Modeling the Phenotype of Spinal Muscular Atrophy by the Direct Conversion of Human Fibroblasts to Motor Neurons
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Regulation of Adult Neurogenesis in Mammalian Brain
International Journal of Molecular Sciences Review Regulation of Adult Neurogenesis in Mammalian Brain 1,2, 3, 3,4 Maria Victoria Niklison-Chirou y, Massimiliano Agostini y, Ivano Amelio and Gerry Melino 3,* 1 Centre for Therapeutic Innovation (CTI-Bath), Department of Pharmacy & Pharmacology, University of Bath, Bath BA2 7AY, UK; [email protected] 2 Blizard Institute of Cell and Molecular Science, Barts and the London School of Medicine and Dentistry, Queen Mary University of London, London E1 2AT, UK 3 Department of Experimental Medicine, TOR, University of Rome “Tor Vergata”, 00133 Rome, Italy; [email protected] (M.A.); [email protected] (I.A.) 4 School of Life Sciences, University of Nottingham, Nottingham NG7 2HU, UK * Correspondence: [email protected] These authors contributed equally to this work. y Received: 18 May 2020; Accepted: 7 July 2020; Published: 9 July 2020 Abstract: Adult neurogenesis is a multistage process by which neurons are generated and integrated into existing neuronal circuits. In the adult brain, neurogenesis is mainly localized in two specialized niches, the subgranular zone (SGZ) of the dentate gyrus and the subventricular zone (SVZ) adjacent to the lateral ventricles. Neurogenesis plays a fundamental role in postnatal brain, where it is required for neuronal plasticity. Moreover, perturbation of adult neurogenesis contributes to several human diseases, including cognitive impairment and neurodegenerative diseases. The interplay between extrinsic and intrinsic factors is fundamental in regulating neurogenesis. Over the past decades, several studies on intrinsic pathways, including transcription factors, have highlighted their fundamental role in regulating every stage of neurogenesis. However, it is likely that transcriptional regulation is part of a more sophisticated regulatory network, which includes epigenetic modifications, non-coding RNAs and metabolic pathways. -

E Proteins and ID Proteins: Helix-Loop-Helix Partners in Development and Disease
View metadata, citation and similar papers at core.ac.uk brought to you by CORE provided by Elsevier - Publisher Connector Developmental Cell Review E Proteins and ID Proteins: Helix-Loop-Helix Partners in Development and Disease Lan-Hsin Wang1 and Nicholas E. Baker1,2,3,* 1Department of Genetics, Albert Einstein College of Medicine, 1300 Morris Park Avenue, Bronx, NY 10461, USA 2Department of Developmental and Molecular Biology, Albert Einstein College of Medicine, 1300 Morris Park Avenue, Bronx, NY 10461, USA 3Department of Ophthalmology and Visual Sciences, Albert Einstein College of Medicine, 1300 Morris Park Avenue, Bronx, NY 10461, USA *Correspondence: [email protected] http://dx.doi.org/10.1016/j.devcel.2015.10.019 The basic Helix-Loop-Helix (bHLH) proteins represent a well-known class of transcriptional regulators. Many bHLH proteins act as heterodimers with members of a class of ubiquitous partners, the E proteins. A widely expressed class of inhibitory heterodimer partners—the Inhibitor of DNA-binding (ID) proteins—also exists. Genetic and molecular analyses in humans and in knockout mice implicate E proteins and ID proteins in a wide variety of diseases, belying the notion that they are non-specific partner proteins. Here, we explore relationships of E proteins and ID proteins to a variety of disease processes and highlight gaps in knowledge of disease mechanisms. E proteins and Inhibitor of DNA-binding (ID) proteins are widely conferring DNA-binding specificity and transcriptional activation expressed transcriptional regulators with very general functions. on heterodimers with the ubiquitous E proteins (Figure 1). They are implicated in diseases by evidence ranging from Another class of pervasive HLH proteins acts in opposition to confirmed Mendelian inheritance, association studies, and E proteins. -

