Xo GENE PANEL
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Limited Presence of IL-22 Binding Protein, a Natural IL-22 Inhibitor, Strengthens Psoriatic Skin Inflammation
Limited Presence of IL-22 Binding Protein, a Natural IL-22 Inhibitor, Strengthens Psoriatic Skin Inflammation This information is current as Jérôme C. Martin, Kerstin Wolk, Gaëlle Bériou, Ahmed of September 25, 2021. Abidi, Ellen Witte-Händel, Cédric Louvet, Georgios Kokolakis, Lucile Drujont, Laure Dumoutier, Jean-Christophe Renauld, Robert Sabat and Régis Josien J Immunol 2017; 198:3671-3678; Prepublished online 29 March 2017; Downloaded from doi: 10.4049/jimmunol.1700021 http://www.jimmunol.org/content/198/9/3671 References This article cites 47 articles, 12 of which you can access for free at: http://www.jimmunol.org/ http://www.jimmunol.org/content/198/9/3671.full#ref-list-1 Why The JI? Submit online. • Rapid Reviews! 30 days* from submission to initial decision • No Triage! Every submission reviewed by practicing scientists by guest on September 25, 2021 • Fast Publication! 4 weeks from acceptance to publication *average Subscription Information about subscribing to The Journal of Immunology is online at: http://jimmunol.org/subscription Permissions Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html Email Alerts Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts The Journal of Immunology is published twice each month by The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2017 by The American Association of Immunologists, Inc. All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. The Journal of Immunology Limited Presence of IL-22 Binding Protein, a Natural IL-22 Inhibitor, Strengthens Psoriatic Skin Inflammation Je´roˆme C. -
(12) Patent Application Publication (10) Pub. No.: US 2012/0070450 A1 Ishikawa Et Al
US 20120070450A1 (19) United States (12) Patent Application Publication (10) Pub. No.: US 2012/0070450 A1 Ishikawa et al. (43) Pub. Date: Mar. 22, 2012 (54) LEUKEMA STEM CELLMARKERS Publication Classification (51) Int. Cl. A 6LX 39/395 (2006.01) (75) Inventors: Fumihiko Ishikawa, Kanagawa CI2O I/68 (2006.01) (JP): Osamu Ohara, Kanagawa GOIN 2L/64 (2006.01) (JP); Yoriko Saito, Kanagawa (JP); A6IP35/02 (2006.01) Hiroshi Kitamura, Kanagawa (JP); C40B 30/04 (2006.01) Atsushi Hijikata, Kanagawa (JP); A63L/7088 (2006.01) Hidetoshi Ozawa, Kanagawa (JP); C07K 6/8 (2006.01) Leonard D. Shultz, Bar Harbor, C7H 2L/00 (2006.01) A6II 35/12 (2006.01) ME (US) CI2N 5/078 (2010.01) (52) U.S. Cl. .................. 424/173.1; 424/178.1; 424/93.7: (73) Assignee: RIKEN, Wako-shi (JP) 435/6.14; 435/723; 435/375; 506/9: 514/44 A: 530/389.6; 530/391.7:536/24.5 (57) ABSTRACT (21) Appl. No.: 13/258,993 The invention provides a test method for predicting the initial onset or a recurrence of acute myeloid leukemia (AML) com PCT Fled: prising (1) measuring the expression level of human leukemic (22) Mar. 24, 2010 stem cell (LSC) marker genes in a biological sample collected from a Subject for a transcription product or translation prod uct of the gene as an analyte and (2) comparing the expression (86) PCT NO.: PCT/UP2010/0551.31 level with a reference value; an LSC-targeting therapeutic agent for AML capable of Suppressing the expression of a S371 (c)(1), gene selected from among LSC marker genes or a Substance (2), (4) Date: Dec. -

Table S1. List of Proteins in the BAHD1 Interactome
Table S1. List of proteins in the BAHD1 interactome BAHD1 nuclear partners found in this work yeast two-hybrid screen Name Description Function Reference (a) Chromatin adapters HP1α (CBX5) chromobox homolog 5 (HP1 alpha) Binds histone H3 methylated on lysine 9 and chromatin-associated proteins (20-23) HP1β (CBX1) chromobox homolog 1 (HP1 beta) Binds histone H3 methylated on lysine 9 and chromatin-associated proteins HP1γ (CBX3) chromobox homolog 3 (HP1 gamma) Binds histone H3 methylated on lysine 9 and chromatin-associated proteins MBD1 methyl-CpG binding domain protein 1 Binds methylated CpG dinucleotide