A Versatile Eif3d in Translational Control of Stress Adaptation
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Composition of Herpesvirus Ribonucleoprotein Complexes †
Abstract Composition of Herpesvirus Ribonucleoprotein Complexes † Eric S. Pringle 1,2,* and Craig McCormick 1,2 1 Department of Microbiology and Immunology, Dalhousie University, Halifax, NS B3H 4R2, Canada; [email protected] 2 Beatrice Hunter Cancer Research Institute, Halifax, NS B3H 4R2, Canada * Correspondence: [email protected] † Presented at Viruses 2020—Novel Concepts in Virology, Barcelona, Spain, 5–7 February 2020. Published: 4 July 2020 Abstract: Herpesvirus genomes are decoded by host RNA polymerase enzymes, generating messenger ribonucleotides (mRNA) that are post-transcriptionally modified and exported to the cytoplasm through the combined work of host and viral factors. These viral mRNA bear 5′-m7GTP caps and poly(A) tails that should permit the assembly of canonical host eIF4F cap-binding complexes to initiate protein synthesis. However, the precise mechanisms of translation initiation remain to be investigated for Kaposi’s sarcoma-associated herpesvirus (KSHV) and other herpesviruses. During KSHV lytic replication in lymphoid cells, the activation of caspases leads to the cleavage of eIF4G and depletion of eIF4F. Translating mRNPs depleted of eIF4F retain viral mRNA, suggesting that non-eIF4F translation initiation is sufficient to support viral protein synthesis. To identify proteins required to support viral protein synthesis, we isolated and characterized actively translating messenger ribonucleoprotein (mRNP) complexes by ultracentrifugation and sucrose-gradient fractionation followed by quantitative mass spectrometry. The abundance of host translation initiation factors available to initiate viral protein synthesis were comparable between cells undergoing KSHV lytic or latent replication. The translation initiation factors eIF4E2, NCBP1, eIF4G2, and eIF3d were detected in association with actively translating mRNP complexes during KSHV lytic replication, but their depletion by RNA silencing did not affect virion production. -

Structural Characterization of the Human Eukaryotic Initiation Factor 3 Protein Complex by Mass Spectrometry*□S
Supplemental Material can be found at: http://www.mcponline.org/cgi/content/full/M600399-MCP200 /DC1 Research Structural Characterization of the Human Eukaryotic Initiation Factor 3 Protein Complex by Mass Spectrometry*□S Eugen Damoc‡, Christopher S. Fraser§, Min Zhou¶, Hortense Videler¶, Greg L. Mayeurʈ, John W. B. Hersheyʈ, Jennifer A. Doudna§, Carol V. Robinson¶**, and Julie A. Leary‡ ‡‡ Protein synthesis in mammalian cells requires initiation The initiation phase of eukaryotic protein synthesis involves factor eIF3, an ϳ800-kDa protein complex that plays a formation of an 80 S ribosomal complex containing the initi- Downloaded from central role in binding of initiator methionyl-tRNA and ator methionyl-tRNAi bound to the initiation codon in the mRNA to the 40 S ribosomal subunit to form the 48 S mRNA. This is a multistep process promoted by proteins initiation complex. The eIF3 complex also prevents pre- called eukaryotic initiation factors (eIFs).1 Currently at least 12 mature association of the 40 and 60 S ribosomal subunits eIFs, composed of at least 29 distinct subunits, have been and interacts with other initiation factors involved in start identified (1). Mammalian eIF3, the largest initiation factor, is a codon selection. The molecular mechanisms by which multisubunit complex with an apparent molecular mass of www.mcponline.org eIF3 exerts these functions are poorly understood. Since ϳ800 kDa. This protein complex plays an essential role in its initial characterization in the 1970s, the exact size, translation by binding directly to the 40 S ribosomal subunit composition, and post-translational modifications of and promoting formation of the 43 S preinitiation complex ⅐ ⅐ mammalian eIF3 have not been rigorously determined. -

