Multifunctional Delivery Systems for Peptide Nucleic Acids
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Rnai in Primary Cells and Difficult-To-Transfect Cell Lines
Automated High Throughput Nucleofection® RNAi in Primary Cells and Difficult-to-Transfect Cell Lines Claudia Merz, Bayer Schering Pharma AG, Berlin, Germany; Andreas Schroers, amaxa AG, Cologne, Germany; Eric Willimann, Tecan AG, Männedorf, Switzerland. Introduction Materials & Methods - Workflow Using primary cells for RNAi based applications such as target identification or – validation, requires a highly efficient transfection displaying the essential steps of the automated Nucleofector® Process: technology in combination with a reliable and robust automation system. To accomplish these requirements we integrated the amaxa 1. Transfer of the cells to the Nucleocuvette™ plate, 96-well Shuttle® in a Tecan Freedom EVO® cell transfection workstation which is based on Tecan’s Freedom EVO® liquid handling 2. Addition of the siRNA, (Steps 1 and 2 could be exchanged), platform and include all the necessary components and features for unattended cell transfection. 3. Nucleofection® process, 4. Addition of medium, Count Cells 5. Transfer of transfected cells to cell culture plate for incubation ® Nucleofector Technology prior to analysis. Remove Medium The 96-well Shuttle® combines high-throughput compatibility with the Nucleofector® Technology, which is a non-viral transfection method ideally suited for primary cells and hard-to-transfect cell lines based on a combination of buffers and electrical parameters. Nucleocuvette Plate Add Nucleofector +– The basic principle and benefits of the (empty) Solution Cell of interest Gene of interest Nucleofector® -

Peptide Nucleic Acids, Morpholinos and Related Antisense Biomolecules
MEDICAL INTELLIGENCE UNIT Peptide Nucleic Acids, Morpholinos and Related Antisense Biomolecules Christopher G. Janson, M.D. Departments of Neurosurgery, Neurology and Molecular Genetics Cell and Gene Therapy Center UMDNJ-Robert Wood Johnson Medical School Camden, New Jersey, U.S.A. Matthew J. During, M.D., ScD. Department of Molecular Medicine and Pathology University of Auckland Auckland, New Zealand LANDES BIOSCIENCE / EUREKAH.COM KLUWER ACADEMIC / PLENUM PUBLISHERS GEORGETOWN, TEXAS NEW YORK, NEW YORK USA U.SA PEPTIDE NUCLEIC ACIDS, MORPHOLINOS AND RELATED ANTISENSE BIOMOLECULES Medical Intelligence Unit Landes Bioscience / Eurekah.com Kluwer Academic / Plenum Publishers Copyright ©2006 Eurekah.com and Kluwer Academic / Plenum Publishers All rights reserved. No part of this book may be reproduced or transmitted in any form or by any means, electronic or mechanical, including photocopy, recording, or any information storage and retrieval system, without permission in writing from the publisher, vdth the exception of any material supplied specifically for the purpose of being entered and executed on a computer system; for exclusive use by the Purchaser of the work. Printed in the U.S.A. Kluwer Academic / Plenum PubHshers, 233 Spring Street, New York, New York, U.S.A. 10013 http ://www.wkap .nl/ Please address all inquiries to the Publishers: Landes Bioscience / Eurekah.com, 810 South Church Street, Georgetown, Texas, U.S.A. 78626 Phone: 512/ 863 7762; FAX: 512/ 863 0081 http ://v^^ww.eurekah. com http://www.landesbioscience.com Peptide -
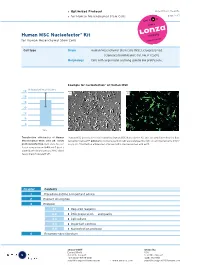
Optimized Protocol for Human Mesenchymal Stem Cells
› Optimized Protocol DPE-1001 Vs. 05-2005 › for Human Mesenchymal Stem Cells page 1 of 7 Human MSC Nucleofector® Kit for Human Mesenchymal Stem Cells Cell type Origin Human Mesenchymal Stem Cells (MSC), cryopreserved [Clonetics/BioWhittaker; Cat. No. PT-2501]. Morphology Cells with large nuclei and long spindle like protrusions.. Example for nucleofection® of Human MSC % transfection efficiency 60 A B 50 40 30 20 10 0 24 h Transfection efficiencies of Human Human MSC were nucleofected using the Human MSC Nucleofector Kit and a plasmid encoding the fluo- Mesenchymal Stem cells 24 hours rescent protein eGFP. 24 hours post-nucleofection cells were analyzed by light (A) and fluorescence micro- post nucleofection. Cells were nucleo- scopy (B). Transfection efficiencies of around 80% can be reached with eGFP. fected using program U-23 and 5 µg of a plasmid encoding the mouse MHC class I heavy chain molecule H-2Kk. Chapter Contents 1 Procedure outline & important advice 2 Product description 3 Protocol 3.1 › Required reagents 3.2 › DNA preparation and quality 3.3 › Cell culture 3.4 › Important controls 3.5 › Nucleofection protocol 4 Recommended literature 5 amaxa GmbH amaxa Inc. Europe/World USA Scientific Support Scientific Support +49 (0)221-99199-400 (240) 632-9110 [email protected] › www.amaxa.com [email protected] › Optimized Protocol DPE-1001 Vs. 05-2005 › for Human Mesenchymal Stem Cells page 2 of 7 1 Procedure outline & important advice Procedure outline Important advice 1. Preparation of cells. › Use MSCGM BulletKit (stored < 2 d at 4°C) (For details see 3.3.) › Passage interval: after reaching 70% confluency. -

