Adherence to and Invasion of Host Cells by Spotted Fever Group Rickettsia Species
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

(Batch Learning Self-Organizing Maps), to the Microbiome Analysis of Ticks
Title A novel approach, based on BLSOMs (Batch Learning Self-Organizing Maps), to the microbiome analysis of ticks Nakao, Ryo; Abe, Takashi; Nijhof, Ard M; Yamamoto, Seigo; Jongejan, Frans; Ikemura, Toshimichi; Sugimoto, Author(s) Chihiro The ISME Journal, 7(5), 1003-1015 Citation https://doi.org/10.1038/ismej.2012.171 Issue Date 2013-03 Doc URL http://hdl.handle.net/2115/53167 Type article (author version) File Information ISME_Nakao.pdf Instructions for use Hokkaido University Collection of Scholarly and Academic Papers : HUSCAP A novel approach, based on BLSOMs (Batch Learning Self-Organizing Maps), to the microbiome analysis of ticks Ryo Nakao1,a, Takashi Abe2,3,a, Ard M. Nijhof4, Seigo Yamamoto5, Frans Jongejan6,7, Toshimichi Ikemura2, Chihiro Sugimoto1 1Division of Collaboration and Education, Research Center for Zoonosis Control, Hokkaido University, Kita-20, Nishi-10, Kita-ku, Sapporo, Hokkaido 001-0020, Japan 2Nagahama Institute of Bio-Science and Technology, Nagahama, Shiga 526-0829, Japan 3Graduate School of Science & Technology, Niigata University, 8050, Igarashi 2-no-cho, Nishi- ku, Niigata 950-2181, Japan 4Institute for Parasitology and Tropical Veterinary Medicine, Freie Universität Berlin, Königsweg 67, 14163 Berlin, Germany 5Miyazaki Prefectural Institute for Public Health and Environment, 2-3-2 Gakuen Kibanadai Nishi, Miyazaki 889-2155, Japan 6Utrecht Centre for Tick-borne Diseases (UCTD), Department of Infectious Diseases and Immunology, Faculty of Veterinary Medicine, Utrecht University, Yalelaan 1, 3584 CL Utrecht, The Netherlands 7Department of Veterinary Tropical Diseases, Faculty of Veterinary Science, University of Pretoria, Private Bag X04, 0110 Onderstepoort, South Africa aThese authors contributed equally to this work. Keywords: BLSOMs/emerging diseases/metagenomics/microbiomes/symbionts/ticks Running title: Tick microbiomes revealed by BLSOMs Subject category: Microbe-microbe and microbe-host interactions Abstract Ticks transmit a variety of viral, bacterial and protozoal pathogens, which are often zoonotic. -

Babela Massiliensis, a Representative of a Widespread Bacterial
Babela massiliensis, a representative of a widespread bacterial phylum with unusual adaptations to parasitism in amoebae Isabelle Pagnier, Natalya Yutin, Olivier Croce, Kira S Makarova, Yuri I Wolf, Samia Benamar, Didier Raoult, Eugene V. Koonin, Bernard La Scola To cite this version: Isabelle Pagnier, Natalya Yutin, Olivier Croce, Kira S Makarova, Yuri I Wolf, et al.. Babela mas- siliensis, a representative of a widespread bacterial phylum with unusual adaptations to parasitism in amoebae. Biology Direct, BioMed Central, 2015, 10 (13), 10.1186/s13062-015-0043-z. hal-01217089 HAL Id: hal-01217089 https://hal-amu.archives-ouvertes.fr/hal-01217089 Submitted on 19 Oct 2015 HAL is a multi-disciplinary open access L’archive ouverte pluridisciplinaire HAL, est archive for the deposit and dissemination of sci- destinée au dépôt et à la diffusion de documents entific research documents, whether they are pub- scientifiques de niveau recherche, publiés ou non, lished or not. The documents may come from émanant des établissements d’enseignement et de teaching and research institutions in France or recherche français ou étrangers, des laboratoires abroad, or from public or private research centers. publics ou privés. Pagnier et al. Biology Direct (2015) 10:13 DOI 10.1186/s13062-015-0043-z RESEARCH Open Access Babela massiliensis, a representative of a widespread bacterial phylum with unusual adaptations to parasitism in amoebae Isabelle Pagnier1, Natalya Yutin2, Olivier Croce1, Kira S Makarova2, Yuri I Wolf2, Samia Benamar1, Didier Raoult1, Eugene V Koonin2 and Bernard La Scola1* Abstract Background: Only a small fraction of bacteria and archaea that are identifiable by metagenomics can be grown on standard media. -