CHARACTERIZING the INTERACTION BETWEEN PDCD4 and Eif3 with RESPECT to TRANSLATION REGULATION
CHARACTERIZING THE INTERACTION BETWEEN PDCD4 AND eIF3 WITH RESPECT TO TRANSLATION REGULATION DIVYA SHARMA KHANDIGA Master of Science, Bangalore University, India 2009 A Thesis/Project Submitted to the School of Graduate Studies of the University of Lethbridge in Partial Fulfillment of the Requirements for the Degree MASTER OF SCIENCE Department of Chemistry and Biochemistry University of Lethbridge LETHBRIDGE, ALBERTA, CANADA © Divya Sharma Khandiga, 2017 CHARACTERIZING THE INTERACTION BETWEEN PDCD4 AND eIF3 WITH RESPECT TO TRANSLATION REGULATION DIVYA SHARMA KHANDIGA Date of Defense: December 12, 2017 Dr. N. Thakor Assistant Professor Ph.D. Thesis Supervisor Dr. M. Roussel Professor Ph.D. Thesis Co-supervisor Dr. U. Kothe Associate Professor Ph.D. Thesis Examination Committee Member Dr. R. Golsteyn Associate Professor Ph.D. Thesis Examination Committee Member Dr. R. Fahlman Professor Ph.D. External Examiner University of Alberta Edmonton, Alberta Dr. M. Gerken Professor Ph.D. Chair, Thesis Examination Committee Dedication To my beloved family and friends, My inspiration, my parents Subraya Sharma and Kamala Sharma My dearly loved husband Samarth, sister Dr. Lakshmi and brother-in-law Dr. Pradeep My cute little niece Mithali and nephew Aathreya My adorable brother Dr. Ganesh, sister Dr. Sharadha, Silly Vidya and little angels My loving cousins and in-laws I am grateful to have them in my life, it is their well wishes, teachings, support and love that have enabled me to achieve success and happiness in life. iii Abstract Programmed cell death protein 4 (PDCD4) inhibits IRES-mediated translation of anti- apoptotic proteins such as XIAP. PDCD4 was shown to directly interact with the XIAP IRES element and inhibit translation initiation. -

Structural Studies of the Eif4e–Vpg Complex Reveal a Direct Competition for Capped RNA: Implications for Translation
Structural studies of the eIF4E–VPg complex reveal a direct competition for capped RNA: Implications for translation Luciana Coutinho de Oliveiraa,1,2, Laurent Volpona,1, Amanda K. Rahardjoa, Michael J. Osbornea, Biljana Culjkovic-Kraljacica, Christian Trahanb, Marlene Oeffingerb,c,d, Benjamin H. Kwoka, and Katherine L. B. Bordena,3 aInstitute of Research in Immunology and Cancer, Department of Pathology and Cell Biology, Université de Montréal, Pavilion Marcelle-Coutu, Chemin Polytechnique, Montréal, QC H3T 1J4, Canada; bDepartment for Systems Biology, Institut de Recherches Cliniques de Montréal, Montréal, QC H2W 1R7, Canada; cDépartement de Biochimie et Médecine Moléculaire, Université de Montréal, Montréal, QC H3T 1J4, Canada; and dDivision of Experimental Medicine, McGill University, Montréal, QC H3A 1A3, Canada Edited by Lynne E. Maquat, University of Rochester School of Medicine and Dentistry, Rochester, NY, and approved October 16, 2019 (received for review March 19, 2019) Viruses have transformed our understanding of mammalian RNA of eIF4E can be targeted in patients to provide clinical benefit, processing, including facilitating the discovery of the methyl-7- highlighting its critical importance. guanosine (m7G)caponthe5′ end of RNAs. The m7Gcapisrequired Viruses have paved the way for our understanding of many for RNAs to bind the eukaryotic translation initiation factor eIF4E aspects of host-cell RNA processing, including m7G capping. and associate with the translation machinery across plant and ani- Indeed, studies into cytoplasmic polyhedrosis virus (CPV) infec- mal kingdoms. The potyvirus-derived viral genome-linked protein tion in silkworm and vaccinia virus (VV) in mammalian cells were (VPg) is covalently bound to the 5′ end of viral genomic RNA (gRNA) critical for the elucidation of the m7G cap structure over 40 y ago and associates with host eIF4E for successful infection. -