New Constrained Amino Acids and Peptide Nucleic Acid Building Blocks for the Construction of Bio-Polymers Alexandra Gresika
New constrained amino acids and peptide nucleic acid building blocks for the construction of bio-polymers Alexandra Gresika To cite this version: Alexandra Gresika. New constrained amino acids and peptide nucleic acid building blocks for the construction of bio-polymers. Other. Université Côte d’Azur, 2018. English. NNT : 2018AZUR4104. tel-02073232 HAL Id: tel-02073232 https://tel.archives-ouvertes.fr/tel-02073232 Submitted on 19 Mar 2019 HAL is a multi-disciplinary open access L’archive ouverte pluridisciplinaire HAL, est archive for the deposit and dissemination of sci- destinée au dépôt et à la diffusion de documents entific research documents, whether they are pub- scientifiques de niveau recherche, publiés ou non, lished or not. The documents may come from émanant des établissements d’enseignement et de teaching and research institutions in France or recherche français ou étrangers, des laboratoires abroad, or from public or private research centers. publics ou privés. THÈSE DE DOCTORAT Nouveaux synthons contraints de type - amino acides et PNA en vue de l´élaboration de bio-foldamères New constrained amino acids and peptide nucleic acid building blocks for the construction of biopolymers Alexandra Gresika Molécules Bioactives Présentée en vue de l’obtention Devant le jury, composé de : du grade de docteur en Chimie Muriel Amblard, Directrice de Recherche d’Université Côte d’Azur CNRS, IBMM UMR 5247, Université de Dirigée par : Nadia Patino Montpellier Karine Alvarez, Chargée de Recherche Soutenue le : 16.11.2018 CNRS, AFBM UMR 7257, Université Aix- Marseille Mohamed Mehiri, Maître de Conférences, ICN UMR 7272, Université Côte d’Azur Nadia Patino, Professeur des Universités, ICN UMR 7272, Université Côte d’Azur 1 New constrained amino acids and peptide nucleic acid building blocks for the construction of bio-polymers. -

Methods of Transfection with Messenger RNA Gene Vectors
View metadata, citation and similar papers at core.ac.uk brought to you by CORE provided by Spiral - Imperial College Digital Repository Chapter 2 Methods of Transfection with Messenger RNA Gene Vectors Oleg E. Tolmachov and Tanya Tolmachova Additional information is available at the end of the chapter http://dx.doi.org/10.5772/61688 Abstract Non-viral gene delivery vectors with messenger RNA (mRNA) as a carrier of genetic information are among the staple gene transfer vectors for research in gene therapy, gene vaccination and cell fate reprogramming. As no passage of genetic cargo in and out of the nucleus is required, mRNA-based vectors typically offer the following five advantages: 1) fast start of transgene expression; 2) ability to express genes in non- dividing cells with an intact nuclear envelope; 3) insensitivity to the major gene silencing mechanisms, which operate in the nucleus; 4) absence of potentially mutagenic genomic insertions; 5) high cell survival rate after transfection procedures, which do not need to disturb nuclear envelope. In addition, mRNA-based vectors offer a simple combination of various transgenes through mixing of several mRNAs in a single multi-gene cocktail or expression of a number of proteins from a single mRNA molecule using internal ribosome entry sites (IRESes), ribosome skipping sequences and proteolytic signals. However, on the downside, uncontrolled extrac‐ ellular and intracellular decay of mRNA can be a substantial hurdle for mRNA- mediated gene transfer. Procedures for mRNA delivery are analogous to DNA transfer methods, which are well-established. In general, there are three actors in the gene delivery play, namely, the vector, the cell and the transfer environment. -

