CD4 T-Cell Cytokines Synergize to Induce Proliferation of Malignant and Nonmalignant Innate Intraepithelial Lymphocytes
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-
(12) Patent Application Publication (10) Pub. No.: US 2012/0070450 A1 Ishikawa Et Al
US 20120070450A1 (19) United States (12) Patent Application Publication (10) Pub. No.: US 2012/0070450 A1 Ishikawa et al. (43) Pub. Date: Mar. 22, 2012 (54) LEUKEMA STEM CELLMARKERS Publication Classification (51) Int. Cl. A 6LX 39/395 (2006.01) (75) Inventors: Fumihiko Ishikawa, Kanagawa CI2O I/68 (2006.01) (JP): Osamu Ohara, Kanagawa GOIN 2L/64 (2006.01) (JP); Yoriko Saito, Kanagawa (JP); A6IP35/02 (2006.01) Hiroshi Kitamura, Kanagawa (JP); C40B 30/04 (2006.01) Atsushi Hijikata, Kanagawa (JP); A63L/7088 (2006.01) Hidetoshi Ozawa, Kanagawa (JP); C07K 6/8 (2006.01) Leonard D. Shultz, Bar Harbor, C7H 2L/00 (2006.01) A6II 35/12 (2006.01) ME (US) CI2N 5/078 (2010.01) (52) U.S. Cl. .................. 424/173.1; 424/178.1; 424/93.7: (73) Assignee: RIKEN, Wako-shi (JP) 435/6.14; 435/723; 435/375; 506/9: 514/44 A: 530/389.6; 530/391.7:536/24.5 (57) ABSTRACT (21) Appl. No.: 13/258,993 The invention provides a test method for predicting the initial onset or a recurrence of acute myeloid leukemia (AML) com PCT Fled: prising (1) measuring the expression level of human leukemic (22) Mar. 24, 2010 stem cell (LSC) marker genes in a biological sample collected from a Subject for a transcription product or translation prod uct of the gene as an analyte and (2) comparing the expression (86) PCT NO.: PCT/UP2010/0551.31 level with a reference value; an LSC-targeting therapeutic agent for AML capable of Suppressing the expression of a S371 (c)(1), gene selected from among LSC marker genes or a Substance (2), (4) Date: Dec. -
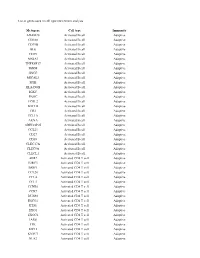
List of Genes Used in Cell Type Enrichment Analysis
List of genes used in cell type enrichment analysis Metagene Cell type Immunity ADAM28 Activated B cell Adaptive CD180 Activated B cell Adaptive CD79B Activated B cell Adaptive BLK Activated B cell Adaptive CD19 Activated B cell Adaptive MS4A1 Activated B cell Adaptive TNFRSF17 Activated B cell Adaptive IGHM Activated B cell Adaptive GNG7 Activated B cell Adaptive MICAL3 Activated B cell Adaptive SPIB Activated B cell Adaptive HLA-DOB Activated B cell Adaptive IGKC Activated B cell Adaptive PNOC Activated B cell Adaptive FCRL2 Activated B cell Adaptive BACH2 Activated B cell Adaptive CR2 Activated B cell Adaptive TCL1A Activated B cell Adaptive AKNA Activated B cell Adaptive ARHGAP25 Activated B cell Adaptive CCL21 Activated B cell Adaptive CD27 Activated B cell Adaptive CD38 Activated B cell Adaptive CLEC17A Activated B cell Adaptive CLEC9A Activated B cell Adaptive CLECL1 Activated B cell Adaptive AIM2 Activated CD4 T cell Adaptive BIRC3 Activated CD4 T cell Adaptive BRIP1 Activated CD4 T cell Adaptive CCL20 Activated CD4 T cell Adaptive CCL4 Activated CD4 T cell Adaptive CCL5 Activated CD4 T cell Adaptive CCNB1 Activated CD4 T cell Adaptive CCR7 Activated CD4 T cell Adaptive DUSP2 Activated CD4 T cell Adaptive ESCO2 Activated CD4 T cell Adaptive ETS1 Activated CD4 T cell Adaptive EXO1 Activated CD4 T cell Adaptive EXOC6 Activated CD4 T cell Adaptive IARS Activated CD4 T cell Adaptive ITK Activated CD4 T cell Adaptive KIF11 Activated CD4 T cell Adaptive KNTC1 Activated CD4 T cell Adaptive NUF2 Activated CD4 T cell Adaptive PRC1 Activated -

Single-Cell RNA Sequencing Demonstrates the Molecular and Cellular Reprogramming of Metastatic Lung Adenocarcinoma