Structure of Ah
Eliseev, B., Yeramala, L., Leitner, A., Karuppasamy, M., Raimondeau, E., Huard, K., ... Schaffitzel, C. (2018). Structure of a human cap-dependent 48S translation pre-initiation complex. Nucleic Acids Research, 46(5), 2678- 2689. [gky054]. https://doi.org/10.1093/nar/gky054 Publisher's PDF, also known as Version of record License (if available): CC BY Link to published version (if available): 10.1093/nar/gky054 Link to publication record in Explore Bristol Research PDF-document This is the final published version of the article (version of record). It first appeared online via Oxford University Press at https://academic.oup.com/nar/advance-article/doi/10.1093/nar/gky054/4833217 . Please refer to any applicable terms of use of the publisher. University of Bristol - Explore Bristol Research General rights This document is made available in accordance with publisher policies. Please cite only the published version using the reference above. Full terms of use are available: http://www.bristol.ac.uk/pure/about/ebr-terms 2678–2689 Nucleic Acids Research, 2018, Vol. 46, No. 5 Published online 1 February 2018 doi: 10.1093/nar/gky054 Structure of a human cap-dependent 48S translation pre-initiation complex Boris Eliseev1, Lahari Yeramala1, Alexander Leitner2, Manikandan Karuppasamy1, Etienne Raimondeau1, Karine Huard1, Elena Alkalaeva3, Ruedi Aebersold2,4 and Christiane Schaffitzel1,5,* 1European Molecular Biology Laboratory, Grenoble Outstation, 71 Avenue des Martyrs, 38042 Grenoble, France, 2ETH Zurich,¨ Institute of Molecular Systems Biology, -
Type of the Paper (Article
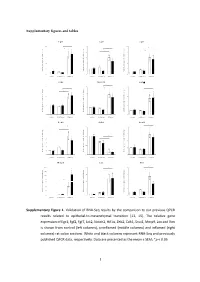
Supplementary figures and tables E g r 1 F g f2 F g f7 1 0 * 5 1 0 * * e e e * g g g * n n n * a a a 8 4 * 8 h h h * c c c d d d * l l l o o o * f f f * n n n o o o 6 3 6 i i i s s s s s s e e e r r r p p p x x x e e e 4 2 4 e e e n n n e e e g g g e e e v v v i i i t t t 2 1 2 a a a l l l e e e R R R 0 0 0 c o n tro l u n in fla m e d in fla m e d c o n tro l u n in fla m e d in fla m e d c o n tro l u n in fla m e d in fla m e d J a k 2 N o tc h 2 H if1 * 3 4 6 * * * e e e g g g n n n a a * * a * h h * h c c c 3 * d d * d l l l * o o o f f 2 f 4 n n n o o o i i i s s s s s s e e e r r 2 r p p p x x x e e e e e e n n n e e 1 e 2 g g g e e 1 e v v v i i i t t t a a a l l l e e e R R R 0 0 0 c o n tro l u n in fla m e d in fla m e d c o n tro l u n in fla m e d in fla m e d c o n tro l u n in fla m e d in fla m e d Z e b 2 C d h 1 S n a i1 * * 7 1 .5 4 * * e e e g g g 6 n n n * a a a * h h h c c c 3 * d d d l l l 5 o o o f f f 1 .0 * n n n * o o o i i i 4 * s s s s s s e e e r r r 2 p p p x x x 3 e e e e e e n n n e e e 0 .5 g g g 2 e e e 1 v v v i i i t t t a a a * l l l e e e 1 * R R R 0 0 .0 0 c o n tro l u n in fla m e d in fla m e d c o n tro l u n in fla m e d in fla m e d c o n tro l u n in fla m e d in fla m e d M m p 9 L o x V im 2 0 0 2 0 8 * * * e e e * g g g 1 5 0 * n n n * a a a * h h h * c c c 1 5 * 6 d d d l l l 1 0 0 o o o f f f n n n o o o i i i 5 0 s s s s s s * e e e r r r 1 0 4 3 0 p p p * x x x e e e * e e e n n n e e e 2 0 g g g e e e 5 2 v v v i i i t t t a a a l l l 1 0 e e e R R R 0 0 0 c o n tro l u n in fla m e d in fla m e d c o n tro l u n in fla m e d in fla m e d c o n tro l u n in fla m e d in fla m e d Supplementary Figure 1. -