Lineage Transcription Factors Co-Regulate Subtype-Specific Genes Providing a Roadmap For
bioRxiv preprint doi: https://doi.org/10.1101/2020.08.13.249029; this version posted August 14, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. Lineage transcription factors co-regulate subtype-specific genes providing a roadmap for systematic identification of small cell lung cancer vulnerabilities Karine Pozo1,2, Rahul K. Kollipara3, Demetra P. Kelenis1, Kathia E. Rodarte1, Xiaoyang Zhang4, John D. Minna5,6,7,8 and Jane E. Johnson1,6,8 1Department of Neuroscience, 2Department of Surgery, 3McDermott Center for Human Growth and Development, UT Southwestern Medical Center, Dallas, TX 75390, USA 4Department of Oncological Sciences, Huntsman Cancer Institute, University of Utah, Salt Lake City, UT 5Hamon Center for Therapeutic Oncology Research, 6Simmons Comprehensive Cancer Center, 7Department of Internal Medicine, 8Department of Pharmacology, UT Southwestern Medical Center, Dallas, TX 75390, USA JDM receives licensing fees for lung cancer lines from the NIH and UT Southwestern. 1 bioRxiv preprint doi: https://doi.org/10.1101/2020.08.13.249029; this version posted August 14, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. ABSTRACT Lineage-defining transcription factors (LTFs) play key roles in tumor cell growth, making them highly attractive, but currently “undruggable”, small cell lung cancer (SCLC) vulnerabilities. -

Retinoblastoma Tumor Suppressor Protein in Pancreatic Progenitors Controls Α- and Β-Cell Fate
Retinoblastoma tumor suppressor protein in pancreatic progenitors controls α- and β-cell fate Erica P. Caia,b, Xiaohong Wuc, Stephanie A. Schroera, Andrew J. Eliad,e, M. Cristina Nostroa,f, Eldad Zacksenhausa, and Minna Wooa,b,e,g,1 aToronto General Research Institute and fMcEwen Centre for Regenerative Medicine, University Health Network, Toronto, ON, Canada M5G 1L7; bInstitute of Medical Science, University of Toronto, Toronto, ON, Canada M5S 1A8; cThe First Affiliated Hospital, Nanjing Medical University, Nanjing, Jiangsu 210029, China; dCampbell Family Institute for Breast Cancer Research, University Health Network, Toronto, ON, Canada M5G 2M9; eDepartment of Medical Biophysics, University of Toronto, Toronto, ON, Canada M5G 2M9; and gDivision of Endocrinology, Department of Medicine, Toronto General Hospital, University Health Network, University of Toronto, Toronto, ON, Canada M5G 2C4 Edited by Tak W. Mak, The Campbell Family Institute for Breast Cancer Research, Ontario Cancer Institute at Princess Margaret Hospital, University Health Network, Toronto, ON, Canada, and approved July 16, 2013 (received for review February 20, 2013) Pancreatic endocrine cells expand rapidly during embryogenesis apoptosis (15), in a highly cell-specific and context-dependent by neogenesis and proliferation, but during adulthood, islet cells manner (16, 17). Although Rb is critical in regulating cell cycle have a very slow turnover. Disruption of murine retinoblastoma entry in proliferating cells, its role in postmitotic cells is more tumor suppressor protein (Rb) in mature pancreatic β-cells has limited. For example, Rb deficiency in proliferating myoblasts a limited effect on cell proliferation. Here we show that deletion induces increased proliferation and apoptosis (18), whereas fi fi of Rb during embryogenesis in islet progenitors leads to an in- Rb de ciency in postmitotic muscle bers does not lead to any crease in the neurogenin 3-expressing precursor cell population, defects (19). -
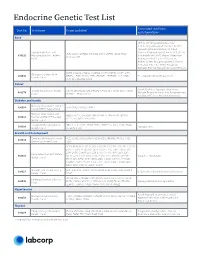
Endocrine Genetic Test List
Endocrine Genetic Test List Associated conditions Test No. Test Name Genes Included and phenotypes Bone HPP, X-Linked Hypophosphatemia, X-Linked Hypophosphatemic Rickets, XLH, Hypophosphatemic Rickets, X-Linked Hypophosphatasia and Dominant Hypophosphatemic Rickets (XLHR), ALPL, CLCN5, CYP2R1, CYP27B1, DMP1, ENPP1, FGF23, PHEX, 630292 Hypophosphatemic Rickets X-Linked Rickets (XLR), Vitamin D-Resistant SLC34A3, VDR Panel Rickets, X-Linked Vitamin D-Resistant Rickets (VDRR), Hypophosphatemic Vitamin D-Resistant Rickets (HPDR), Phosphate Diabetes, Familial Hypophosphatemic Rickets BMP1, COL1A1, COL1A2, CREB3L1, CRTAP, FKBP10, IFITM5, LRP5, Osteogenesis Imperfecta 630543 MBTPS2, P3H1, PLOD2, PPIB, SERPINF1, SERPINH1, SP7, SPARC, OI, Juvenile Primary Osteoporosis Genetic Panel TENT5A, TMEM38B, WNT1 Cancer Familial Isolated Hyperparathyroidism, VistaSeq® Endocrine Cancer CDC73, MAX, MEN1, NF1, PRKAR1A, PTEN, RET, SDHB, SDHC, SDHD, 481374 Multiple Endocrine Neoplasia, Paraganglioma, Panel* TMEM127, TP53 and VHL Parathyroid Cancer, Pheochromocytoma Diabetes and Insulin Maturity-Onset Diabetes of the 630568 GCK, HNF1A, HNF1B, HNF4A Young (MODY) 4-gene Panel Maturity-Onset Diabetes of ABCC8, APPL1, BLK, GCK, HNF1A, HNF1B, HNF4A, INS, KCNJ11, 630513 the Young (MODY) Expanded KLF11, NEUROD1, PAX4, PDX1 Genetic Panel Congenital Hyperinsulinism ABCC8, GCK, GLUD1, HADH, HNF1A, HNF4A, KCNJ11, PGM1, PMM2, 630500 Hypoglycemia Genetic Panel SLC16A1, UCP2 Growth and Development Combined Pituitary Hormone GLI1, HESX1, LHX3, LHX4, OTX2, POU1F1, PROKR2, PROP1, -