Nucleofection Protocol
NUCLEOFECTION PROTOCOL Nucleofection (electroporation) of Cas9/ synthetic RNA ribonucleoprotein (RNP) complexes for CRISPR/Cas9 genome editing (Lonza Nucleofection™ System) BACKGROUND This protocol describes how to transfect cultured cells with ribonucleoprotein (RNP) complexes that consist of purified Cas9 nuclease duplexed with synthetic guide RNA (gRNA; synthetic sgRNA or annealed crRNA and tracrRNA) using the Lonza 4D Nucleofector™ unit with 16-well Nucleocuvette™ Strips. Delivery of RNPs means that CRISPR components exist only transiently inside the cell, limiting Cas9 and guide RNA expression – this allows for the highest levels of editing efficiency and greatly reduces the chances of possible off-target and toxic effects. Furthermore, the use of synthetic guide RNAs eliminates the risk of incorporating foreign DNA into the host genome, which can occur when using plasmid-based guides. Synthego synthetic guide RNAs are of the highest quality and offer a superior alternative to in vitro transcribed (IVT) guide RNAs that are of variable quality and produce inconsistent editing results. MATERIALS REQUIRED Reagent/Material Vendor Synthego CRISPRevolution synthetic guide RNA Synthego (sgRNA or crRNA/tracrRNA) Synthego 2NLS-Cas9 nuclease Synthego Lonza Nucleofector 4D Electroporation System Lonza (#AAF-1002B and variants) Lonza SF Cell Line 4D - Nucleofector™ X Kit S Lonza (#V4XC-2032) with 16-well Nucleocuvette™ Strip DMEM, high glucose, with GlutaMAX™ Thermo Fisher (#10566 and variants) Fetal Bovine Serum (FBS) Thermo Fisher (#10437010 and variants) TrypLE™ Express Enzyme Thermo Fisher (#12605010 and variants) 1X PBS (without Ca²⁺ and Mg²⁺) Common lab supplier, or make in house Sterile tissue culture plates (24-well) Common lab supplier Microcentrifuge tubes Common lab supplier 1 synthego.com | [email protected] 20161208 NUCLEOFECTION PROTOCOL IMPORTANT CONSIDERATIONS • Wearing gloves and using nuclease-free tubes and reagents is recommended in order to avoid RNase contamination. -

Physical Non-Viral Gene Delivery Methods for Tissue Engineering
HHS Public Access Author manuscript Author ManuscriptAuthor Manuscript Author Ann Biomed Manuscript Author Eng. Author Manuscript Author manuscript; available in PMC 2016 November 09. Published in final edited form as: Ann Biomed Eng. 2013 March ; 41(3): 446–468. doi:10.1007/s10439-012-0678-1. Physical non-viral gene delivery methods for tissue engineering Adam J. Mellott, B.S.1, M. Laird Forrest, Ph.D.2, and Michael S. Detamore, Ph.D.1,3 1Bioengineering Program, University of Kansas, Lawrence, Kansas 66045 2Department of Pharmaceutical Chemistry, University of Kansas, Lawrence, Kansas 66047 3Department of Chemical and Petroleum Engineering, University of Kansas, Lawrence, Kansas 66045 Abstract The integration of gene therapy into tissue engineering to control differentiation and direct tissue formation is not a new concept; however, successful delivery of nucleic acids into primary cells, progenitor cells, and stem cells has proven exceptionally challenging. Viral vectors are generally highly effective at delivering nucleic acids to a variety of cell populations, both dividing and non- dividing, yet these viral vectors are marred by significant safety concerns. Non-viral vectors are preferred for gene therapy, despite lower transfection efficiencies, and possess many customizable attributes that are desirable for tissue engineering applications. However, there is no single non- viral gene delivery strategy that “fits-all” cell types and tissues. Thus, there is a compelling opportunity to examine different non-viral vectors, especially physical vectors, and compare their relative degrees of success. This review examines the advantages and disadvantages of physical non-viral methods (i.e., microinjection, ballistic gene delivery, electroporation, sonoporation, laser irradiation, magnetofection, and electric field-induced molecular vibration), with particular attention given to electroporation because of its versatility, with further special emphasis on Nucleofection™. -