ARTICLE https://doi.org/10.1038/s41467-020-16164-1 OPEN Single-cell RNA sequencing demonstrates the molecular and cellular reprogramming of metastatic lung adenocarcinoma Nayoung Kim 1,2,3,13, Hong Kwan Kim4,13, Kyungjong Lee 5,13, Yourae Hong 1,6, Jong Ho Cho4, Jung Won Choi7, Jung-Il Lee7, Yeon-Lim Suh8,BoMiKu9, Hye Hyeon Eum 1,2,3, Soyean Choi 1, Yoon-La Choi6,10,11, Je-Gun Joung1, Woong-Yang Park 1,2,6, Hyun Ae Jung12, Jong-Mu Sun12, Se-Hoon Lee12, ✉ ✉ Jin Seok Ahn12, Keunchil Park12, Myung-Ju Ahn 12 & Hae-Ock Lee 1,2,3,6 1234567890():,; Advanced metastatic cancer poses utmost clinical challenges and may present molecular and cellular features distinct from an early-stage cancer. Herein, we present single-cell tran- scriptome profiling of metastatic lung adenocarcinoma, the most prevalent histological lung cancer type diagnosed at stage IV in over 40% of all cases. From 208,506 cells populating the normal tissues or early to metastatic stage cancer in 44 patients, we identify a cancer cell subtype deviating from the normal differentiation trajectory and dominating the metastatic stage. In all stages, the stromal and immune cell dynamics reveal ontological and functional changes that create a pro-tumoral and immunosuppressive microenvironment. Normal resident myeloid cell populations are gradually replaced with monocyte-derived macrophages and dendritic cells, along with T-cell exhaustion. This extensive single-cell analysis enhances our understanding of molecular and cellular dynamics in metastatic lung cancer and reveals potential diagnostic and therapeutic targets in cancer-microenvironment interactions. 1 Samsung Genome Institute, Samsung Medical Center, Seoul 06351, Korea. -

Human CD123 / IL3RA Protein (Fc Tag)
Human CD123 / IL3RA Protein (Fc Tag) Catalog Number: 10518-H02H General Information SDS-PAGE: Gene Name Synonym: CD123; hIL-3Ra; IL3R; IL3RAY; IL3RX; IL3RY Protein Construction: A DNA sequence encoding the human IL3RA (NP_002174.1) (Met1- Arg305) was expressed with the Fc region of human IgG1 at the C- terminus. Source: Human Expression Host: HEK293 Cells QC Testing Purity: > 95 % as determined by SDS-PAGE. Endotoxin: Protein Description < 1.0 EU per μg protein as determined by the LAL method. Interleukin-3 receptor subunit alpha, also known as IL-3 receptor subunit alpha, IL-3R-alpha, CD123, and IL3RA, is a single-pass type I membrane Stability: protein which belongs to the type I cytokine receptor family and Type 5 subfamily. The specific alpha subunit of the interleukin-3 receptor (IL- Samples are stable for up to twelve months from date of receipt at -70 ℃ 3Ralpha, CD123) is strongly expressed in various leukemic blasts and leukemic stem cells and seems to be an excellent target for the therapy of Predicted N terminal: Thr 19 leukemias. The WSXWS motif of IL3RA appears to be necessary for Molecular Mass: proper protein folding and thereby efficient intracellular transport and cell- surface receptor binding. The box one motif of IL3RA is required for JAK The recombinant human IL3RA consists of 525 amino acids and predicts a interaction and / or activation. IL3RA represents a unique marker for molecular mass of 59.8 kDa. primitive leukemic stem cells. Targeting of IL3RA may be a promising strategy for the preferential ablation of AML cells. Aberrant IL3RA Formulation: expression is a good marker for monitoring of minimal residual disease. -

Immune Checkpoint Blockade Enhances Immune Activity of Therapeutic Lung Cancer Vaccine