Systematically Profiling the Expression of Eif3 Subunits in Glioma Reveals
Chai et al. Cancer Cell Int (2019) 19:155 https://doi.org/10.1186/s12935-019-0867-1 Cancer Cell International PRIMARY RESEARCH Open Access Systematically profling the expression of eIF3 subunits in glioma reveals the expression of eIF3i has prognostic value in IDH-mutant lower grade glioma Rui‑Chao Chai1,4,6†, Ning Wang2†, Yu‑Zhou Chang3, Ke‑Nan Zhang1,6, Jing‑Jun Li1,6, Jun‑Jie Niu5, Fan Wu1,6*, Yu‑Qing Liu1,6* and Yong‑Zhi Wang1,3,4,6* Abstract Background: Abnormal expression of the eukaryotic initiation factor 3 (eIF3) subunits plays critical roles in tumo‑ rigenesis and progression, and also has potential prognostic value in cancers. However, the expression and clinical implications of eIF3 subunits in glioma remain unknown. Methods: Expression data of eIF3 for patients with gliomas were obtained from the Chinese Glioma Genome Atlas (CGGA) (n 272) and The Cancer Genome Atlas (TCGA) (n 595). Cox regression, the receiver operating characteristic (ROC) curves= and Kaplan–Meier analysis were used to study= the prognostic value. Gene oncology (GO) and gene set enrichment analysis (GSEA) were utilized for functional prediction. Results: In both the CGGA and TCGA datasets, the expression levels of eIF3d, eIF3e, eIF3f, eIF3h and eIF3l highly were associated with the IDH mutant status of gliomas. The expression of eIF3b, eIF3i, eIF3k and eIF3m was increased with the tumor grade, and was associated with poorer overall survival [All Hazard ratio (HR) > 1 and P < 0.05]. By contrast, the expression of eIF3a and eIF3l was decreased in higher grade gliomas and was associated with better overall sur‑ vival (Both HR < 1 and P < 0.05). -

Translation Initiation Factor Eif3h Targets Specific Transcripts To
Translation initiation factor eIF3h targets specific transcripts to polysomes during embryogenesis Avik Choudhuria,b, Umadas Maitraa,1, and Todd Evansb,1 aDepartment of Developmental and Molecular Biology, Albert Einstein College of Medicine of Yeshiva University, Bronx, NY 10461; and bDepartment of Surgery, Weill Cornell Medical College, New York, NY 10065 Edited by Igor B. Dawid, The Eunice Kennedy Shriver National Institute of Child Health and Human Development, National Institutes of Health, Bethesda, MD, and approved April 30, 2013 (received for review February 14, 2013) Eukaryotic translation initiation factor 3 (eIF3) plays a central role eukaryotes. These—eIF3d, eIF3e, eIF3f, eIF3h, eIF3j, eIF3k, in translation initiation and consists of five core (conserved) sub- eIF3l, and eIF3m—were designated “non-core” subunits (4). units present in both budding yeast and higher eukaryotes. Higher In contrast to the budding yeast, the genome of the fission eukaryotic eIF3 contains additional (noncore or nonconserved) yeast Schizosaccharomyces pombe contains structural homologs subunits of poorly defined function, including sub-unit h (eIF3h), of at least five noncore (nonconserved) eIF3 subunits—eIF3d, which in zebrafish is encoded by two distinct genes (eif3ha and eIF3e, eIF3f, eIF3h, and eIF3m. The gene encoding eIF3f is eif3hb). Previously we showed that eif3ha encodes the predominant essential for growth, whereas eIF3d, eIF3e, and eIF3h are dis- isoform during zebrafish embryogenesis and that depletion of this pensable for growth and viability (5–11). However, deleted factor causes defects in the development of the brain and eyes. To strains show specific phenotypes including defects in meiosis/ investigate the molecular mechanism governing this regulation, we sporulation (6, 9, 11). -

Relevance of Translation Initiation in Diffuse Glioma Biology and Its