Characterization of the Interaction Between R. Conorii and Human
Louisiana State University LSU Digital Commons LSU Doctoral Dissertations Graduate School 4-5-2018 Characterization of the Interaction Between R. Conorii and Human Host Vitronectin in Rickettsial Pathogenesis Abigail Inez Fish Louisiana State University and Agricultural and Mechanical College, [email protected] Follow this and additional works at: https://digitalcommons.lsu.edu/gradschool_dissertations Part of the Bacteria Commons, Bacteriology Commons, Biology Commons, Immunology of Infectious Disease Commons, and the Pathogenic Microbiology Commons Recommended Citation Fish, Abigail Inez, "Characterization of the Interaction Between R. Conorii and Human Host Vitronectin in Rickettsial Pathogenesis" (2018). LSU Doctoral Dissertations. 4566. https://digitalcommons.lsu.edu/gradschool_dissertations/4566 This Dissertation is brought to you for free and open access by the Graduate School at LSU Digital Commons. It has been accepted for inclusion in LSU Doctoral Dissertations by an authorized graduate school editor of LSU Digital Commons. For more information, please [email protected]. CHARACTERIZATION OF THE INTERACTION BETWEEN R. CONORII AND HUMAN HOST VITRONECTIN IN RICKETTSIAL PATHOGENESIS A Dissertation Submitted to the Graduate Faculty of the Louisiana State University and Agricultural and Mechanical College in partial fulfillment of the requirements for the degree of Doctor of Philosophy in The Interdepartmental Program in Biomedical and Veterinary Medical Sciences Through the Department of Pathobiological Sciences by Abigail Inez -

Transition of Serum Cytokine Concentration in Rickettsia Japonica Infection
Article Transition of Serum Cytokine Concentration in Rickettsia japonica Infection Makoto Kondo, Yoshiaki Matsushima, Kento Mizutani, Shohei Iida, Koji Habe and Keiichi Yamanaka * Department of Dermatology, Mie University Graduate School of medicine, 2-174 Edobashi, Tsu, Mie 514-8507, Japan; [email protected] (M.K.); [email protected] (Y.M.); [email protected] (K.M.); [email protected] (S.I.); [email protected] (K.H.) * Correspondence: [email protected]; Tel.: +81-59-231-5025; Fax: +81-59-231-5206 Received: 31 October 2020; Accepted: 8 December 2020; Published: 11 December 2020 Abstract: (1) Background. Rickettsia japonica (R. japonica) infection induces severe inflammation, and the disappearance of eosinophil in the acute stage is one of the phenomena. (2) Materials and Methods. In the current study, we measured the serum concentrations of Th1, Th2, and Th17 cytokines in the acute and recovery stages. (3) Results. In the acute phase, IL-6 and IFN-γ levels were elevated and we speculated that they played a role as a defense mechanism against R. japonica. The high concentration of IFN-γ suppressed the differentiation of eosinophil and induced apoptosis of eosinophil, leading to the disappearance of eosinophil. On day 7, IL-6 and IFN-γ concentrations were decreased, and Th2 cytokines such as IL-5 and IL-9 were slightly increased. On day 14, eosinophil count recovered to the normal level. The transition of serum cytokine concentration in R. japonica infection was presented. (4) Conclusions. -