Article Immune Checkpoint Blockade Enhances Immune Activity of Therapeutic Lung Cancer Vaccine Pournima Kadam 1, Ram P. Singh 1, Michael Davoodi 1 , Jay M. Lee 2,3, Maie St. John 3,4,5 and Sherven Sharma 1,2,3,* 1 Department of Medicine, Veterans Affairs Greater Los Angeles Healthcare System, Los Angeles, CA 90073, USA; [email protected] (P.K.); [email protected] (R.P.S.); [email protected] (M.D.) 2 Department of Medicine, UCLA Lung Cancer Research Program, David Geffen School of Medicine at UCLA, Los Angeles, CA 90095, USA; [email protected] 3 Jonsson Comprehensive Cancer Center, David Geffen School of Medicine at UCLA, Los Angeles, CA 90095, USA; [email protected] 4 Department of Head and Neck Surgery, David Geffen School of Medicine at UCLA, Los Angeles, CA 90095, USA 5 UCLA Head and Neck Cancer Program, David Geffen School of Medicine at UCLA, Los Angeles, CA 90095, USA * Correspondence: [email protected]; Tel.: +1-310-268-4296; Fax: +1-310-268-4807 Received: 31 August 2020; Accepted: 3 October 2020; Published: 5 November 2020 Abstract: Background: Immune checkpoint blockade that downregulates T cell evasion for effective immunity has provided a renewed interest in therapeutic cancer vaccines. Methods: Utilizing murine lung cancer models, we determined: tumor burden, TIL cytolysis, immunohistochemistry, flow cytometry, RNA Sequencing, CD4 T cells, CD8 T cells, CXCL9 chemokine, and CXCL10 chemokine neutralization to evaluate the efficacy of Programmed cell death protein 1 (PD-1) blockade combined with chemokine (C-C motif) ligand 21-dendritic cell tumor antigen (CCL21-DC tumor Ag) vaccine. -

Single-Cell Analysis of Crohn's Disease Lesions Identifies
bioRxiv preprint doi: https://doi.org/10.1101/503102; this version posted December 20, 2018. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Single-cell analysis of Crohn’s disease lesions identifies a pathogenic cellular module associated with resistance to anti-TNF therapy JC Martin1,2,3, G Boschetti1,2,3, C Chang1,2,3, R Ungaro4, M Giri5, LS Chuang5, S Nayar5, A Greenstein6, M. Dubinsky7, L Walker1,2,5,8, A Leader1,2,3, JS Fine9, CE Whitehurst9, L Mbow9, S Kugathasan10, L.A. Denson11, J.Hyams12, JR Friedman13, P Desai13, HM Ko14, I Laface1,2,8, Guray Akturk1,2,8, EE Schadt15,16, S Gnjatic1,2,8, A Rahman1,2,5,8, , M Merad1,2,3,8,17,18*, JH Cho5,17,*, E Kenigsberg1,15,16,17* 1 Precision Immunology Institute, Icahn School of Medicine at Mount Sinai, New York, NY 10029, USA. 2 Tisch Cancer Institute, Icahn School of Medicine at Mount Sinai, New York, NY 10029, USA. 3 Department of Oncological Sciences, Icahn School of Medicine at Mount Sinai, New York, NY 10029, USA. 4 The Dr. Henry D. Janowitz Division of Gastroenterology, Icahn School of Medicine at Mount Sinai, New York City, NY 10029, USA. 5 Charles Bronfman Institute for Personalized Medicine, Icahn School of Medicine at Mount Sinai, New York, NY 10029, USA. 6 Department of Colorectal Surgery, Icahn School of Medicine at Mount Sinai, New York, NY 10029, USA 7 Department of Pediatrics, Susan and Leonard Feinstein IBD Clinical Center, Icahn School of Medicine at Mount Sinai, New York, NY 10029, USA. -

Evolutionary Divergence and Functions of the Human Interleukin (IL) Gene Family Chad Brocker,1 David Thompson,2 Akiko Matsumoto,1 Daniel W
