Interactome Mapping of Eif3a in a Colon Cancer and an Immortalized Embryonic Cell Line Using Proximity-Dependent Biotin Identification
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Composition of Herpesvirus Ribonucleoprotein Complexes †
Abstract Composition of Herpesvirus Ribonucleoprotein Complexes † Eric S. Pringle 1,2,* and Craig McCormick 1,2 1 Department of Microbiology and Immunology, Dalhousie University, Halifax, NS B3H 4R2, Canada; [email protected] 2 Beatrice Hunter Cancer Research Institute, Halifax, NS B3H 4R2, Canada * Correspondence: [email protected] † Presented at Viruses 2020—Novel Concepts in Virology, Barcelona, Spain, 5–7 February 2020. Published: 4 July 2020 Abstract: Herpesvirus genomes are decoded by host RNA polymerase enzymes, generating messenger ribonucleotides (mRNA) that are post-transcriptionally modified and exported to the cytoplasm through the combined work of host and viral factors. These viral mRNA bear 5′-m7GTP caps and poly(A) tails that should permit the assembly of canonical host eIF4F cap-binding complexes to initiate protein synthesis. However, the precise mechanisms of translation initiation remain to be investigated for Kaposi’s sarcoma-associated herpesvirus (KSHV) and other herpesviruses. During KSHV lytic replication in lymphoid cells, the activation of caspases leads to the cleavage of eIF4G and depletion of eIF4F. Translating mRNPs depleted of eIF4F retain viral mRNA, suggesting that non-eIF4F translation initiation is sufficient to support viral protein synthesis. To identify proteins required to support viral protein synthesis, we isolated and characterized actively translating messenger ribonucleoprotein (mRNP) complexes by ultracentrifugation and sucrose-gradient fractionation followed by quantitative mass spectrometry. The abundance of host translation initiation factors available to initiate viral protein synthesis were comparable between cells undergoing KSHV lytic or latent replication. The translation initiation factors eIF4E2, NCBP1, eIF4G2, and eIF3d were detected in association with actively translating mRNP complexes during KSHV lytic replication, but their depletion by RNA silencing did not affect virion production. -

Structural Characterization of the Human Eukaryotic Initiation Factor 3 Protein Complex by Mass Spectrometry*□S
Supplemental Material can be found at: http://www.mcponline.org/cgi/content/full/M600399-MCP200 /DC1 Research Structural Characterization of the Human Eukaryotic Initiation Factor 3 Protein Complex by Mass Spectrometry*□S Eugen Damoc‡, Christopher S. Fraser§, Min Zhou¶, Hortense Videler¶, Greg L. Mayeurʈ, John W. B. Hersheyʈ, Jennifer A. Doudna§, Carol V. Robinson¶**, and Julie A. Leary‡ ‡‡ Protein synthesis in mammalian cells requires initiation The initiation phase of eukaryotic protein synthesis involves factor eIF3, an ϳ800-kDa protein complex that plays a formation of an 80 S ribosomal complex containing the initi- Downloaded from central role in binding of initiator methionyl-tRNA and ator methionyl-tRNAi bound to the initiation codon in the mRNA to the 40 S ribosomal subunit to form the 48 S mRNA. This is a multistep process promoted by proteins initiation complex. The eIF3 complex also prevents pre- called eukaryotic initiation factors (eIFs).1 Currently at least 12 mature association of the 40 and 60 S ribosomal subunits eIFs, composed of at least 29 distinct subunits, have been and interacts with other initiation factors involved in start identified (1). Mammalian eIF3, the largest initiation factor, is a codon selection. The molecular mechanisms by which multisubunit complex with an apparent molecular mass of www.mcponline.org eIF3 exerts these functions are poorly understood. Since ϳ800 kDa. This protein complex plays an essential role in its initial characterization in the 1970s, the exact size, translation by binding directly to the 40 S ribosomal subunit composition, and post-translational modifications of and promoting formation of the 43 S preinitiation complex ⅐ ⅐ mammalian eIF3 have not been rigorously determined. -