Identification of C2CD4A As a Human Diabetes Susceptibility Gene with a Role in Β Cell Insulin Secretion
Identification of C2CD4A as a human diabetes susceptibility gene with a role in β cell insulin secretion Taiyi Kuoa, Michael J. Kraakmana, Manashree Damleb,c, Richard Gilla, Mitchell A. Lazarb,c,1, and Domenico Accilia,1 aDepartment of Medicine, Berrie Diabetes Center, Columbia University College of Physicians and Surgeons, New York, NY 10032; bThe Institute for Diabetes, Obesity, and Metabolism, University of Pennsylvania Perelman School of Medicine, Philadelphia, PA 19104; and cDivision of Endocrinology, Diabetes, and Metabolism, Department of Medicine, University of Pennsylvania Perelman School of Medicine, Philadelphia, PA 19104 Contributed by Mitchell A. Lazar, July 31, 2019 (sent for review March 14, 2019; reviewed by Alvin C. Powers and Andrew F. Stewart) Fine mapping and validation of genes causing β cell failure from targets. To circumvent this obstacle, we generated FoxO1- susceptibility loci identified in type 2 diabetes genome-wide asso- GFPVenus (Venus) reporter knockin mice, and utilized 2-photon ciation studies (GWAS) poses a significant challenge. The VPS13C- microscopy to track its subcellular localization in pancreatic β C2CD4A-C2CD4B locus on chromosome 15 confers diabetes suscep- cells. We next performed genome-wide FoxO1 chromatin immu- tibility in every ethnic group studied to date. However, the causative noprecipitation sequencing (ChIP-seq) to identify its genomic gene is unknown. FoxO1 is involved in the pathogenesis of β cell targets as well as superenhancers encompassing FoxO1 sites. A dysfunction, but its link to human diabetes GWAS has not been comparative analysis of human islet and murine β cell super- explored. Here we generated a genome-wide map of FoxO1 super- enhancers revealed C2CD4A, a gene encoding an IL-1β–induced β enhancers in chemically identified cells using 2-photon live-cell nuclear protein (18) embedded among several SNPs conferring imaging to monitor FoxO1 localization. -

TWIST1 Homo- and Heterodimers Orchestrate Specificity Control in Embryonic Stem Cell
bioRxiv preprint doi: https://doi.org/10.1101/672824; this version posted June 16, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. TWIST1 homo- and heterodimers orchestrate specificity control in embryonic stem cell lineage differentiation and craniofacial development Xiaochen Fan1,2, a+, Ashley J. Waardenberg3, b, Madeleine Demuth1, c, Pierre Osteil1, Jane Sun1, David A.F. Loebel1,2, Mark Graham4, Patrick P.L. Tam1,2 and Nicolas Fossat1,2, d 1 Embryology Unit, Children’s Medical Research Institute, The University of Sydney, 2 The University of Sydney, School of Medical Sciences, Faculty of Medicine and Health, 3 Bioinformatics Group, and 4 Synapse Proteomics Group, Children’s Medical Research Institute, The University of Sydney. +Corresponding author: Xiaochen Fan email: [email protected] Present address: a Department of Bioengineering, University of California, San Diego, La Jolla, CA 92093, USA b Centre for Tropical Bioinformatics and Molecular Biology, James Cook University, QLD 4870 Australia. c Developmental Dynamics Laboratory, The Francis Crick Institute, London NW1 1AT, UK. d Copenhagen Hepatitis C Program (CO-HEP), Department of Immunology and Microbiology, University of Copenhagen, and Department of Infectious Diseases, Hvidovre Hospital, Denmark. 1 bioRxiv preprint doi: https://doi.org/10.1101/672824; this version posted June 16, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. -