and chromatin-associated proteins (22, 24-26) Chromatin modification enzymes CHD1 chromodomain helicase DNA binding protein 1 ATP-dependent chromatin remodeling activity (27-28) HDAC5 histone deacetylase 5 Histone deacetylase activity (23,29,30) SETDB1 (ESET;KMT1E) SET domain, bifurcated 1 Histone-lysine N-methyltransferase activity (31-34) Transcription factors GTF3C2 general transcription factor IIIC, polypeptide 2, beta 110kDa Required for RNA polymerase III-mediated transcription HEYL (Hey3) hairy/enhancer-of-split related with YRPW motif-like DNA-binding transcription factor with basic helix-loop-helix domain (35) KLF10 (TIEG1) Kruppel-like factor 10 DNA-binding transcription factor with C2H2 zinc finger domain (36) NR2F1 (COUP-TFI) nuclear receptor subfamily 2, group F, member 1 DNA-binding transcription factor with C4 type zinc finger domain (ligand-regulated) (36) PEG3 paternally expressed 3 DNA-binding transcription factor with -

Activated Peripheral-Blood-Derived Mononuclear Cells
Transcription factor expression in lipopolysaccharide- activated peripheral-blood-derived mononuclear cells Jared C. Roach*†, Kelly D. Smith*‡, Katie L. Strobe*, Stephanie M. Nissen*, Christian D. Haudenschild§, Daixing Zhou§, Thomas J. Vasicek¶, G. A. Heldʈ, Gustavo A. Stolovitzkyʈ, Leroy E. Hood*†, and Alan Aderem* *Institute for Systems Biology, 1441 North 34th Street, Seattle, WA 98103; ‡Department of Pathology, University of Washington, Seattle, WA 98195; §Illumina, 25861 Industrial Boulevard, Hayward, CA 94545; ¶Medtronic, 710 Medtronic Parkway, Minneapolis, MN 55432; and ʈIBM Computational Biology Center, P.O. Box 218, Yorktown Heights, NY 10598 Contributed by Leroy E. Hood, August 21, 2007 (sent for review January 7, 2007) Transcription factors play a key role in integrating and modulating system. In this model system, we activated peripheral-blood-derived biological information. In this study, we comprehensively measured mononuclear cells, which can be loosely termed ‘‘macrophages,’’ the changing abundances of mRNAs over a time course of activation with lipopolysaccharide (LPS). We focused on the precise mea- of human peripheral-blood-derived mononuclear cells (‘‘macro- surement of mRNA concentrations. There is currently no high- phages’’) with lipopolysaccharide. Global and dynamic analysis of throughput technology that can precisely and sensitively measure all transcription factors in response to a physiological stimulus has yet to mRNAs in a system, although such technologies are likely to be be achieved in a human system, and our efforts significantly available in the near future. To demonstrate the potential utility of advanced this goal. We used multiple global high-throughput tech- such technologies, and to motivate their development and encour- nologies for measuring mRNA levels, including massively parallel age their use, we produced data from a combination of two distinct signature sequencing and GeneChip microarrays. -

Supplementary Material
Coagulation & Its Disorders SUPPLEMENTARY APPENDIX Dexamethasone promotes durable factor VIII-specific tolerance in hemophilia A mice via thymic mechanisms Maria T. Georgescu, 1 Paul C. Moorehead, 2,3 Alice S. van Velzen, 4 Kate Nesbitt, 1 Birgit M. Reipert, 5 Katharina N. Steinitz, 5 Maria Schuster, 5 Christine Hough 1 and David Lillicrap 1 1Department of Pathology and Molecular Medicine, Queen’s University, Kingston, ON, Canada; 2Janeway Children’s Health and Reha - bilitation Centre, St. John’s, NL, Canada; 3Faculty of Medicine, Memorial University, St. John’s, NL, Canada; 4Department of Pediatric Hematology, Immunology and Infectious Diseases, Emma Children’s Hospital, Amsterdam, the Netherlands and 5Baxalta Innovations GmbH, Vienna, Austria ©2018 Ferrata Storti Foundation. This is an open-access paper. doi:10.3324/haematol. 