CHARACTERIZING the INTERACTION BETWEEN PDCD4 and Eif3 with RESPECT to TRANSLATION REGULATION
CHARACTERIZING THE INTERACTION BETWEEN PDCD4 AND eIF3 WITH RESPECT TO TRANSLATION REGULATION DIVYA SHARMA KHANDIGA Master of Science, Bangalore University, India 2009 A Thesis/Project Submitted to the School of Graduate Studies of the University of Lethbridge in Partial Fulfillment of the Requirements for the Degree MASTER OF SCIENCE Department of Chemistry and Biochemistry University of Lethbridge LETHBRIDGE, ALBERTA, CANADA © Divya Sharma Khandiga, 2017 CHARACTERIZING THE INTERACTION BETWEEN PDCD4 AND eIF3 WITH RESPECT TO TRANSLATION REGULATION DIVYA SHARMA KHANDIGA Date of Defense: December 12, 2017 Dr. N. Thakor Assistant Professor Ph.D. Thesis Supervisor Dr. M. Roussel Professor Ph.D. Thesis Co-supervisor Dr. U. Kothe Associate Professor Ph.D. Thesis Examination Committee Member Dr. R. Golsteyn Associate Professor Ph.D. Thesis Examination Committee Member Dr. R. Fahlman Professor Ph.D. External Examiner University of Alberta Edmonton, Alberta Dr. M. Gerken Professor Ph.D. Chair, Thesis Examination Committee Dedication To my beloved family and friends, My inspiration, my parents Subraya Sharma and Kamala Sharma My dearly loved husband Samarth, sister Dr. Lakshmi and brother-in-law Dr. Pradeep My cute little niece Mithali and nephew Aathreya My adorable brother Dr. Ganesh, sister Dr. Sharadha, Silly Vidya and little angels My loving cousins and in-laws I am grateful to have them in my life, it is their well wishes, teachings, support and love that have enabled me to achieve success and happiness in life. iii Abstract Programmed cell death protein 4 (PDCD4) inhibits IRES-mediated translation of anti- apoptotic proteins such as XIAP. PDCD4 was shown to directly interact with the XIAP IRES element and inhibit translation initiation. -

A Versatile Eif3d in Translational Control of Stress Adaptation
ll Spotlight A Versatile eIF3d in Translational Control of Stress Adaptation Longfei Jia1 and Shu-Bing Qian1,* 1Division of Nutritional Sciences, Cornell University, Ithaca, NY, USA *Correspondence: [email protected] https://doi.org/10.1016/j.molcel.2020.12.016 Lamper et al. (2020) reported that eIF3d-mediated cap-dependent translation is subject to regulation by phosphorylation during chronic glucose deprivation, providing a mechanism underlying selective translation of stress genes essential for cell survival. A remarkable feature of all living organ- tion, however, overall translation is because eIF3d contains a natural HIV-1 isms is the ability to sense fluctuations reduced only by 60% (An et al., protease cleavage site. Remarkably, of environmental cues and respond to 2020). This does not necessarily mean glucose starvation led to an increased adverse conditions by adjusting cellular that the remaining 40% of translation cap-binding activity of eIF3d by 10- activities. Protein synthesis consumes solely relies on cap-independent mech- fold. Since phosphorylation is a common a lion’s share of cellular energy. Not sur- anisms. Notably, most mRNAs are still mechanism in stress signaling pathways, prisingly, many stress conditions sup- capped when the canonical eIF4F is in- the authors confirmed nutrient-depen- press global protein synthesis as part hibited. Although cap-dependent trans- dent phosphorylation of eIF3d. Using of the stress response. However, trans- lation is widely believed to be driven biochemical and genetic approaches, lation of certain mRNAs needs to be through eIF4F, alternative mechanisms they not only mapped the phosphoryla- maintained or even upregulated to sus- likely exist to mediate cap-dependent tion sites but also identified CK2 as tain vital functions. -