UPDATE ON GENE COMPLETIONS AND ANNOTATIONS Evolutionary divergence and functions of the human interleukin (IL) gene family Chad Brocker,1 David Thompson,2 Akiko Matsumoto,1 Daniel W. Nebert3* and Vasilis Vasiliou1 1Molecular Toxicology and Environmental Health Sciences Program, Department of Pharmaceutical Sciences, University of Colorado Denver, Aurora, CO 80045, USA 2Department of Clinical Pharmacy, University of Colorado Denver, Aurora, CO 80045, USA 3Department of Environmental Health and Center for Environmental Genetics (CEG), University of Cincinnati Medical Center, Cincinnati, OH 45267–0056, USA *Correspondence to: Tel: þ1 513 821 4664; Fax: þ1 513 558 0925; E-mail: [email protected]; [email protected] Date received (in revised form): 22nd September 2010 Abstract Cytokines play a very important role in nearly all aspects of inflammation and immunity. The term ‘interleukin’ (IL) has been used to describe a group of cytokines with complex immunomodulatory functions — including cell proliferation, maturation, migration and adhesion. These cytokines also play an important role in immune cell differentiation and activation. Determining the exact function of a particular cytokine is complicated by the influence of the producing cell type, the responding cell type and the phase of the immune response. ILs can also have pro- and anti-inflammatory effects, further complicating their characterisation. These molecules are under constant pressure to evolve due to continual competition between the host’s immune system and infecting organisms; as such, ILs have undergone significant evolution. This has resulted in little amino acid conservation between orthologous proteins, which further complicates the gene family organisation. Within the literature there are a number of overlapping nomenclature and classification systems derived from biological function, receptor-binding properties and originating cell type. -

Open Chen-Thesis Finalv5.Pdf
The Pennsylvania State University The Graduate School Department of Neural and Behavioral Sciences EPIGENETIC ANALYSIS OF IMMUNE ASSOCIATED SIGNALING MOLECULES DURING MOUSE RETINA DEVELOPMENT A Thesis in Anatomy by Chen Yang © 2013 Chen Yang Submitted in Partial Fulfillment of the Requirements for the Degree of Master of Science May 2013 The thesis of Chen Yang was reviewed and approved* by the following: Samuel Shao-Min Zhang Assistant Professor of Neural and Behavioral Sciences Thesis Advisor Colin J. Barnstable Department Head of Neural and Behavioral Sciences Professor of Neural and Behavioral Sciences Patricia J. McLaughlin Professor of Neural and Behavioral Sciences Director of Graduate Program in Anatomy *Signatures are on file in the Graduate School. ii ABSTRACT The retina is an immune-privileged organ. Many autoimmune diseases, such as AMD, glaucoma, and diabetic retinopathy, are caused by excessive inflammatory responses targeting self-tissue. The physiological functions of extracellular and intracellular signaling molecules of immune responses have been well characterized. The epigenetic aspects of these molecules in the retina, however, have not been well elucidated. In this study, we examined the expression of selected immune-related genes, and their transcriptional accessibility via epigenetic mapping, cluster analysis, and RT-PCR. Among these genes, interleukin receptor related genes and intracellular signaling molecules exhibit higher transcriptional accessibility. Epigenetic mapping of the toll-like receptor (TLR) family revealed that 3 out of 13 TLRs exhibit H3K4me2 accumulation during retina development, suggesting that TLR2, TLR3, and TLR9 are the only TLR members expressed in the retina. Most of the NF-κB signaling molecules exhibited transcriptional accessibility, implying their essential roles in inflammatory regulation during retina maturation. -