2018.189852 Received: January 30, 2018. Accepted: April 19, 2018. Pre-published: April 19, 2018. Correspondence: [email protected] Supplementary Material 100 80 60 40 20 Anti-VWF IgG Positive IgG Mice (%)Anti-VWF 0 FVIII/FVIII FVIII+Dex/FVIII FVIII+Dex/intFVIII+FVIII n=4 n=13 n=12 Figure S1. Administration of Dex during initial FVIII exposure does not impair the immune response to VWF. Anti-VWF IgG incidence at week 22, following exposure to VWF (Week 18-21) in mice initially treated with FVIII or FVIII+Dex and with no evidence of anti-FVIII IgG at week 4. Table S1. Genes down-regulated following FVIII+Dex treatment. Transcript Count Ratio Gene Name FVIII+Dex vs FVIII Dex vs HBSS Ccr4 -5.82 -14.45 Rag1 -4.18 -16.00 Cd69 -3.51 -4.40 Ccr9 -3.46 -9.90 Slamf1 -3.20 -6.50 Cd8b1 -3.18 -7.77 Cd40lg -2.99 -2.56 Cxcl1 -2.90 -2.37 Rag2 -2.84 -9.21 Rorc -2.78 -5.65 Cd8a -2.67 -7.74 Cd4 -2.66 -5.06 Il9 -2.64 -2.26 Il12b -2.60 -2.49 Bcl6 -2.59 -3.30 Il27 -2.54 -3.06 Mr1 -2.48 -3.26 Sh2d1a -2.39 -6.68 Il13 -2.23 -2.38 Ccl22 -2.22 -2.51 Il16 -2.22 -3.05 Socs1 -2.20 -4.55 Card9 -2.12 -3.32 Cxcr4 -2.12 -3.98 Tcf7 -2.12 -5.16 Lck -2.06 -4.47 Icam2 -2.02 -3.09 Table S2. -

Molecular Profile of Tumor-Specific CD8+ T Cell Hypofunction in a Transplantable Murine Cancer Model
Downloaded from http://www.jimmunol.org/ by guest on September 25, 2021 T + is online at: average * The Journal of Immunology , 34 of which you can access for free at: 2016; 197:1477-1488; Prepublished online 1 July from submission to initial decision 4 weeks from acceptance to publication 2016; doi: 10.4049/jimmunol.1600589 http://www.jimmunol.org/content/197/4/1477 Molecular Profile of Tumor-Specific CD8 Cell Hypofunction in a Transplantable Murine Cancer Model Katherine A. Waugh, Sonia M. Leach, Brandon L. Moore, Tullia C. Bruno, Jonathan D. Buhrman and Jill E. Slansky J Immunol cites 95 articles Submit online. Every submission reviewed by practicing scientists ? is published twice each month by Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts http://jimmunol.org/subscription Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html http://www.jimmunol.org/content/suppl/2016/07/01/jimmunol.160058 9.DCSupplemental This article http://www.jimmunol.org/content/197/4/1477.full#ref-list-1 Information about subscribing to The JI No Triage! Fast Publication! Rapid Reviews! 30 days* Why • • • Material References Permissions Email Alerts Subscription Supplementary The Journal of Immunology The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2016 by The American Association of Immunologists, Inc. All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. This information is current as of September 25, 2021. The Journal of Immunology Molecular Profile of Tumor-Specific CD8+ T Cell Hypofunction in a Transplantable Murine Cancer Model Katherine A. -
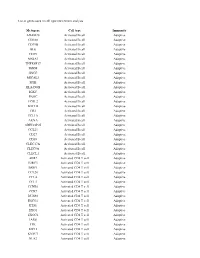
List of Genes Used in Cell Type Enrichment Analysis
List of genes used in cell type enrichment analysis Metagene Cell type Immunity ADAM28 Activated B cell Adaptive CD180 Activated B cell Adaptive CD79B Activated B cell Adaptive BLK Activated B cell Adaptive CD19 Activated B cell Adaptive MS4A1 Activated B cell Adaptive TNFRSF17 Activated B cell Adaptive IGHM Activated B cell Adaptive GNG7 Activated B cell Adaptive MICAL3 Activated B cell Adaptive SPIB Activated B cell Adaptive HLA-DOB Activated B cell Adaptive IGKC Activated B cell Adaptive PNOC Activated B cell Adaptive FCRL2 Activated B cell Adaptive BACH2 Activated B cell Adaptive CR2 Activated B cell Adaptive TCL1A Activated B cell Adaptive AKNA Activated B cell Adaptive ARHGAP25 Activated B cell Adaptive CCL21 Activated B cell Adaptive CD27 Activated B cell Adaptive CD38 Activated B cell Adaptive CLEC17A Activated B cell Adaptive CLEC9A Activated B cell Adaptive CLECL1 Activated B cell Adaptive AIM2 Activated CD4 T cell Adaptive BIRC3 Activated CD4 T cell Adaptive BRIP1 Activated CD4 T cell Adaptive CCL20 Activated CD4 T cell Adaptive CCL4 Activated CD4 T cell Adaptive CCL5 Activated CD4 T cell Adaptive CCNB1 Activated CD4 T cell Adaptive CCR7 Activated CD4 T cell Adaptive DUSP2 Activated CD4 T cell Adaptive ESCO2 Activated CD4 T cell Adaptive ETS1 Activated CD4 T cell Adaptive EXO1 Activated CD4 T cell Adaptive EXOC6 Activated CD4 T cell Adaptive IARS Activated CD4 T cell Adaptive ITK Activated CD4 T cell Adaptive KIF11 Activated CD4 T cell Adaptive KNTC1 Activated CD4 T cell Adaptive NUF2 Activated CD4 T cell Adaptive PRC1 Activated -

