaara, Dagmar Bouwera, George M. C. Janssenb, Allan Thompsona, Martijn H. Brugmana, Frederike Schmitza, Arnoud H. de Rub, Tom van Gilsc, Gerd Boumac, Jon J. van Rooda,1, Peter A. van Veelenb, M. Luisa Mearind, Chris J. Mulderc, Frits Koninga, and Jeroen van Bergena,1 aDepartment of Immunohematology and Blood Transfusion, Leiden University Medical Center, Leiden 2333 ZA, The Netherlands; bCenter for Proteomics and Metabolomics, Leiden University Medical Center, Leiden 2333 ZA, The Netherlands; cDepartment of Gastroenterology and Hepatology, VU University Medical Center, Amsterdam 1081 HZ, The Netherlands; and dDepartment of Pediatrics, Leiden University Medical Center, Leiden 2333 ZA, The Netherlands Contributed by Jon J. van Rood, December 7, 2016 (sent for review January 6, 2016; reviewed by Georg Gasteiger and Bana Jabri) − Refractory celiac disease type II (RCDII) is a severe complication of lymphoma, because the Lin IELs expanded in RCDII often give celiac disease (CD) characterized by the presence of an enlarged rise to type I enteropathy-associated T-cell lymphoma (EATL). − clonal population of innate intraepithelial lymphocytes (IELs) lacking The main treatment goal in RCDII is to eliminate the Lin IEL − classical B-, T-, and natural killer (NK)-cell lineage markers (Lin IELs) population before its transformation into a high-grade lymphoma. in the duodenum. In ∼50% of patients with RCDII, these Lin−IELs Cladribine (2-CDA) is thought to be especially active against low- develop into a lymphoma for which no effective treatment is avail- grade malignancies with limited proliferative capacity, and reduces − able. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Single-Cell RNA Sequencing Demonstrates the Molecular and Cellular Reprogramming of Metastatic Lung Adenocarcinoma
ARTICLE https://doi.org/10.1038/s41467-020-16164-1 OPEN Single-cell RNA sequencing demonstrates the molecular and cellular reprogramming of metastatic lung adenocarcinoma Nayoung Kim 1,2,3,13, Hong Kwan Kim4,13, Kyungjong Lee 5,13, Yourae Hong 1,6, Jong Ho Cho4, Jung Won Choi7, Jung-Il Lee7, Yeon-Lim Suh8,BoMiKu9, Hye Hyeon Eum 1,2,3, Soyean Choi 1, Yoon-La Choi6,10,11, Je-Gun Joung1, Woong-Yang Park 1,2,6, Hyun Ae Jung12, Jong-Mu Sun12, Se-Hoon Lee12, ✉ ✉ Jin Seok Ahn12, Keunchil Park12, Myung-Ju Ahn 12 & Hae-Ock Lee 1,2,3,6 1234567890():,; Advanced metastatic cancer poses utmost clinical challenges and may present molecular and cellular features distinct from an early-stage cancer. Herein, we present single-cell tran- scriptome profiling of metastatic lung adenocarcinoma, the most prevalent histological lung cancer type diagnosed at stage IV in over 40% of all cases. From 208,506 cells populating the normal tissues or early to metastatic stage cancer in 44 patients, we identify a cancer cell subtype deviating from the normal differentiation trajectory and dominating the metastatic stage. In all stages, the stromal and immune cell dynamics reveal ontological and functional changes that create a pro-tumoral and immunosuppressive microenvironment. Normal resident myeloid cell populations are gradually replaced with monocyte-derived macrophages and dendritic cells, along with T-cell exhaustion. This extensive single-cell analysis enhances our understanding of molecular and cellular dynamics in metastatic lung cancer and reveals potential diagnostic and therapeutic targets in cancer-microenvironment interactions. 1 Samsung Genome Institute, Samsung Medical Center, Seoul 06351, Korea. -

B Cell Checkpoints in Autoimmune Rheumatic Diseases
REVIEWS B cell checkpoints in autoimmune rheumatic diseases Samuel J. S. Rubin1,2,3, Michelle S. Bloom1,2,3 and William H. Robinson1,2,3* Abstract | B cells have important functions in the pathogenesis of autoimmune diseases, including autoimmune rheumatic diseases. In addition to producing autoantibodies, B cells contribute to autoimmunity by serving as professional antigen- presenting cells (APCs), producing cytokines, and through additional mechanisms. B cell activation and effector functions are regulated by immune checkpoints, including both activating and inhibitory checkpoint receptors that contribute to the regulation of B cell tolerance, activation, antigen presentation, T cell help, class switching, antibody production and cytokine production. The various activating checkpoint receptors include B cell activating receptors that engage with cognate receptors on T cells or other cells, as well as Toll-like receptors that can provide dual stimulation to B cells via co- engagement with the B cell receptor. Furthermore, various inhibitory checkpoint receptors, including B cell inhibitory receptors, have important functions in regulating B cell development, activation and effector functions. Therapeutically targeting B cell checkpoints represents a promising strategy for the treatment of a variety of autoimmune rheumatic diseases. Antibody- dependent B cells are multifunctional lymphocytes that contribute that serve as precursors to and thereby give rise to acti- cell- mediated cytotoxicity to the pathogenesis of autoimmune diseases -