Structural Studies of the Eif4e–Vpg Complex Reveal a Direct Competition for Capped RNA: Implications for Translation
Structural studies of the eIF4E–VPg complex reveal a direct competition for capped RNA: Implications for translation Luciana Coutinho de Oliveiraa,1,2, Laurent Volpona,1, Amanda K. Rahardjoa, Michael J. Osbornea, Biljana Culjkovic-Kraljacica, Christian Trahanb, Marlene Oeffingerb,c,d, Benjamin H. Kwoka, and Katherine L. B. Bordena,3 aInstitute of Research in Immunology and Cancer, Department of Pathology and Cell Biology, Université de Montréal, Pavilion Marcelle-Coutu, Chemin Polytechnique, Montréal, QC H3T 1J4, Canada; bDepartment for Systems Biology, Institut de Recherches Cliniques de Montréal, Montréal, QC H2W 1R7, Canada; cDépartement de Biochimie et Médecine Moléculaire, Université de Montréal, Montréal, QC H3T 1J4, Canada; and dDivision of Experimental Medicine, McGill University, Montréal, QC H3A 1A3, Canada Edited by Lynne E. Maquat, University of Rochester School of Medicine and Dentistry, Rochester, NY, and approved October 16, 2019 (received for review March 19, 2019) Viruses have transformed our understanding of mammalian RNA of eIF4E can be targeted in patients to provide clinical benefit, processing, including facilitating the discovery of the methyl-7- highlighting its critical importance. guanosine (m7G)caponthe5′ end of RNAs. The m7Gcapisrequired Viruses have paved the way for our understanding of many for RNAs to bind the eukaryotic translation initiation factor eIF4E aspects of host-cell RNA processing, including m7G capping. and associate with the translation machinery across plant and ani- Indeed, studies into cytoplasmic polyhedrosis virus (CPV) infec- mal kingdoms. The potyvirus-derived viral genome-linked protein tion in silkworm and vaccinia virus (VV) in mammalian cells were (VPg) is covalently bound to the 5′ end of viral genomic RNA (gRNA) critical for the elucidation of the m7G cap structure over 40 y ago and associates with host eIF4E for successful infection. -

Genes with 5' Terminal Oligopyrimidine Tracts Preferentially Escape Global Suppression of Translation by the SARS-Cov-2 NSP1 Protein
Downloaded from rnajournal.cshlp.org on September 28, 2021 - Published by Cold Spring Harbor Laboratory Press Genes with 5′ terminal oligopyrimidine tracts preferentially escape global suppression of translation by the SARS-CoV-2 Nsp1 protein Shilpa Raoa, Ian Hoskinsa, Tori Tonna, P. Daniela Garciaa, Hakan Ozadama, Elif Sarinay Cenika, Can Cenika,1 a Department of Molecular Biosciences, University of Texas at Austin, Austin, TX 78712, USA 1Corresponding author: [email protected] Key words: SARS-CoV-2, Nsp1, MeTAFlow, translation, ribosome profiling, RNA-Seq, 5′ TOP, Ribo-Seq, gene expression 1 Downloaded from rnajournal.cshlp.org on September 28, 2021 - Published by Cold Spring Harbor Laboratory Press Abstract Viruses rely on the host translation machinery to synthesize their own proteins. Consequently, they have evolved varied mechanisms to co-opt host translation for their survival. SARS-CoV-2 relies on a non-structural protein, Nsp1, for shutting down host translation. However, it is currently unknown how viral proteins and host factors critical for viral replication can escape a global shutdown of host translation. Here, using a novel FACS-based assay called MeTAFlow, we report a dose-dependent reduction in both nascent protein synthesis and mRNA abundance in cells expressing Nsp1. We perform RNA-Seq and matched ribosome profiling experiments to identify gene-specific changes both at the mRNA expression and translation level. We discover that a functionally-coherent subset of human genes are preferentially translated in the context of Nsp1 expression. These genes include the translation machinery components, RNA binding proteins, and others important for viral pathogenicity. Importantly, we uncovered a remarkable enrichment of 5′ terminal oligo-pyrimidine (TOP) tracts among preferentially translated genes. -