Homozygous Mutations in NEUROD1 Are Responsible for a Novel Syndrome of Permanent Neonatal Diabetes and Neurological Abnormalities Oscar Rubio-Cabezas,1,2 Jayne A.L
BRIEF REPORT Homozygous Mutations in NEUROD1 Are Responsible for a Novel Syndrome of Permanent Neonatal Diabetes and Neurological Abnormalities Oscar Rubio-Cabezas,1,2 Jayne A.L. Minton,1 Iren Kantor,3 Denise Williams,4 Sian Ellard,1 and Andrew T. Hattersley1 OBJECTIVE—NEUROD1 is expressed in both developing and mature -cells. Studies in mice suggest that this basic helix-loop- helix transcription factor is critical in the development of endo- onogenic permanent neonatal diabetes crine cell lineage. Heterozygous mutations have previously been (PNDM) is typically diagnosed within the identified as a rare cause of maturity-onset diabetes of the young first 6 months of birth in contrast to polygenic (MODY). We aimed to explore the potential contribution of Mautoimmune type 1 diabetes, which is usually NEUROD1 mutations in patients with permanent neonatal diagnosed later in childhood or in young adults (1,2). diabetes. PNDM is both phenotypically and genetically heteroge- RESEARCH DESIGN AND METHODS—We sequenced the neous. Most patients present with isolated diabetes, but in NEUROD1 gene in 44 unrelated patients with permanent neona- some cases diabetes appears in the context of a more tal diabetes of unknown genetic etiology. complex multisystemic syndrome. Dominant mutations in three genes (KCNJ11, ABCC8, and INS) are the cause RESULTS—Two homozygous mutations in NEUROD1 (c.427_ of PNDM in ϳ50% of cases, and in the majority diabetes 428del and c.364dupG) were identified in two patients. Both is an isolated finding (3,4). Recessive mutations, auto- mutations introduced a frameshift that would be predicted to generate a truncated protein completely lacking the activating somal or X-linked, have been described in 10 genes domain. -

NEUROD1 Is Required for the Early and Endocrine Differentiation in The
International Journal of Molecular Sciences Article NEUROD1 Is Required for the Early α and β Endocrine Differentiation in the Pancreas Romana Bohuslavova 1, Ondrej Smolik 1,2 , Jessica Malfatti 1,2, Zuzana Berkova 3 , Zaneta Novakova 1, Frantisek Saudek 3 and Gabriela Pavlinkova 1,* 1 Institute of Biotechnology CAS, 25250 Vestec, Czech Republic; [email protected] (R.B.); [email protected] (O.S.); [email protected] (J.M.); [email protected] (Z.N.) 2 Department of Cell Biology, Faculty of Science, Charles University, 12843 Prague, Czech Republic 3 Laboratory of Pancreatic Islets, Institute for Clinical and Experimental Medicine, 14021 Prague, Czech Republic; [email protected] (Z.B.); [email protected] (F.S.) * Correspondence: [email protected]; Tel.: +420-32587-3794 Abstract: Diabetes is a metabolic disease that involves the death or dysfunction of the insulin- secreting β cells in the pancreas. Consequently, most diabetes research is aimed at understanding the molecular and cellular bases of pancreatic development, islet formation, β-cell survival, and insulin secretion. Complex interactions of signaling pathways and transcription factor networks regulate the specification, growth, and differentiation of cell types in the developing pancreas. Many of the same regulators continue to modulate gene expression and cell fate of the adult pancreas. The transcription factor NEUROD1 is essential for the maturation of β cells and the expansion of the pancreatic islet cell mass. Mutations of the Neurod1 gene cause diabetes in humans and mice. However, the different aspects of the requirement of NEUROD1 for pancreas development are not fully understood. -