Xo GENE PANEL
xO GENE PANEL Targeted panel of 1714 genes | Tumor DNA Coverage: 500x | RNA reads: 50 million Onco-seq panel includes clinically relevant genes and a wide array of biologically relevant genes Genes A-C Genes D-F Genes G-I Genes J-L AATK ATAD2B BTG1 CDH7 CREM DACH1 EPHA1 FES G6PC3 HGF IL18RAP JADE1 LMO1 ABCA1 ATF1 BTG2 CDK1 CRHR1 DACH2 EPHA2 FEV G6PD HIF1A IL1R1 JAK1 LMO2 ABCB1 ATM BTG3 CDK10 CRK DAXX EPHA3 FGF1 GAB1 HIF1AN IL1R2 JAK2 LMO7 ABCB11 ATR BTK CDK11A CRKL DBH EPHA4 FGF10 GAB2 HIST1H1E IL1RAP JAK3 LMTK2 ABCB4 ATRX BTRC CDK11B CRLF2 DCC EPHA5 FGF11 GABPA HIST1H3B IL20RA JARID2 LMTK3 ABCC1 AURKA BUB1 CDK12 CRTC1 DCUN1D1 EPHA6 FGF12 GALNT12 HIST1H4E IL20RB JAZF1 LPHN2 ABCC2 AURKB BUB1B CDK13 CRTC2 DCUN1D2 EPHA7 FGF13 GATA1 HLA-A IL21R JMJD1C LPHN3 ABCG1 AURKC BUB3 CDK14 CRTC3 DDB2 EPHA8 FGF14 GATA2 HLA-B IL22RA1 JMJD4 LPP ABCG2 AXIN1 C11orf30 CDK15 CSF1 DDIT3 EPHB1 FGF16 GATA3 HLF IL22RA2 JMJD6 LRP1B ABI1 AXIN2 CACNA1C CDK16 CSF1R DDR1 EPHB2 FGF17 GATA5 HLTF IL23R JMJD7 LRP5 ABL1 AXL CACNA1S CDK17 CSF2RA DDR2 EPHB3 FGF18 GATA6 HMGA1 IL2RA JMJD8 LRP6 ABL2 B2M CACNB2 CDK18 CSF2RB DDX3X EPHB4 FGF19 GDNF HMGA2 IL2RB JUN LRRK2 ACE BABAM1 CADM2 CDK19 CSF3R DDX5 EPHB6 FGF2 GFI1 HMGCR IL2RG JUNB LSM1 ACSL6 BACH1 CALR CDK2 CSK DDX6 EPOR FGF20 GFI1B HNF1A IL3 JUND LTK ACTA2 BACH2 CAMTA1 CDK20 CSNK1D DEK ERBB2 FGF21 GFRA4 HNF1B IL3RA JUP LYL1 ACTC1 BAG4 CAPRIN2 CDK3 CSNK1E DHFR ERBB3 FGF22 GGCX HNRNPA3 IL4R KAT2A LYN ACVR1 BAI3 CARD10 CDK4 CTCF DHH ERBB4 FGF23 GHR HOXA10 IL5RA KAT2B LZTR1 ACVR1B BAP1 CARD11 CDK5 CTCFL DIAPH1 ERCC1 FGF3 GID4 -

Transcriptomic Landscape of Early Age Onset of Colorectal Cancer Identifies
www.nature.com/scientificreports OPEN Transcriptomic landscape of early age onset of colorectal cancer identifes novel genes and pathways in Indian CRC patients Manish Pratap Singh, Sandhya Rai, Nand K. Singh & Sameer Srivastava* Past decades of the current millennium have witnessed an unprecedented rise in Early age Onset of Colo Rectal Cancer (EOCRC) cases in India as well as across the globe. Unfortunately, EOCRCs are diagnosed at a more advanced stage of cancer. Moreover, the aetiology of EOCRC is not fully explored and still remains obscure. This study is aimed towards the identifcation of genes and pathways implicated in the EOCRC. In the present study, we performed high throughput RNA sequencing of colorectal tumor tissues for four EOCRC (median age 43.5 years) samples with adjacent mucosa and performed subsequent bioinformatics analysis to identify novel deregulated pathways and genes. Our integrated analysis identifes 17 hub genes (INSR, TNS1, IL1RAP, CD22, FCRLA, CXCL3, HGF, MS4A1, CD79B, CXCR2, IL1A, PTPN11, IRS1, IL1B, MET, TCL1A, and IL1R1). Pathway analysis of identifed genes revealed that they were involved in the MAPK signaling pathway, hematopoietic cell lineage, cytokine–cytokine receptor pathway and PI3K-Akt signaling pathway. Survival and stage plot analysis identifed four genes CXCL3, IL1B, MET and TNS1 genes (p = 0.015, 0.038, 0.049 and 0.011 respectively), signifcantly associated with overall survival. Further, diferential expression of TNS1 and MET were confrmed on the validation cohort of the 5 EOCRCs (median age < 50 years and sporadic origin). This is the frst approach to fnd early age onset biomarkers in Indian CRC patients. -