cells Review Relevance of Translation Initiation in Diffuse Glioma Biology and its Therapeutic Potential Digregorio Marina 1, Lombard Arnaud 1,2, Lumapat Paul Noel 1, Scholtes Felix 1,2, Rogister Bernard 1,3 and Coppieters Natacha 1,* 1 Laboratory of Nervous System Disorders and Therapy, GIGA-Neurosciences Research Centre, University of Liège, 4000 Liège, Belgium; [email protected] (D.M.); [email protected] (L.A.); [email protected] (L.P.N.); [email protected] (S.F.); [email protected] (R.B.) 2 Department of Neurosurgery, CHU of Liège, 4000 Liège, Belgium 3 Department of Neurology, CHU of Liège, 4000 Liège, Belgium * Correspondence: [email protected] Received: 18 October 2019; Accepted: 26 November 2019; Published: 29 November 2019 Abstract: Cancer cells are continually exposed to environmental stressors forcing them to adapt their protein production to survive. The translational machinery can be recruited by malignant cells to synthesize proteins required to promote their survival, even in times of high physiological and pathological stress. This phenomenon has been described in several cancers including in gliomas. Abnormal regulation of translation has encouraged the development of new therapeutics targeting the protein synthesis pathway. This approach could be meaningful for glioma given the fact that the median survival following diagnosis of the highest grade of glioma remains short despite current therapy. The identification of new targets for the development of novel therapeutics is therefore needed in order to improve this devastating overall survival rate. This review discusses current literature on translation in gliomas with a focus on the initiation step covering both the cap-dependent and cap-independent modes of initiation. -

Stress Granule Formation, Disassembly, and Composition Are Regulated by Alphavirus ADP-Ribosylhydrolase Activity
Stress granule formation, disassembly, and composition are regulated by alphavirus ADP-ribosylhydrolase activity Aravinth Kumar Jayabalana, Srivathsan Adivarahanb, Aakash Koppulac, Rachy Abrahamd, Mona Batishc,e, Daniel Zenklusenb, Diane E. Griffind, and Anthony K. L. Leunga,f,g,1 aDepartment of Biochemistry and Molecular Biology, Bloomberg School of Public Health, Johns Hopkins University, Baltimore, MD 21205; bDépartment de Biochimie et Médecine Moléculaire, Université de Montréal, Montréal, QC H3T 1J4, Canada; cDepartment of Biological Sciences, University of Delaware, Newark, DE 19716; dW. Harry Feinstone Department of Molecular Microbiology and Immunology, Bloomberg School of Public Health, Johns Hopkins University, Baltimore, MD 21205; eDepartment of Medical and Molecular Sciences, University of Delaware, Newark, DE 19716; fDepartment of Molecular Biology and Genetics, School of Medicine, Johns Hopkins University, Baltimore, MD 21205; and gDepartment of Oncology, School of Medicine, Johns Hopkins University, Baltimore, MD 21205 Edited by Thomas Shenk, Princeton University, Princeton, NJ, and approved December 31, 2020 (received for review October 22, 2020) While biomolecular condensates have emerged as an important later infection stages, many viruses instead suppress SG forma- biological phenomenon, mechanisms regulating their composition tion or disassemble SGs altogether. The mechanisms underlying and the ways that viruses hijack these mechanisms remain unclear. this switch, and its physiological function, remain unclear. The mosquito-borne alphaviruses cause a range of diseases from SG formation and disassembly are regulated by posttransla- rashes and arthritis to encephalitis, and no licensed drugs are tional modifications of proteins, including those that conjugate available for treatment or vaccines for prevention. The alphavirus simple chemical groups, attach polypeptides, and add nucleotides virulence factor nonstructural protein 3 (nsP3) suppresses the for- as in the case of ADP-ribosylation (15–21). -