Rickettsia Prowazekii Common Human Exposure Routes
APPENDIX 2 Rickettsia prowazekii Common Human Exposure Routes: Disease Agent: • Exposure to the feces of infected body lice. The lice are infected by a human blood meal. The rickettsiae • Rickettsia prowazekii reproduce in the louse gut epithelium. Infection occurs when louse feces are scratched into the skin, Disease Agent Characteristics: inoculated onto mucous membrane or inhaled. • As a bioweapon, the agent can be aerosolized, with • Rickettsiae are obligate intracellular Gram-negative intent of infection through inhalation. bacteria. • Sporadic cases occur after exposure to flying squir- • Order: Rickettsiales; Family: Rickettsiaceae rels, most likely as a result of exposure to squirrel flea • Size: 0.3 ¥ 1.0 mm intracellular bacteria that take up feces. The organism has also been identified in ticks Gram stain poorly feeding on livestock in Africa. • Nucleic acid: Rickettsial genomes are among the Likelihood of Secondary Transmission: smallest of bacteria at 1000-1600 kb. The R. prowazekii genome is 1100 kb. • No evidence of direct person-to-person transmission • Physicochemical properties: Susceptible to 1% • Under crowded conditions, where bathing and sodium hypochlorite, 70% ethanol, glutaraldehyde, washing clothes are difficult, and where lice are formaldehyde and quaternary ammonium disinfec- present, typhus can spread explosively. tants. Sensitive to moist heat (121°C for at least • Recent outbreaks have occurred in a number of areas 15 min) and dry heat (160-170°C for at least 1 hour). in the world under conditions of war and population The organism is stable in tick tissues or blood under displacement. ambient environmental conditions, surviving up to 1 year; sensitive to drying (feces of infected ticks At-Risk Populations: quickly lose their infectivity on drying). -
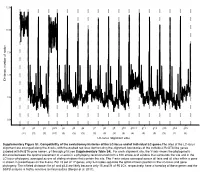
LC-Locus Alignment Sites Distance, Number of Nodes Supplementary
12.5 10.0 7.5 5.0 Distance, number of nodes 2.5 0.0 g1 g2 g3 g3.5 g4 g5 g6 g7 g8 g9 g10 g10.1 g11 g12 g13 g14 g15 (11) (3) (3) (10) (3) (3) (3) (3) (3) (3) (3) (4) (3) (3) (3) (2) (3) LC-locus alignment sites Supplementary Figure S1. Compatibility of the evolutionary histories of the LC-locus and of individual LC genes.The sites of the LC-locus alignment are arranged along the X-axis, with the dashed red lines demarcating the alignment boundaries of the individual RcGTA-like genes (labeled with RcGTA gene names, g1 through g15; see Supplementary Table S4). For each alignment site, the Y-axis shows the phylogenetic distance between the optimal placement of a taxon in a phylogeny reconstructed from a 100 amino-acid window that surrounds the site and in the LC-locus phylogeny, averaged across all sliding windows that contain the site. The Y-axis values averaged across all taxa and all sites within a gene is shown in parentheses on the X-axis. For 15 out of 17 genes, only 2-4 nodes separate the optimal taxon position in the LC-locus and gene phylogeny. The inflated distances for g1 and g3.5 are likely because only 15 and 21 of 95 LCs, respectively, have a homolog of these genes and the SSPB analysis is highly sensitive to missing data (Berger et al. 2011). a. Bacteria Unassigned Thermotogae Tenericutes Synergistetes Spirochaetes Proteobacteria ylum Planctomycetes h p Firmicutes Deferribacteres Cyanobacteria Chloroflexi Bacteroidetes Actinobacteria Acidobacteria 1(11,750) 2(1,750) 3(2,538) 4(168) 5(51) 6(54) 7(43) 8(32) 9(26) 10(40) 11(33) 12(198) 13(173) 14(101) 15(98) 16(43) 17(114) Number of rcc01682−rcc01698 homologs in a cluster b. -

CD Alert Monthly Newsletter of National Centre for Disease Control, Directorate General of Health Services, Government of India
CD Alert Monthly Newsletter of National Centre for Disease Control, Directorate General of Health Services, Government of India May - July 2009 Vol. 13 : No. 1 SCRUB TYPHUS & OTHER RICKETTSIOSES it lacks lipopolysaccharide and peptidoglycan RICKETTSIAL DISEASES and does not have an outer slime layer. It is These are the diseases caused by rickettsiae endowed with a major surface protein (56kDa) which are small, gram negative bacilli adapted and some minor surface protein (110, 80, 46, to obligate intracellular parasitism, and 43, 39, 35, 25 and 25kDa). There are transmitted by arthropod vectors. These considerable differences in virulence and organisms are primarily parasites of arthropods antigen composition among individual strains such as lice, fleas, ticks and mites, in which of O.tsutsugamushi. O.tsutsugamushi has they are found in the alimentary canal. In many serotypes (Karp, Gillian, Kato and vertebrates, including humans, they infect the Kawazaki). vascular endothelium and reticuloendothelial GLOBAL SCENARIO cells. Commonly known rickettsial disease is Scrub Typhus. Geographic distribution of the disease occurs within an area of about 13 million km2 including- The family Rickettsiaeceae currently comprises Afghanistan and Pakistan to the west; Russia of three genera – Rickettsia, Orientia and to the north; Korea and Japan to the northeast; Ehrlichia which appear to have descended Indonesia, Papua New Guinea, and northern from a common ancestor. Former members Australia to the south; and some smaller of the family, Coxiella burnetii, which causes islands in the western Pacific. It was Q fever and Rochalimaea quintana causing first observed in Japan where it was found to trench fever have been excluded because the be transmitted by mites. -