Human CD123 / IL3RA Protein (Fc Tag)
Human CD123 / IL3RA Protein (Fc Tag) Catalog Number: 10518-H02H General Information SDS-PAGE: Gene Name Synonym: CD123; hIL-3Ra; IL3R; IL3RAY; IL3RX; IL3RY Protein Construction: A DNA sequence encoding the human IL3RA (NP_002174.1) (Met1- Arg305) was expressed with the Fc region of human IgG1 at the C- terminus. Source: Human Expression Host: HEK293 Cells QC Testing Purity: > 95 % as determined by SDS-PAGE. Endotoxin: Protein Description < 1.0 EU per μg protein as determined by the LAL method. Interleukin-3 receptor subunit alpha, also known as IL-3 receptor subunit alpha, IL-3R-alpha, CD123, and IL3RA, is a single-pass type I membrane Stability: protein which belongs to the type I cytokine receptor family and Type 5 subfamily. The specific alpha subunit of the interleukin-3 receptor (IL- Samples are stable for up to twelve months from date of receipt at -70 ℃ 3Ralpha, CD123) is strongly expressed in various leukemic blasts and leukemic stem cells and seems to be an excellent target for the therapy of Predicted N terminal: Thr 19 leukemias. The WSXWS motif of IL3RA appears to be necessary for Molecular Mass: proper protein folding and thereby efficient intracellular transport and cell- surface receptor binding. The box one motif of IL3RA is required for JAK The recombinant human IL3RA consists of 525 amino acids and predicts a interaction and / or activation. IL3RA represents a unique marker for molecular mass of 59.8 kDa. primitive leukemic stem cells. Targeting of IL3RA may be a promising strategy for the preferential ablation of AML cells. Aberrant IL3RA Formulation: expression is a good marker for monitoring of minimal residual disease. -

Review Article Molecular Functions of Thyroid Hormones and Their Clinical Significance in Liver-Related Diseases
Hindawi Publishing Corporation BioMed Research International Volume 2013, Article ID 601361, 16 pages http://dx.doi.org/10.1155/2013/601361 Review Article Molecular Functions of Thyroid Hormones and Their Clinical Significance in Liver-Related Diseases Hsiang Cheng Chi,1 Cheng-Yi Chen,1 Ming-Ming Tsai,2 Chung-Ying Tsai,1 and Kwang-Huei Lin1 1 Department of Biochemistry, School of Medicine, Chang-Gung University, Taoyuan 333, Taiwan 2 Department of Nursing, Chang-Gung University of Science and Technology, Taoyuan 333, Taiwan Correspondence should be addressed to Kwang-Huei Lin; [email protected] Received 4 February 2013; Revised 14 May 2013; Accepted 28 May 2013 Academic Editor: Elena Orlova Copyright © 2013 Hsiang Cheng Chi et al. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. Thyroid hormones (THs) are potent mediators of several physiological processes, including embryonic development, cellular differentiation, metabolism, and cell growth. Triiodothyronine3 (T ) is the most biologically active TH form. Thyroid hormone receptors (TRs) belong to the nuclear receptor superfamily and mediate the biological functions of T3 via transcriptional regulation. TRs generally form heterodimers with the retinoid X receptor (RXR) and regulate target genes upon T3 stimulation. Research over the past few decades has revealed that disruption of cellular TH signaling triggers chronic liver diseases, including alcoholic or nonalcoholic fatty liver disease and hepatocellular carcinoma (HCC). Animal model experiments and epidemiologic studies to date imply close associations between high TH levels and prevention of liver disease. -