Interactome Mapping of Eif3a in a Colon Cancer and an Immortalized Embryonic Cell Line Using Proximity-Dependent Biotin Identification
cancers Article Interactome Mapping of eIF3A in a Colon Cancer and an Immortalized Embryonic Cell Line Using Proximity-Dependent Biotin Identification Diep-Khanh Vo 1 , Alexander Engler 2 , Darko Stoimenovski 1, Roland Hartig 3 , Thilo Kaehne 2, Thomas Kalinski 1, Michael Naumann 2 , Johannes Haybaeck 1,4,5,6,† and Norbert Nass 1,*,†,‡ 1 Department of Pathology, Medical Faculty, Otto-von-Guericke University Magdeburg, D-39120 Magdeburg, Germany; [email protected] (D.-K.V.); [email protected] (D.S.); [email protected] (T.K.); [email protected] (J.H.) 2 Institute of Experimental Internal Medicine, Medical Faculty, Otto von Guericke University, D-39120 Magdeburg, Germany; [email protected] (A.E.); [email protected] (T.K.); [email protected] (M.N.) 3 Institute of Molecular and Clinical Immunology, Otto von Guericke University Magdeburg, Leipziger Str. 44, D-39120 Magdeburg, Germany; [email protected] 4 Department of Pathology, Neuropathology, and Molecular Pathology, Medical University of Innsbruck, A-6020 Innsbruck, Austria 5 Department of Pathology, Diagnostic & Research Center for Molecular BioMedicine, Institute of Pathology, Medical University of Graz, A-8010 Graz, Austria 6 Center for Biomarker Research in Medicine, A-8010 Graz, Austria * Correspondence: [email protected] † These authors contributed equally to the supervision of the project. ‡ Current address: Department of Internal Medicine I, Dessau Medical Center, D-06847 Dessau, Germany. Citation: Vo, D.-K.; Engler, A.; Stoimenovski, D.; Hartig, R.; Kaehne, Simple Summary: The behavior of a cancer cell is greatly influenced by its proteome, which is the T.; Kalinski, T.; Naumann, M.; Haybaeck, result of protein biosynthesis, modification and degradation. -

Vaccine-Increased Seq ID Unigene ID Uniprot ID Gene Names
BMJ Publishing Group Limited (BMJ) disclaims all liability and responsibility arising from any reliance Supplemental material placed on this supplemental material which has been supplied by the author(s) J Immunother Cancer Vaccine-Increased Seq ID Unigene ID Uniprot ID Gene Names 1_HSPA1A_3303 Hs.274402 P0DMV8 HSPA1A HSP72 HSPA1 HSX70 100_AKAP17A_8227 Hs.522572 Q02040 AKAP17A CXYorf3 DXYS155E SFRS17A XE7 1000_H2AFY_9555 Hs.420272 O75367 H2AFY MACROH2A1 1001_ITPK1_3705 Hs.308122 Q13572 ITPK1 1002_PTPN11_5781 Hs.506852 Q06124 PTPN11 PTP2C SHPTP2 1003_EIF3J_8669 Hs.404056 O75822 EIF3J EIF3S1 PRO0391 1004_TRIP12_9320 Hs.591633 Q14669 TRIP12 KIAA0045 ULF 1006_YEATS2_55689 Hs.632575 Q9ULM3 YEATS2 KIAA1197 1007_SEL1L3_23231 Hs.479384 Q68CR1 SEL1L3 KIAA0746 1008_IDH1_3417 Hs.593422 O75874 IDH1 PICD 101_HSPH1_10808 Hs.36927 Q92598 HSPH1 HSP105 HSP110 KIAA0201 1010_LDLR_3949 Hs.213289 P01130 LDLR 1011_FAM129B_64855 Hs.522401 Q96TA1 NIBAN2 C9orf88 FAM129B 1012_MAP3K5_4217 Hs.186486 Q99683 MAP3K5 ASK1 MAPKKK5 MEKK5 1013_NEFH_4744 Hs.198760 P12036 NEFH KIAA0845 NFH 1014_RAP1B_5908 Hs.369920 P61224 RAP1B OK/SW-cl.11 1015_MCCC1_56922 Hs.47649 Q96RQ3 MCCC1 MCCA 1017_MT1E_4493 Hs.534330 P04732 MT1E 1022_TXNDC5_81567 Hs.150837 Q8NBS9 TXNDC5 TLP46 UNQ364/PRO700 1023_STRA13_201254 Hs.37616 O14503 BHLHE40 BHLHB2 DEC1 SHARP2 STRA13 1024_NPEPPS_9520 Hs.443837 P55786 NPEPPS PSA 1025_YIPF6_286451 Hs.82719 Q96EC8 YIPF6 1026_CLIP1_6249 Hs.524809 P30622 CLIP1 CYLN1 RSN 1027_SRSF7_6432 Hs.309090 Q16629 SRSF7 SFRS7 103_RPS25_6230 Hs.512676 P62851 RPS25 1031_SOCS7_30837