Health: Epidemiology Subchapter 41A
CHAPTER 41 – HEALTH: EPIDEMIOLOGY SUBCHAPTER 41A – COMMUNICABLE DISEASE CONTROL SECTION .0100 – REPORTING OF COMMUNICABLE DISEASES 10A NCAC 41A .0101 REPORTABLE DISEASES AND CONDITIONS (a) The following named diseases and conditions are declared to be dangerous to the public health and are hereby made reportable within the time period specified after the disease or condition is reasonably suspected to exist: (1) acquired immune deficiency syndrome (AIDS) - 24 hours; (2) anthrax - immediately; (3) botulism - immediately; (4) brucellosis - 7 days; (5) campylobacter infection - 24 hours; (6) chancroid - 24 hours; (7) chikungunya virus infection - 24 hours; (8) chlamydial infection (laboratory confirmed) - 7 days; (9) cholera - 24 hours; (10) Creutzfeldt-Jakob disease - 7 days; (11) cryptosporidiosis - 24 hours; (12) cyclosporiasis - 24 hours; (13) dengue - 7 days; (14) diphtheria - 24 hours; (15) Escherichia coli, shiga toxin-producing - 24 hours; (16) ehrlichiosis - 7 days; (17) encephalitis, arboviral - 7 days; (18) foodborne disease, including Clostridium perfringens, staphylococcal, Bacillus cereus, and other and unknown causes - 24 hours; (19) gonorrhea - 24 hours; (20) granuloma inguinale - 24 hours; (21) Haemophilus influenzae, invasive disease - 24 hours; (22) Hantavirus infection - 7 days; (23) Hemolytic-uremic syndrome – 24 hours; (24) Hemorrhagic fever virus infection - immediately; (25) hepatitis A - 24 hours; (26) hepatitis B - 24 hours; (27) hepatitis B carriage - 7 days; (28) hepatitis C, acute - 7 days; (29) human immunodeficiency -

Ohio Department of Health, Bureau of Infectious Diseases Disease Name Class A, Requires Immediate Phone Call to Local Health
Ohio Department of Health, Bureau of Infectious Diseases Reporting specifics for select diseases reportable by ELR Class A, requires immediate phone Susceptibilities specimen type Reportable test name (can change if Disease Name other specifics+ call to local health required* specifics~ state/federal case definition or department reporting requirements change) Culture independent diagnostic tests' (CIDT), like BioFire panel or BD MAX, E. histolytica Stain specimen = stool, bile results should be sent as E. histolytica DNA fluid, duodenal fluid, 260373001^DETECTED^SCT with E. histolytica Antigen Amebiasis (Entamoeba histolytica) No No tissue large intestine, disease/organism-specific DNA LOINC E. histolytica Antibody tissue small intestine codes OR a generic CIDT-LOINC code E. histolytica IgM with organism-specific DNA SNOMED E. histolytica IgG codes E. histolytica Total Antibody Ova and Parasite Anthrax Antibody Anthrax Antigen Anthrax EITB Acute Anthrax EITB Convalescent Anthrax Yes No Culture ELISA PCR Stain/microscopy Stain/spore ID Eastern Equine Encephalitis virus Antibody Eastern Equine Encephalitis virus IgG Antibody Eastern Equine Encephalitis virus IgM Arboviral neuroinvasive and non- Eastern Equine Encephalitis virus RNA neuroinvasive disease: Eastern equine California serogroup virus Antibody encephalitis virus disease; LaCrosse Equivocal results are accepted for all California serogroup virus IgG Antibody virus disease (other California arborviral diseases; California serogroup virus IgM Antibody specimen = blood, serum, serogroup -

Mechanisms of Rickettsia Parkeri Invasion of Host Cells and Early Actin-Based Motility
Mechanisms of Rickettsia parkeri invasion of host cells and early actin-based motility By Shawna Reed A dissertation submitted in partial satisfaction of the requirements for the degree of Doctor of Philosophy in Microbiology in the Graduate Division of the University of California, Berkeley Committee in charge: Professor Matthew D. Welch, Chair Professor David G. Drubin Professor Daniel Portnoy Professor Kathleen R. Ryan Fall 2012 Mechanisms of Rickettsia parkeri invasion of host cells and early actin-based motility © 2012 By Shawna Reed ABSTRACT Mechanisms of Rickettsia parkeri invasion of host cells and early actin-based motility by Shawna Reed Doctor of Philosophy in Microbiology University of California, Berkeley Professor Matthew D. Welch, Chair Rickettsiae are obligate intracellular pathogens that are transmitted to humans by arthropod vectors and cause diseases such as spotted fever and typhus. Spotted fever group (SFG) Rickettsia hijack the host actin cytoskeleton to invade, move within, and spread between eukaryotic host cells during their obligate intracellular life cycle. Rickettsia express two bacterial proteins that can activate actin polymerization: RickA activates the host actin-nucleating Arp2/3 complex while Sca2 directly nucleates actin filaments. In this thesis, I aimed to resolve which host proteins were required for invasion and intracellular motility, and to determine how the bacterial proteins RickA and Sca2 contribute to these processes. Although rickettsiae require the host cell actin cytoskeleton for invasion, the cytoskeletal proteins that mediate this process have not been completely described. To identify the host factors important during cell invasion by Rickettsia parkeri, a member of the SFG, I performed an RNAi screen targeting 105 proteins in Drosophila melanogaster S2R+ cells. -