Cd4/Cd8 Panel, Blood
Lab Dept: Flow and Immunology Test Name: CD4/CD8 PANEL, BLOOD General Information Lab Order Codes: C48P Synonyms: Helper/Suppressor Ratio; T- cells; T-cell subsets; T-cell phenotyping See also: Immune Status Panel and Comprehensive Immune Status Panel CPT Codes: 86359 – T cells, total count 86360 – T cells; absolute CD4 and CD8 count, including ratio Test Includes: CD4(CD3+) and CD8(CD3+) relative percentages, absolute values and a calculated Helper/Suppressor ratio. Logistics Test Indications: This is a minimal antibody panel to monitor immune status. Lab Testing Sections: Flow Cytometry Phone Numbers: MIN Lab: 612-813-6280 STP Lab: 651-220-6550 Test Availability: 3 times weekly determined by volume. Transport collected specimen immediately to Flow Cytometry. Routine testing is not available on weekends or holidays. Therefore, specimens cannot be used if drawn the day before a 3 day weekend such as Memorial Day, Labor Day or major holiday that falls on a Monday or Friday. Turnaround Time: 1 – 3 days Special Instructions: See Test Availability Specimen Specimen Type: Whole blood Container: Lavender top (EDTA) tube Draw Volume: 2 mL blood in a 2 mL Lavender (EDTA) tube Minimum volume: 0.5 mL in an EDTA microtainer Collection: Routine venipuncture Special Processing: Keep sample at room temperature and forward promptly to the laboratory. Do not centrifuge, refrigerate, or freeze sample. Patient Preparation: None Sample Rejection: Specimens will not be processed that are clotted; hemolyzed; greater than 72 hours old; collected in the wrong tube type (0.5 mL in a 2 mL tube), or that have been held or handled at a temperature other than room temperature Interpretive Reference Range: Age-dependant reference ranges provided with results. -

Rational Design of a CD4 Mimic That Inhibits HIV-1 Entry and Exposes Cryptic Neutralization Epitopes
RESEARCH ARTICLE Rational design of a CD4 mimic that inhibits HIV-1 entry and exposes cryptic neutralization epitopes Loïc Martin1, François Stricher1, Dorothée Missé2, Francesca Sironi3, Martine Pugnière4, Philippe Barthe5, Rafael Prado-Gotor5, Isabelle Freulon1, Xavier Magne2, Christian Roumestand5, André Ménez1, Paolo Lusso3, Francisco Veas2, and Claudio Vita1* Published online 16 December 2002; doi:10.1038/nbt768 The conserved surfaces of the human immunodeficiency virus (HIV)-1 envelope involved in receptor binding represent potential targets for the development of entry inhibitors and neutralizing antibodies. Using structural information on a CD4-gp120-17b antibody complex, we have designed a 27-amino acid CD4 mimic, CD4M33, that presents optimal interactions with gp120 and binds to viral particles and diverse HIV-1 envelopes with CD4-like affinity.This mini-CD4 inhibits infection of both immortalized and primary cells by HIV-1, including pri- http://www.nature.com/naturebiotechnology mary patient isolates that are generally resistant to inhibition by soluble CD4. Furthermore, CD4M33 pos- sesses functional properties of CD4, including the ability to unmask conserved neutralization epitopes of gp120 that are cryptic on the unbound glycoprotein. CD4M33 is a prototype of inhibitors of HIV-1 entry and, in complex with envelope proteins, a potential component of vaccine formulations, or a molecular target in phage display technology to develop broad-spectrum neutralizing antibodies. HIV-1 avoids the host immune surveillance by exposing -

Immune Checkpoint Blockade Enhances Immune Activity of Therapeutic Lung Cancer Vaccine
Article Immune Checkpoint Blockade Enhances Immune Activity of Therapeutic Lung Cancer Vaccine Pournima Kadam 1, Ram P. Singh 1, Michael Davoodi 1 , Jay M. Lee 2,3, Maie St. John 3,4,5 and Sherven Sharma 1,2,3,* 1 Department of Medicine, Veterans Affairs Greater Los Angeles Healthcare System, Los Angeles, CA 90073, USA; [email protected] (P.K.); [email protected] (R.P.S.); [email protected] (M.D.) 2 Department of Medicine, UCLA Lung Cancer Research Program, David Geffen School of Medicine at UCLA, Los Angeles, CA 90095, USA; [email protected] 3 Jonsson Comprehensive Cancer Center, David Geffen School of Medicine at UCLA, Los Angeles, CA 90095, USA; [email protected] 4 Department of Head and Neck Surgery, David Geffen School of Medicine at UCLA, Los Angeles, CA 90095, USA 5 UCLA Head and Neck Cancer Program, David Geffen School of Medicine at UCLA, Los Angeles, CA 90095, USA * Correspondence: [email protected]; Tel.: +1-310-268-4296; Fax: +1-310-268-4807 Received: 31 August 2020; Accepted: 3 October 2020; Published: 5 November 2020 Abstract: Background: Immune checkpoint blockade that downregulates T cell evasion for effective immunity has provided a renewed interest in therapeutic cancer vaccines. Methods: Utilizing murine lung cancer models, we determined: tumor burden, TIL cytolysis, immunohistochemistry, flow cytometry, RNA Sequencing, CD4 T cells, CD8 T cells, CXCL9 chemokine, and CXCL10 chemokine neutralization to evaluate the efficacy of Programmed cell death protein 1 (PD-1) blockade combined with chemokine (C-C motif) ligand 21-dendritic cell tumor antigen (CCL21-DC tumor Ag) vaccine.