CCAAT/Enhancer Binding Protein Epsilon) Thomas Burmeister Charite, Med
Atlas of Genetics and Cytogenetics in Oncology and Haematology OPEN ACCESS JOURNAL INIST-CNRS Gene Section Short Communication CEBPE (CCAAT/enhancer binding protein epsilon) Thomas Burmeister Charite, Med. Klinik fur Hamatologie, Onkologie und Tumorimmunologie, Hindenburgdamm 30, 12200 Berlin, Germany; [email protected] Published in Atlas Database: March 2017 Online updated version : http://AtlasGeneticsOncology.org/Genes/CEBPEID42984ch14q11.html Printable original version : http://documents.irevues.inist.fr/bitstream/handle/2042/69005/03-2017-CEBPEID42984ch14q11.pdf DOI: 10.4267/2042/69005 This work is licensed under a Creative Commons Attribution-Noncommercial-No Derivative Works 2.0 France Licence. © 2017 Atlas of Genetics and Cytogenetics in Oncology and Haematology alternative 3-exon-organization of the human Abstract CEBPE gene (Figure 1b). However, exon 1, as described by Yamanaka et al. contains a frameshift Review on CEBPE, with data on DNA, on the according to the GRCh38.p7 NCBI assembly. protein encoded, and where the gene is implicated. Transcription Keywords CEBPE; Transcription factor; Neutrophil specific Various transcripts have been reported, resulting in granule deficiency; Acute lymphoblastic leukemia; four protein isoforms (Lekstrom-Himes 2001, Translocation. Yamanaka 1997; Figure 1c). All transcripts share a common 3' end. Identity Protein Other names: CRP1 Description HGNC (Hugo): CEBPE CEBPE is a member of the CCAAT/enhancer- Location: 14q11.2 binding protein (C/EBP) family, which also Location (base pair) includes CEBPA, CEBPB, CEBPG, CEBPD and Starts at 23117306 and ends at 23119611 bp from CEBPZ (Ramji & Foka; 2002). A common pter (according to GRCh38.p7 Annotation Release structural feature of the C/EBP proteins is the 108, May 5 2016) presence of a highly conserved 55-65 amino acid sequence at the C-terminus which encodes a basic DNA/RNA leucine zipper motif (bZIP domain) that functions as a dimerization domain. -

Interplay Between Epigenetics and Metabolism in Oncogenesis: Mechanisms and Therapeutic Approaches
OPEN Oncogene (2017) 36, 3359–3374 www.nature.com/onc REVIEW Interplay between epigenetics and metabolism in oncogenesis: mechanisms and therapeutic approaches CC Wong1, Y Qian2,3 and J Yu1 Epigenetic and metabolic alterations in cancer cells are highly intertwined. Oncogene-driven metabolic rewiring modifies the epigenetic landscape via modulating the activities of DNA and histone modification enzymes at the metabolite level. Conversely, epigenetic mechanisms regulate the expression of metabolic genes, thereby altering the metabolome. Epigenetic-metabolomic interplay has a critical role in tumourigenesis by coordinately sustaining cell proliferation, metastasis and pluripotency. Understanding the link between epigenetics and metabolism could unravel novel molecular targets, whose intervention may lead to improvements in cancer treatment. In this review, we summarized the recent discoveries linking epigenetics and metabolism and their underlying roles in tumorigenesis; and highlighted the promising molecular targets, with an update on the development of small molecule or biologic inhibitors against these abnormalities in cancer. Oncogene (2017) 36, 3359–3374; doi:10.1038/onc.2016.485; published online 16 January 2017 INTRODUCTION metabolic genes have also been identified as driver genes It has been appreciated since the early days of cancer research mutated in some cancers, such as isocitrate dehydrogenase 1 16 17 that the metabolic profiles of tumor cells differ significantly from and 2 (IDH1/2) in gliomas and acute myeloid leukemia (AML), 18 normal cells. Cancer cells have high metabolic demands and they succinate dehydrogenase (SDH) in paragangliomas and fuma- utilize nutrients with an altered metabolic program to support rate hydratase (FH) in hereditary leiomyomatosis and renal cell 19 their high proliferative rates and adapt to the hostile tumor cancer (HLRCC). -