Gene Gain and Loss Events in Rickettsia and Orientia Species Kalliopi Georgiades1,2, Vicky Merhej1, Khalid El Karkouri1, Didier Raoult1, Pierre Pontarotti2*
Georgiades et al. Biology Direct 2011, 6:6 http://www.biology-direct.com/content/6/1/6 RESEARCH Open Access Gene gain and loss events in Rickettsia and Orientia species Kalliopi Georgiades1,2, Vicky Merhej1, Khalid El Karkouri1, Didier Raoult1, Pierre Pontarotti2* Abstract Background: Genome degradation is an ongoing process in all members of the Rickettsiales order, which makes these bacterial species an excellent model for studying reductive evolution through interspecies variation in genome size and gene content. In this study, we evaluated the degree to which gene loss shaped the content of some Rickettsiales genomes. We shed light on the role played by horizontal gene transfers in the genome evolution of Rickettsiales. Results: Our phylogenomic tree, based on whole-genome content, presented a topology distinct from that of the whole core gene concatenated phylogenetic tree, suggesting that the gene repertoires involved have different evolutionary histories. Indeed, we present evidence for 3 possible horizontal gene transfer events from various organisms to Orientia and 6 to Rickettsia spp., while we also identified 3 possible horizontal gene transfer events from Rickettsia and Orientia to other bacteria. We found 17 putative genes in Rickettsia spp. that are probably the result of de novo gene creation; 2 of these genes appear to be functional. On the basis of these results, we were able to reconstruct the gene repertoires of “proto-Rickettsiales” and “proto-Rickettsiaceae”, which correspond to the ancestors of Rickettsiales and Rickettsiaceae, respectively. Finally, we found that 2,135 genes were lost during the evolution of the Rickettsiaceae to an intracellular lifestyle. Conclusions: Our phylogenetic analysis allowed us to track the gene gain and loss events occurring in bacterial genomes during their evolution from a free-living to an intracellular lifestyle. -

Scrub Typhus and Molecular Characterization of Orientia Tsutsugamushi from Central Nepal
pathogens Article Scrub Typhus and Molecular Characterization of Orientia tsutsugamushi from Central Nepal Rajendra Gautam 1, Keshab Parajuli 1, Mythili Tadepalli 2, Stephen Graves 2, John Stenos 2,* and Jeevan Bahadur Sherchand 1 1 Department of Microbiology, Maharajgunj Medical Campus, Institute of Medicine, Kathmandu 44600, Nepal; [email protected] (R.G.); [email protected] (K.P.); [email protected] (J.B.S.) 2 Australian Rickettsial Reference Laboratory, Geelong, VIC 3220, Australia; [email protected] (M.T.); [email protected] (S.G.) * Correspondence: [email protected]; Tel.: +61-342151357 Abstract: Scrub typhus is a vector-borne, acute febrile illness caused by Orientia tsutsugamushi. Scrub typhus continues to be an important but neglected tropical disease in Nepal. Information on this pathogen in Nepal is limited to serological surveys with little information available on molecular methods to detect O. tsutsugamushi. Limited information exists on the genetic diversity of this pathogen. A total of 282 blood samples were obtained from patients with suspected scrub typhus from central Nepal and 84 (30%) were positive for O. tsutsugamushi by 16S rRNA qPCR. Positive samples were further subjected to 56 kDa and 47 kDa molecular typing and molecularly compared to other O. tsutsugamushi strains. Phylogenetic analysis revealed that Nepalese O. tsutsugamushi strains largely cluster together and cluster away from other O. tsutsugamushi strains from Asia and elsewhere. One exception was the sample of Nepal_1, with its partial 56 kDa sequence clustering Citation: Gautam, R.; Parajuli, K.; more closely with non-Nepalese O. tsutsugamushi 56 kDa sequences, potentially indicating that Tadepalli, M.; Graves, S.; Stenos, J.; homologous recombination may influence the genetic diversity of strains in this region.