Designing an Rnai Experiment Using Nucleofection™ Technical Reference Guide
BioResearch Designing an RNAi Experiment Using Nucleofection™ Technical Reference Guide The Nucleofector™ Technology is well suited for the transfection of siRNA duplexes or shRNA vectors into both primary cells and difficult-to-transfect cell lines. Choose siRNA Choose Cell Type and Transfection Protocol – Select gene target(s) – Select cell type(s) to maximize physiological relevance – Select control siRNA of results – Negative control siRNA – – Find Optimized Nucleofection™ Protocols at e.g., Thermo Scientific siGENOME Non-Targeting Control www.lonza.com/cell-database – Positive control siRNA – – Select transfection controls e.g., siGENOME® GAPDH siRNA – Untreated sample (no siRNA and transfection) – Mock-transfection (no siRNA, only transfection) ➔ ➔ Confirm siRNA Delivery Choose Detection Assay – Confirm siRNA delivery efficiency using: – Select detection assay(s) – Fluorescently-labeled siRNA – mRNA – branched-DNA, RT-PCR or – Protein – ELISA, Western, FACS analysis – Fluorescent expression plasmid (e.g., pmaxGFP™ Vector) – Phenotype – viability, apoptosis or – pmaxGFP™ Vector and maxGFP™ Reporter Protein siRNA or siRNA targeting housekeeping gene ➔ ➔ Optimize Target Knockdown Adapt Assay Conditions – Determine optimal siRNA concentration – Optimize detection assay(s) conditions for – Nucleofector™ Device ➞ 0.2 – 200 pmol (2 nM – 2 µM) specific system ➞ – 96-well Shuttle™ Device 0.04 – 40 pmol (2 nM – 2 µM) – Determine optimal cell densities for linear detection range – Correlate results from multiple assays ➔ ➔ Optimize Assay Conditions -

An Efficient Transfection Method for Mouse Embryonic Stem Cells
Gene Therapy (2009) 16, 154–158 & 2009 Macmillan Publishers Limited All rights reserved 0969-7128/09 $32.00 www.nature.com/gt SHORT COMMUNICATION An efficient transfection method for mouse embryonic stem cells BS Ko1,2,5, TC Chang1,5, SK Shyue3, YC Chen4 and JY Liou1 1National Health Research Institutes, Zhunan Town, Miaoli County, Taiwan; 2Department of Internal Medicine, National Taiwan University Hospital, Taipei, Taiwan; 3Institutes of Biomedical Science, Academic Sinica, Taipei, Taiwan and 4Department of Laboratory Medicine, National Taiwan University Hospital, Taipei, Taiwan Embryonic stem (ES) cells are considered to have potentials Our results show that transfection by Effectene achieves the for tissue regeneration and treatment of diverse human efficiency of 498% in CCE and 480% in D3 cells. The diseases. ES cells are capable of indefinite renewal and optimal ratio of DNA:Effectene for EGFP transfection is proliferation, which can be induced to differentiate into tissues between 1:4 and 1:8. Transient-expressed EGFP or endo- of all three germ lines. Despite these exciting potential, it genous protein kinase A (PKA) were significantly knocked remains unclear as to how the renewal and differentiation down by Effectene transfection of specific siRNA. High EGFP programs are operated and regulated at the genetic level. level expression and accumulation in mES cells induces Genetic manipulation such as delivery of exogenous gene minor cytotoxicity but can be reduced by introducing siRNA of expression or knockdown with small interfering RNA (siRNA) EGFP. Further, this transfection method did not significantly is commonly used in most of cancer or transformed cells but affect mES properties of proliferation or differentiation. -