Immune Suppressive Landscape in the Human Esophageal Squamous Cell
ARTICLE https://doi.org/10.1038/s41467-020-20019-0 OPEN Immune suppressive landscape in the human esophageal squamous cell carcinoma microenvironment ✉ Yingxia Zheng 1,2,10 , Zheyi Chen1,10, Yichao Han 3,10, Li Han1,10, Xin Zou4, Bingqian Zhou1, Rui Hu5, Jie Hao4, Shihao Bai4, Haibo Xiao5, Wei Vivian Li6, Alex Bueker7, Yanhui Ma1, Guohua Xie1, Junyao Yang1, ✉ ✉ ✉ Shiyu Chen1, Hecheng Li 3 , Jian Cao 7,8 & Lisong Shen 1,9 1234567890():,; Cancer immunotherapy has revolutionized cancer treatment, and it relies heavily on the comprehensive understanding of the immune landscape of the tumor microenvironment (TME). Here, we obtain a detailed immune cell atlas of esophageal squamous cell carcinoma (ESCC) at single-cell resolution. Exhausted T and NK cells, regulatory T cells (Tregs), alternatively activated macrophages and tolerogenic dendritic cells are dominant in the TME. Transcriptional profiling coupled with T cell receptor (TCR) sequencing reveal lineage con- nections in T cell populations. CD8 T cells show continuous progression from pre-exhausted to exhausted T cells. While exhausted CD4, CD8 T and NK cells are major proliferative cell components in the TME, the crosstalk between macrophages and Tregs contributes to potential immunosuppression in the TME. Our results indicate several immunosuppressive mechanisms that may be simultaneously responsible for the failure of immuno-surveillance. Specific targeting of these immunosuppressive pathways may reactivate anti-tumor immune responses in ESCC. 1 Department of Laboratory Medicine, Xin Hua Hospital, Shanghai Jiao Tong University School of Medicine, Shanghai, China. 2 Institute of Biliary Tract Diseases Research, Shanghai Jiao Tong University School of Medicine, Shanghai, China. 3 Department of Thoracic Surgery, Ruijin Hospital, Shanghai Jiao Tong University School of Medicine, Shanghai, China. -

Supplemental Figure 1. No Lymphoid Cell Number Changes Are Observed in the Joint 2-And-4 Weeks Post ACLT
Supplemental Figure 1. No lymphoid cell number changes are observed in the joint 2-and-4 weeks post ACLT. Numbers of IFNy+ CD4+ T cells in the joint are similar to no surgery at 1 week (A), 2 weeks (B), and 4 weeks (C). (D) is an example dot plot from an ACLT joint 2 weeks post surgery demonstrating the low number of IL-4+ cells (30) for quantitation. This sample is representative of most joint samples for IL-4 levels. Lymphoid cell numbers in the joint are similar to no surgery at 2 weeks (E) and 4 weeks (F) post surgery. Supplemental Figure 2. (A) Immunofluorescence of human healthy and OA synovium. Only OA tissue exhibits IL-17 and IL-23 staining. Joint and inguinal lymph node post injury gene expression, flow cytometry, and immunohistochemistry characterization. (B) Nanostring pathway analysis of ILCs sorted from 2wk post ACLT vs no surgery joints. (C) Inguinal lymph node PCR from young mice with sham or ACLT 1, 2, and 4 wks post surgery (n=3). Stars represent significance compared to the no surgery group, pound signs indicate significance compared to opposite surgical group, and when p is indicated numerically, groups being compared are specified with a line. Groups are only statistically compared within each time point. (D) OARSI scores and representative images of sham joints 4wks post surgery. E. Whole joint PCR from young mice with sham or ACLT 1, 2, and 4 wks post surgery (n=3). Same statistical scheme as used in C. *p<0.05, **p <0.01, ***p<0.001 and ****p<0.001.