The Yeast Eif3 Subunits TIF32/A, NIP1/C, and Eif5 Make Critical Connections with the 40Sribosome in Vivo
Downloaded from genesdev.cshlp.org on October 6, 2021 - Published by Cold Spring Harbor Laboratory Press The yeast eIF3 subunits TIF32/a, NIP1/c, and eIF5 make critical connections with the 40Sribosome in vivo Leoš Valášek, Amy A. Mathew, Byung-Sik Shin, Klaus H. Nielsen, Béla Szamecz, and Alan G. Hinnebusch1 Laboratory of Gene Regulation and Development, National Institute of Child Health and Human Development, Bethesda, Maryland 20892, USA Initiation factor 3 (eIF3) forms a multifactor complex (MFC) with eIF1, eIF2, and eIF5 that stimulates Met Met-tRNAi binding to 40Sribosomes and promotes scanning or AUG recognition. We have previously characterized MFC subcomplexes produced in vivo from affinity-tagged eIF3 subunits lacking discrete binding domains for other MFC components. Here we asked whether these subcomplexes can bind to 40Sribosomes in vivo. We found that the N- and C-terminal domains of NIP1/eIF3c, the N- and C-terminal domains of TIF32/eIF3a, and eIF5 have critical functions in 40Sbinding, with eIF5 an d the TIF32-CTD performing redundant functions. The TIF32-CTD interacted in vitro with helices 16–18 of domain I in 18SrRNA, and the TIF32-NTD and NIP1 interacted with 40Sprotein RPS0A.These results sugge st that eIF3 binds to the solvent side of the 40Ssubunit in a way that provides access to the interface side fo r the two eIF3 segments Met (NIP1-NTD and TIF32-CTD) that interact with eIF1, eIF5, and the eIF2/GTP/Met-tRNAi ternary complex. [Keywords: Eukaryotic translation initiation factor (eIF); multifactor complex (MFC); translational control; protein synthesis; 40S ribosome binding; TIF32/NIP1] Supplemental material is available at http://www.genesdev.org. -

Structure of Ah
Eliseev, B., Yeramala, L., Leitner, A., Karuppasamy, M., Raimondeau, E., Huard, K., ... Schaffitzel, C. (2018). Structure of a human cap-dependent 48S translation pre-initiation complex. Nucleic Acids Research, 46(5), 2678- 2689. [gky054]. https://doi.org/10.1093/nar/gky054 Publisher's PDF, also known as Version of record License (if available): CC BY Link to published version (if available): 10.1093/nar/gky054 Link to publication record in Explore Bristol Research PDF-document This is the final published version of the article (version of record). It first appeared online via Oxford University Press at https://academic.oup.com/nar/advance-article/doi/10.1093/nar/gky054/4833217 . Please refer to any applicable terms of use of the publisher. University of Bristol - Explore Bristol Research General rights This document is made available in accordance with publisher policies. Please cite only the published version using the reference above. Full terms of use are available: http://www.bristol.ac.uk/pure/about/ebr-terms 2678–2689 Nucleic Acids Research, 2018, Vol. 46, No. 5 Published online 1 February 2018 doi: 10.1093/nar/gky054 Structure of a human cap-dependent 48S translation pre-initiation complex Boris Eliseev1, Lahari Yeramala1, Alexander Leitner2, Manikandan Karuppasamy1, Etienne Raimondeau1, Karine Huard1, Elena Alkalaeva3, Ruedi Aebersold2,4 and Christiane Schaffitzel1,5,* 1European Molecular Biology Laboratory, Grenoble Outstation, 71 Avenue des Martyrs, 38042 Grenoble, France, 2ETH Zurich,¨ Institute of Molecular Systems Biology, -