RNA Pull-Down Procedure to Identify RNA Targets of a Long Non-Coding
RNA Pull-down Procedure to Identify RNA Targets of a Long Non-coding RNA Manon Torres, Denis Becquet, Séverine Guillen, Bénédicte Boyer, Mathias Moreno, Marie-Pierre Blanchard, Jean-Louis Franc, Anne-Marie François-Bellan To cite this version: Manon Torres, Denis Becquet, Séverine Guillen, Bénédicte Boyer, Mathias Moreno, et al.. RNA Pull-down Procedure to Identify RNA Targets of a Long Non-coding RNA. Journal of visualized experiments : JoVE, JoVE, 2018, 10.3791/57379. hal-02093065 HAL Id: hal-02093065 https://hal-amu.archives-ouvertes.fr/hal-02093065 Submitted on 20 Dec 2019 HAL is a multi-disciplinary open access L’archive ouverte pluridisciplinaire HAL, est archive for the deposit and dissemination of sci- destinée au dépôt et à la diffusion de documents entific research documents, whether they are pub- scientifiques de niveau recherche, publiés ou non, lished or not. The documents may come from émanant des établissements d’enseignement et de teaching and research institutions in France or recherche français ou étrangers, des laboratoires abroad, or from public or private research centers. publics ou privés. Distributed under a Creative Commons Attribution - NonCommercial - NoDerivatives| 4.0 International License Journal of Visualized Experiments www.jove.com Video Article RNA Pull-down Procedure to Identify RNA Targets of a Long Non-coding RNA Manon Torres1, Denis Becquet1, Séverine Guillen1, Bénédicte Boyer1, Mathias Moreno1, Marie-Pierre Blanchard2, Jean-Louis Franc1, Anne- Marie François-Bellan1 1 CNRS, CRN2M-UMR7286, Faculté -

Advances in Oligonucleotide Drug Delivery
REVIEWS Advances in oligonucleotide drug delivery Thomas C. Roberts 1,2 ✉ , Robert Langer 3 and Matthew J. A. Wood 1,2 ✉ Abstract | Oligonucleotides can be used to modulate gene expression via a range of processes including RNAi, target degradation by RNase H-mediated cleavage, splicing modulation, non-coding RNA inhibition, gene activation and programmed gene editing. As such, these molecules have potential therapeutic applications for myriad indications, with several oligonucleotide drugs recently gaining approval. However, despite recent technological advances, achieving efficient oligonucleotide delivery, particularly to extrahepatic tissues, remains a major translational limitation. Here, we provide an overview of oligonucleotide-based drug platforms, focusing on key approaches — including chemical modification, bioconjugation and the use of nanocarriers — which aim to address the delivery challenge. Oligonucleotides are nucleic acid polymers with the In addition to their ability to recognize specific tar- potential to treat or manage a wide range of diseases. get sequences via complementary base pairing, nucleic Although the majority of oligonucleotide therapeutics acids can also interact with proteins through the for- have focused on gene silencing, other strategies are being mation of three-dimensional secondary structures — a pursued, including splice modulation and gene activa- property that is also being exploited therapeutically. For tion, expanding the range of possible targets beyond example, nucleic acid aptamers are structured -

Guide for Morpholino Users: Toward Therapeutics
Open Access Journal of Drug Discovery, Development and Delivery Special Article - Antisense Drug Research and Development Guide for Morpholino Users: Toward Therapeutics Moulton JD* Gene Tools, LLC, USA Abstract *Corresponding author: Moulton JD, Gene Tools, Morpholino oligos are uncharged molecules for blocking sites on RNA. They LLC, 1001 Summerton Way, Philomath, Oregon 97370, are specific, soluble, non-toxic, stable, and effective antisense reagents suitable USA for development as therapeutics and currently in clinical trials. They are very versatile, targeting a wide range of RNA targets for outcomes such as blocking Received: January 28, 2016; Accepted: April 29, 2016; translation, modifying splicing of pre-mRNA, inhibiting miRNA maturation and Published: May 03, 2016 activity, as well as less common biological targets and diagnostic applications. Solutions have been developed for delivery into a range of cultured cells, embryos and adult animals; with development of a non-toxic and effective system for systemic delivery, Morpholinos have potential for broad therapeutic development targeting pathogens and genetic disorders. Keywords: Splicing; Duchenne muscular dystrophy; Phosphorodiamidate morpholino oligos; Internal ribosome entry site; Nonsense-mediated decay Morpholinos: Research Applications, the transcript from miRNA regulation; Therapeutic Promise • Block regulatory proteins from binding to RNA, shifting Morpholino oligos bind to complementary sequences of RNA alternative splicing; and get in the way of processes. Morpholino oligos are commonly • Block association of RNAs with cytoskeletal motor protein used to prevent a particular protein from being made in an organism complexes, preventing RNA translocation; or cell culture. Morpholinos are not the only tool used for this: a protein’s synthesis can be inhibited by altering DNA to make a null • Inhibit poly-A tailing of pre-mRNA; mutant (called a gene knockout) or by interrupting processes on RNA • Trigger frame shifts at slippery sequences; (called a gene knockdown).