P107 in the Public Eye: an Rb Understudy and More
Wirt and Sage Cell Division 2010, 5:9 http://www.celldiv.com/content/5/1/9 REVIEW Open Access p107Review in the public eye: an Rb understudy and more Stacey E Wirt1,2 and Julien Sage*1,2 Abstract p107 and its related family members Rb and p130 are critical regulators of cellular proliferation and tumorigenesis. Due to the extent of functional overlap within the Rb family, it has been difficult to assess which functions are exclusive to individual members and which are shared. Like its family members, p107 can bind a variety of cellular proteins to affect the expression of many target genes during cell cycle progression. Unlike Rb and p130, p107 is most highly expressed during the G1 to S phase transition of the cell cycle in actively dividing cells and accumulating evidence suggests a role for p107 during DNA replication. The specific roles for p107 during differentiation and development are less clear, although emerging studies suggest that it can cooperate with other Rb family members to control differentiation in multiple cell lineages. As a tumor suppressor, p107 is not as potent as Rb, yet studies in knockout mice have revealed some tumor suppressor functions in mice, depending on the context. In this review, we identify the unique and overlapping functions of p107 during the cell cycle, differentiation, and tumorigenesis. Review suppression. Additionally, the overlapping functions of Introduction each of the three genes further complicates our under- The Rb tumor suppressor was first identified as the gene standing of how they control critical cellular functions whose loss causes hereditary retinoblastoma in children such as exit from and entry into the cell cycle, differentia- [1-4]. -

Patel, a Et Al. 1 Integrative Genomic and Epigenomic Analyses Identify A
bioRxiv preprint doi: https://doi.org/10.1101/852939; this version posted November 29, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Patel, A et al. 1 Integrative genomic and epigenomic analyses identify a distinct role of c-Myc and L-Myc for lineage determination in small cell lung cancer Ayushi S. Patel1,2, Seungyeul Yoo3,4, Ranran Kong1,2,5, Takashi Sato1,2, Maya Fridrikh1,2, Abhilasha Sinha1,2, German Nudelman6, Charles A. Powell1,2, Mary Beth Beasley7, Jun Zhu2,3,4, Hideo Watanabe1,2,3,8* 1Division of Pulmonary, Critical Care and Sleep Medicine, Department of Medicine, Icahn School of Medicine at Mount Sinai, New York, NY 10029, USA 2Tisch Cancer Institute, Icahn School of Medicine at Mount Sinai, New York, NY 10029, USA 3Department of Genetics and Genomic Sciences, Icahn School of Medicine at Mount Sinai, New York, NY 10029, USA 4Sema4, a Mount Sinai venture, Stamford, CT 06902, USA. 5Department of Thoracic Surgery, The Second Affiliated Hospital of Medical School, Xi'an Jiaotong University, Xi'an, Shaanxi, 710004, China 6 Department of Neurology, Icahn School of Medicine at Mount Sinai, New York, NY 10029, USA 7Department of Pathology and Laboratory Medicine, Icahn School of Medicine at Mount Sinai, New York, NY 10029, USA 8Lead Contact *Correspondence: [email protected] bioRxiv preprint doi: https://doi.org/10.1101/852939; this version posted November 29, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. -

17-Β Estradiol Regulates Proglucagon-Derived Peptide Secretion in Mouse and Human Α- and L Cells
17-β Estradiol regulates proglucagon-derived peptide secretion in mouse and human α- and L cells Sandra Handgraaf, … , Jacques Philippe, Yvan Gosmain JCI Insight. 2018;3(7):e98569. https://doi.org/10.1172/jci.insight.98569. Research Article Endocrinology Metabolism Clinical and experimental data indicate a beneficial effect of estrogens on energy and glucose homeostasis associated with improved insulin sensitivity and positive effects on insulin secretion. The aim of the study was to investigate the impact of estrogens on proglucagon-producing cells, pancreatic α cells, and enteroendocrine L cells. The consequences of sexual hormone deprivation were evaluated in ovariectomized mice (ovx). Ovx mice exhibited impaired glucose tolerance during oral glucose tolerance tests (OGTT), which was associated with decreased GLP-1 intestinal and pancreatic secretion and content, an effect that was reversed by estradiol (E2) treatment. Indeed, E2 increased oral glucose–induced GLP-1 secretion in vivo and GLP-1 secretion from primary culture of mouse and human α cells through the activation of all 3 estrogen receptors (ERs), whereas E2-induced GLP-1 secretion from mouse and human intestinal explants occurred only by ERβ activation. Underlying the implication of ERβ, its selective agonist WAY20070 was able to restore glucose tolerance in ovx mice at least partly through plasma GLP-1 increase. We conclude that E2 directly controls both α- and L cells to increase GLP-1 secretion, in addition to its effects on insulin and glucagon secretion, highlighting