GPRC5A Suppresses Protein Synthesis at the Endoplasmic Reticulum to Prevent Radiation-Induced Lung Tumorigenesis
ARTICLE Received 8 Mar 2016 | Accepted 28 Apr 2016 | Published 8 Jun 2016 DOI: 10.1038/ncomms11795 OPEN GPRC5A suppresses protein synthesis at the endoplasmic reticulum to prevent radiation-induced lung tumorigenesis Jian Wang1, Alton B. Farris2, Kaiming Xu1, Ping Wang1, Xiangming Zhang1, Duc M. Duong3, Hong Yi4, Hui-Kuo Shu1, Shi-Yong Sun5 & Ya Wang1 GPRC5A functions as a lung tumour suppressor to prevent spontaneous and environmentally induced lung carcinogenesis; however, the underlying mechanism remains unclear. Here we reveal that GPRC5A at the endoplasmic reticulum (ER) membrane suppresses synthesis of the secreted or membrane-bound proteins including a number of oncogenes, the most important one being Egfr. The ER-located GPRC5A disturbs the assembly of the eIF4F-mediated translation initiation complex on the mRNA cap through directly binding to the eIF4F complex with its two middle extracellular loops. Particularly, suppression of EGFR by GPRC5A contributes significantly to preventing ionizing radiation (IR)-induced lung tumorigenesis. Thus, GPRC5A deletion enhances IR-promoted EGFR expression through an increased translation rate, thereby significantly increasing lung tumour incidence in Gprc5a À / À mice. Our findings indicate that under-expressed GPRC5A during lung tumor- igenesis enhances any transcriptional stimulation through an active translational status, which can be used to control oncogene expression and potentially the resulting related disease. 1 Department of Radiation Oncology, Emory University School of Medicine and the Winship Cancer Institute, Emory University, Atlanta, Georgia 30322, USA. 2 Department of Pathology, Emory University School of Medicine and the Winship Cancer Institute, Emory University, Atlanta, Georgia 30322, USA. 3 Emory Integrated Proteomics Core and Biochemistry Department, Atlanta, Georgia 30322, USA. -
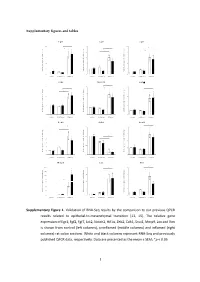
Type of the Paper (Article
Supplementary figures and tables E g r 1 F g f2 F g f7 1 0 * 5 1 0 * * e e e * g g g * n n n * a a a 8 4 * 8 h h h * c c c d d d * l l l o o o * f f f * n n n o o o 6 3 6 i i i s s s s s s e e e r r r p p p x x x e e e 4 2 4 e e e n n n e e e g g g e e e v v v i i i t t t 2 1 2 a a a l l l e e e R R R 0 0 0 c o n tro l u n in fla m e d in fla m e d c o n tro l u n in fla m e d in fla m e d c o n tro l u n in fla m e d in fla m e d J a k 2 N o tc h 2 H if1 * 3 4 6 * * * e e e g g g n n n a a * * a * h h * h c c c 3 * d d * d l l l * o o o f f 2 f 4 n n n o o o i i i s s s s s s e e e r r 2 r p p p x x x e e e e e e n n n e e 1 e 2 g g g e e 1 e v v v i i i t t t a a a l l l e e e R R R 0 0 0 c o n tro l u n in fla m e d in fla m e d c o n tro l u n in fla m e d in fla m e d c o n tro l u n in fla m e d in fla m e d Z e b 2 C d h 1 S n a i1 * * 7 1 .5 4 * * e e e g g g 6 n n n * a a a * h h h c c c 3 * d d d l l l 5 o o o f f f 1 .0 * n n n * o o o i i i 4 * s s s s s s e e e r r r 2 p p p x x x 3 e e e e e e n n n e e e 0 .5 g g g 2 e e e 1 v v v i i i t t t a a a * l l l e e e 1 * R R R 0 0 .0 0 c o n tro l u n in fla m e d in fla m e d c o n tro l u n in fla m e d in fla m e d c o n tro l u n in fla m e d in fla m e d M m p 9 L o x V im 2 0 0 2 0 8 * * * e e e * g g g 1 5 0 * n n n * a a a * h h h * c c c 1 5 * 6 d d d l l l 1 0 0 o o o f f f n n n o o o i i i 5 0 s s s s s s * e e e r r r 1 0 4 3 0 p p p * x x x e e e * e e e n n n e e e 2 0 g g g e e e 5 2 v v v i i i t t t a a a l l l 1 0 e e e R R R 0 0 0 c o n tro l u n in fla m e d in fla m e d c o n tro l u n in fla m e d in fla m e d c o n tro l u n in fla m e d in fla m e d Supplementary Figure 1. -

Systematically Profiling the Expression of Eif3 Subunits in Glioma Reveals
Chai et al. Cancer Cell Int (2019) 19:155 https://doi.org/10.1186/s12935-019-0867-1 Cancer Cell International PRIMARY RESEARCH Open Access Systematically profling the expression of eIF3 subunits in glioma reveals the expression of eIF3i has prognostic value in IDH-mutant lower grade glioma Rui‑Chao Chai1,4,6†, Ning Wang2†, Yu‑Zhou Chang3, Ke‑Nan Zhang1,6, Jing‑Jun Li1,6, Jun‑Jie Niu5, Fan Wu1,6*, Yu‑Qing Liu1,6* and Yong‑Zhi Wang1,3,4,6* Abstract Background: Abnormal expression of the eukaryotic initiation factor 3 (eIF3) subunits plays critical roles in tumo‑ rigenesis and progression, and also has potential prognostic value in cancers. However, the expression and clinical implications of eIF3 subunits in glioma remain unknown. Methods: Expression data of eIF3 for patients with gliomas were obtained from the Chinese Glioma Genome Atlas (CGGA) (n 272) and The Cancer Genome Atlas (TCGA) (n 595). Cox regression, the receiver operating characteristic (ROC) curves= and Kaplan–Meier analysis were used to study= the prognostic value. Gene oncology (GO) and gene set enrichment analysis (GSEA) were utilized for functional prediction. Results: In both the CGGA and TCGA datasets, the expression levels of eIF3d, eIF3e, eIF3f, eIF3h and eIF3l highly were associated with the IDH mutant status of gliomas. The expression of eIF3b, eIF3i, eIF3k and eIF3m was increased with the tumor grade, and was associated with poorer overall survival [All Hazard ratio (HR) > 1 and P < 0.05]. By contrast, the expression of eIF3a and eIF3l was decreased in higher grade gliomas and was associated with better overall sur‑ vival (Both HR < 1 and P < 0.05). -

The J-Subunit of Human Translation Initiation Factor Eif3 Is Required for the Stable Binding of Eif3 and Its Subcomplexes to 40 S Ribosomal Subunits in Vitro*
THE JOURNAL OF BIOLOGICAL CHEMISTRY Vol. 279, No. 10, Issue of March 5, pp. 8946–8956, 2004 © 2004 by The American Society for Biochemistry and Molecular Biology, Inc. Printed in U.S.A. The j-Subunit of Human Translation Initiation Factor eIF3 Is Required for the Stable Binding of eIF3 and Its Subcomplexes to 40 S Ribosomal Subunits in Vitro* Received for publication, November 21, 2003, and in revised form, December 18, 2003 Published, JBC Papers in Press, December 19, 2003, DOI 10.1074/jbc.M312745200 Christopher S. Fraser‡, Jennifer Y. Lee, Greg L. Mayeur, Martin Bushell§, Jennifer A. Doudna¶, and John W. B. Hersheyʈ From the Department of Biological Chemistry, School of Medicine, University of California, Davis, California 95616 Eukaryotic initiation factor 3 (eIF3) is a 12-subunit protein subunits, named in order of decreasing molecular protein complex that plays a central role in binding of weight as recommended (4): eIF3a, eIF3b, eIF3c, eIF3d, eIF3l, initiator methionyl-tRNA and mRNA to the 40 S riboso- eIF3e, eIF3f, eIF3g, eIF3h, eIF3i, eIF3j, and eIF3k (5, 6). Spe- mal subunit to form the 40 S initiation complex. The cific functions for mammalian eIF3 have been identified by a molecular mechanisms by which eIF3 exerts these func- variety of in vitro experiments. It binds directly to 40 S ribo- tions are poorly understood. To learn more about the somal subunits in the absence of other initiation components structure and function of eIF3 we have expressed and (1), and affects the association/dissociation of ribosomes (7–10). purified individual human eIF3 subunits or complexes It promotes the binding of Met-tRNA and mRNA to the 40 S of eIF3 subunits using baculovirus-infected Sf9 cells.