The Potential of Histone Demethylases As Therapeutic Targets
Pharmaceuticals 2012, 5, 963-990; doi:10.3390/ph5090963 OPEN ACCESS pharmaceuticals ISSN 1424-8247 www.mdpi.com/journal/pharmaceuticals Review Epigenetic Control and Cancer: The Potential of Histone Demethylases as Therapeutic Targets Fernando Lizcano * and Jeison Garcia Center of Biomedical Research La Sabana University-CIBUS, School of Medicine, Universidad de La Sabana, Campus Del Puente del Común, km 7 Autopista Norte de Bogota, Chía 250001, Colombia; E-Mail: [email protected] (J.G.) * Author to whom correspondence should be addressed; E-Mail: [email protected]; Tel.: +571-861-5555 (ext. 23328); Fax: +571-861-5555 (ext. 10111). Received: 26 June 2012; in revised form: 21 July 2012 / Accepted: 17 August 2012 / Published: 12 September 2012 Abstract: The development of cancer involves an immense number of factors at the molecular level. These factors are associated principally with alterations in the epigenetic mechanisms that regulate gene expression profiles. Studying the effects of chromatin structure alterations, which are caused by the addition/removal of functional groups to specific histone residues, are of great interest as a promising way to identify markers for cancer diagnosis, classify the disease and determine its prognosis, and these markers could be potential targets for the treatment of this disease in its different forms. This manuscript presents the current point of view regarding members of the recently described family of proteins that exhibit histone demethylase activity; histone demethylases are genetic regulators that play a fundamental role in both the activation and repression of genes and whose expression has been observed to increase in many types of cancer. -

Mutant IDH, (R)-2-Hydroxyglutarate, and Cancer
Downloaded from genesdev.cshlp.org on October 1, 2021 - Published by Cold Spring Harbor Laboratory Press REVIEW What a difference a hydroxyl makes: mutant IDH, (R)-2-hydroxyglutarate, and cancer Julie-Aurore Losman1 and William G. Kaelin Jr.1,2,3 1Department of Medical Oncology, Dana-Farber Cancer Institute, Brigham and Women’s Hospital, Harvard Medical School, Boston, Massachusetts 02215, USA; 2Howard Hughes Medical Institute, Chevy Chase, Maryland 20815, USA Mutations in metabolic enzymes, including isocitrate whether altered cellular metabolism is a cause of cancer dehydrogenase 1 (IDH1) and IDH2, in cancer strongly or merely an adaptive response of cancer cells in the face implicate altered metabolism in tumorigenesis. IDH1 of accelerated cell proliferation is still a topic of some and IDH2 catalyze the interconversion of isocitrate and debate. 2-oxoglutarate (2OG). 2OG is a TCA cycle intermediate The recent identification of cancer-associated muta- and an essential cofactor for many enzymes, including tions in three metabolic enzymes suggests that altered JmjC domain-containing histone demethylases, TET cellular metabolism can indeed be a cause of some 5-methylcytosine hydroxylases, and EglN prolyl-4-hydrox- cancers (Pollard et al. 2003; King et al. 2006; Raimundo ylases. Cancer-associated IDH mutations alter the enzymes et al. 2011). Two of these enzymes, fumarate hydratase such that they reduce 2OG to the structurally similar (FH) and succinate dehydrogenase (SDH), are bone fide metabolite (R)-2-hydroxyglutarate [(R)-2HG]. Here we tumor suppressors, and loss-of-function mutations in FH review what is known about the molecular mechanisms and SDH have been identified in various cancers, in- of transformation by mutant IDH and discuss their im- cluding renal cell carcinomas and paragangliomas. -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated.