Detecting Differential Copy Number Variation Between Groups of Samples
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Supplementary Table 1: Adhesion Genes Data Set
Supplementary Table 1: Adhesion genes data set PROBE Entrez Gene ID Celera Gene ID Gene_Symbol Gene_Name 160832 1 hCG201364.3 A1BG alpha-1-B glycoprotein 223658 1 hCG201364.3 A1BG alpha-1-B glycoprotein 212988 102 hCG40040.3 ADAM10 ADAM metallopeptidase domain 10 133411 4185 hCG28232.2 ADAM11 ADAM metallopeptidase domain 11 110695 8038 hCG40937.4 ADAM12 ADAM metallopeptidase domain 12 (meltrin alpha) 195222 8038 hCG40937.4 ADAM12 ADAM metallopeptidase domain 12 (meltrin alpha) 165344 8751 hCG20021.3 ADAM15 ADAM metallopeptidase domain 15 (metargidin) 189065 6868 null ADAM17 ADAM metallopeptidase domain 17 (tumor necrosis factor, alpha, converting enzyme) 108119 8728 hCG15398.4 ADAM19 ADAM metallopeptidase domain 19 (meltrin beta) 117763 8748 hCG20675.3 ADAM20 ADAM metallopeptidase domain 20 126448 8747 hCG1785634.2 ADAM21 ADAM metallopeptidase domain 21 208981 8747 hCG1785634.2|hCG2042897 ADAM21 ADAM metallopeptidase domain 21 180903 53616 hCG17212.4 ADAM22 ADAM metallopeptidase domain 22 177272 8745 hCG1811623.1 ADAM23 ADAM metallopeptidase domain 23 102384 10863 hCG1818505.1 ADAM28 ADAM metallopeptidase domain 28 119968 11086 hCG1786734.2 ADAM29 ADAM metallopeptidase domain 29 205542 11085 hCG1997196.1 ADAM30 ADAM metallopeptidase domain 30 148417 80332 hCG39255.4 ADAM33 ADAM metallopeptidase domain 33 140492 8756 hCG1789002.2 ADAM7 ADAM metallopeptidase domain 7 122603 101 hCG1816947.1 ADAM8 ADAM metallopeptidase domain 8 183965 8754 hCG1996391 ADAM9 ADAM metallopeptidase domain 9 (meltrin gamma) 129974 27299 hCG15447.3 ADAMDEC1 ADAM-like, -

Identification of Sequence Variants
Wang et al. BMC Medical Genomics (2019) 12:28 https://doi.org/10.1186/s12920-019-0475-x RESEARCH ARTICLE Open Access Identification of sequence variants associated with severe microtia-astresia by targeted sequencing Pu Wang1, Yibei Wang1, Xinmiao Fan1, Yaping Liu2, Yue Fan1, Tao Liu3, Chongjian Chen3, Shuyang Zhang4*† and Xiaowei Chen1*† Abstract Background: Microtia-atresia is characterized by abnormalities of the auricle (microtia) and aplasia or hypoplasia of the external auditory canal, often associated with middle ear abnormalities. To date, no causal genetic mutations or genes have been identified in microtia-atresia patients. Methods: We designed a panel of 131 genes associated with external/middle or inner ear deformity. Targeted genomic capturing combined with next-generation sequencing (NGS) was utilized to screen for mutations in 40 severe microtia-atresia patients. Mutations detected by NGS were filtered and validated. And then mutations were divided into three categories—rare or novel variants, low-frequency variants and common variants—based on their frequency in the public database. The rare or novel mutations were prioritized by pathogenicity analysis. For the low-frequency variants and common variants, we used association studies to explore risk factors of severe microtia-atresia. Results: Sixty-five rare heterozygous mutations of 42 genes were identified in 27 (67.5%) severe microtia-atresia patients. Association studies to determine genes that were potentially pathogenic found that PLEC, USH2A, FREM2, DCHS1, GLI3, POMT1 and GBA genes were significantly associated with severe microtia-atresia. Of these, DCHS1 was strongly suggested to cause severe microtia-atresia as it was identified by both low-frequency and common variants association studies. -

Mutations in DCHS1 Cause Mitral Valve Prolapse Ronen Durst, Kimberly Sauls, David S
Mutations in DCHS1 cause mitral valve prolapse Ronen Durst, Kimberly Sauls, David S. Peal, Annemarieke Devlaming, Katelynn Toomer, Maire Leyne, Monica Salani, Michael E. Talkowski, Harrison Brand, Maëlle Perrocheau, et al. To cite this version: Ronen Durst, Kimberly Sauls, David S. Peal, Annemarieke Devlaming, Katelynn Toomer, et al.. Mutations in DCHS1 cause mitral valve prolapse. Nature, Nature Publishing Group, 2015, Equipe 3, 525 (7567), pp.109–113. 10.1038/nature14670. hal-01830584 HAL Id: hal-01830584 https://hal.archives-ouvertes.fr/hal-01830584 Submitted on 12 Jul 2018 HAL is a multi-disciplinary open access L’archive ouverte pluridisciplinaire HAL, est archive for the deposit and dissemination of sci- destinée au dépôt et à la diffusion de documents entific research documents, whether they are pub- scientifiques de niveau recherche, publiés ou non, lished or not. The documents may come from émanant des établissements d’enseignement et de teaching and research institutions in France or recherche français ou étrangers, des laboratoires abroad, or from public or private research centers. publics ou privés. HHS Public Access Author manuscript Author Manuscript Author ManuscriptNature. Author ManuscriptAuthor manuscript; Author Manuscript available in PMC 2016 March 03. Published in final edited form as: Nature. 2015 September 3; 525(7567): 109–113. doi:10.1038/nature14670. Mutations in DCHS1 Cause Mitral Valve Prolapse Ronen Durst1,2,*, Kimberly Sauls5,*, David S Peal3,*, Annemarieke deVlaming5, Katelynn Toomer5, Maire Leyne1, Monica Salani1, Michael E. Talkowski1, Harrison Brand1, Maëlle Perrocheau9, Charles Simpson1, Christopher Jett1, Matthew R. Stone1, Florie Charles1, Colby Chiang1, Stacey N. Lynch3, Nabila Bouatia-Naji9,10, Francesca N. Delling7, Lisa A. -

Peripheral Nerve Single-Cell Analysis Identifies Mesenchymal Ligands That Promote Axonal Growth
Research Article: New Research Development Peripheral Nerve Single-Cell Analysis Identifies Mesenchymal Ligands that Promote Axonal Growth Jeremy S. Toma,1 Konstantina Karamboulas,1,ª Matthew J. Carr,1,2,ª Adelaida Kolaj,1,3 Scott A. Yuzwa,1 Neemat Mahmud,1,3 Mekayla A. Storer,1 David R. Kaplan,1,2,4 and Freda D. Miller1,2,3,4 https://doi.org/10.1523/ENEURO.0066-20.2020 1Program in Neurosciences and Mental Health, Hospital for Sick Children, 555 University Avenue, Toronto, Ontario M5G 1X8, Canada, 2Institute of Medical Sciences University of Toronto, Toronto, Ontario M5G 1A8, Canada, 3Department of Physiology, University of Toronto, Toronto, Ontario M5G 1A8, Canada, and 4Department of Molecular Genetics, University of Toronto, Toronto, Ontario M5G 1A8, Canada Abstract Peripheral nerves provide a supportive growth environment for developing and regenerating axons and are es- sential for maintenance and repair of many non-neural tissues. This capacity has largely been ascribed to paracrine factors secreted by nerve-resident Schwann cells. Here, we used single-cell transcriptional profiling to identify ligands made by different injured rodent nerve cell types and have combined this with cell-surface mass spectrometry to computationally model potential paracrine interactions with peripheral neurons. These analyses show that peripheral nerves make many ligands predicted to act on peripheral and CNS neurons, in- cluding known and previously uncharacterized ligands. While Schwann cells are an important ligand source within injured nerves, more than half of the predicted ligands are made by nerve-resident mesenchymal cells, including the endoneurial cells most closely associated with peripheral axons. At least three of these mesen- chymal ligands, ANGPT1, CCL11, and VEGFC, promote growth when locally applied on sympathetic axons. -

Using an Atlas of Gene Regulation Across 44 Human Tissues to Inform Complex Disease- and Trait-Associated Variation
Using an atlas of gene regulation across 44 human tissues to inform complex disease- and trait-associated variation The MIT Faculty has made this article openly available. Please share how this access benefits you. Your story matters. Citation Gamazon, Eric R. "Using an atlas of gene regulation across 44 human tissues to inform complex disease- and trait-associated variation." Nature Genetics 50, 7 (July 2018): 956–967 © 2018 The Author(s) As Published http://dx.doi.org/10.1038/S41588-018-0154-4 Publisher Springer Nature Version Author's final manuscript Citable link https://hdl.handle.net/1721.1/121227 Terms of Use Creative Commons Attribution-Noncommercial-Share Alike Detailed Terms http://creativecommons.org/licenses/by-nc-sa/4.0/ Europe PMC Funders Group Author Manuscript Nat Genet. Author manuscript; available in PMC 2018 December 28. Published in final edited form as: Nat Genet. 2018 July ; 50(7): 956–967. doi:10.1038/s41588-018-0154-4. Europe PMC Funders Author Manuscripts Using an atlas of gene regulation across 44 human tissues to inform complex disease- and trait-associated variation Eric R. Gamazon1,2,§,*, Ayellet V. Segrè3,4,§,*, Martijn van de Bunt5,6,§, Xiaoquan Wen7, Hualin S. Xi8, Farhad Hormozdiari9,10, Halit Ongen11,12,13, Anuar Konkashbaev1, Eske M. Derks14, François Aguet3, Jie Quan8, GTEx Consortium15, Dan L. Nicolae16,17,18, Eleazar Eskin9, Manolis Kellis3,19, Gad Getz3,20, Mark I. McCarthy5,6, Emmanouil T. Dermitzakis11,12,13, Nancy J. Cox1, and Kristin G. Ardlie3 1Division of Genetic Medicine, Department of Medicine, -
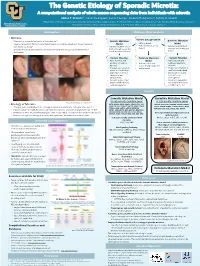
Example of a Scientific Poster
The Genetic Etiology of Sporadic Microtia: A computational analysis of whole exome sequencing data from individuals with microtia Abbas T. Shaikh1,*, Iván E. Rodríguez2, Connor Lester3, Frederic Deleyiannis4, Tamim H. Shaikh1 1Department of Pediatrics, University of Colorado, Anschutz Medical Campus, Aurora, CO; 2Department of Surgery, University of Colorado, Anschutz Medical Campus, Aurora, CO; University of Colorado Denver 3 4 Anschutz Medical Campus Georgetown University School of Medicine, Washington, D.C.; UCHealth Medical Group, Colorado Springs, CO Introduction Methods: Data Analysis • Microtia Variant Categorization • Microtia is a congenital deformity of the outer ear[1] Somatic Mutation Germline Mutation • • Severity ranges from mild structural deformations to complete absence of the auricle (outer Model • SNVs or Indels Model • high, moderate, or low • ear), known as anotia[1] • Variants found in one or • high, moderate, or low • Variants found in both impact affected and unaffected • Sporadic microtia is an isolated occurrence of microtia with no associated syndromes or both affected tissues but affected and unaffected tissues deformities not in unaffected tissues Variant Filtering Common Mutation Variant Filtering • Gene function and Model • Gene function and pathway relevant to • Variants consistent pathway relevant to microtia across multiple patients microtia • Frequency of variant in with microtia • Frequency of variant in unaffected individuals unaffected individuals • Expression in tissues • Expression in tissues relevant to ear relevant to ear development development • Variants in genes that • Variants in genes that matched with microtia matched with microtia candidate gene list candidate gene list (204) Results Somatic Mutation Model Germline Mutation Model 37 high-priority candidate genes: 18 high-priority candidate genes: Image taken from Luquetti et al. -

Characterization of Rare Germline Variants in Familial Multiple Myeloma
Catalano et al. Blood Cancer Journal (2021) 11:33 https://doi.org/10.1038/s41408-021-00422-6 Blood Cancer Journal CORRESPONDENCE Open Access Characterization of rare germline variants in familial multiple myeloma Calogerina Catalano1,2, Nagarajan Paramasivam3,JoannaBlocka2, Sara Giangiobbe1,2,StefanieHuhn2,4, Matthias Schlesner 5, Niels Weinhold2, Rolf Sijmons6, Mirjam de Jong6, Christian Langer7, Klaus-Dieter Preuss8, Björn Nilsson 9, Brian Durie10, Hartmut Goldschmidt 2,4, Obul Reddy Bandapalli1,11,12,KariHemminki1,13,14 and Asta Försti 1,11,12 Dear Editor, exome sequencing9. Altogether, 21 families with 46 Multiple myeloma (MM) is a malignancy of plasma affected and 20 unaffected family members were recruited cells, characterized by the presence of monoclonal (Supplementary Information, Supplementary Fig. 1). Each immunoglobulin, known as M protein1. MM is preceded family had at least two individuals diagnosed with MM or by monoclonal gammopathy of undetermined significance its precursors MGUS and smoldering MM (SMM). After (MGUS) which is also a precursor of immunoglobulin sequencing, detailed bioinformatics analyses using our in- light chain (AL) amyloidosis1. Previous studies have house developed Familial Cancer Variant Prioritization reported a 2- to 4-fold increased risk of MGUS or MM in Pipeline version-2 (FCVPPv2) were conducted to prior- first-degree relatives of MM or MGUS patients, suggest- itize the most likely candidates (Fig. 1 and Supplementary ing the existence of inherited susceptibility2,3. For many Methods). Gradual filtering of variants after sequencing is years, high-risk germline predisposing genes have been shown for each family in Supplementary Table 1. The lacking for MM. However, recent sequencing efforts have functions of the gene products were collected from the 1234567890():,; 1234567890():,; 1234567890():,; 1234567890():,; proposed a few novel candidates, most notably loss-of- UniProtKB database (https://www.uniprot.org/) and function (LoF) variants in the tumor suppressor gene literature search. -

Molecular Network Pathway of ERBB in Diffuse-Type Gastric Cancer
Research Article iMedPub Journals Journal of Clinical Epigenetics 2018 www.imedpub.com ISSN 2472-1158 Vol.4 No.2:13 DOI: 10.21767/2472-1158.100098 Molecular Network Pathway of Shihori Tanabe1*, Kazuhiko Aoyagi2, ERBB In Diffuse-Type Gastric Cancer: Hiroshi Yokozaki3 and Mesenchymal Stem Cells and Epithelial- Hiroki Sasaki2 Mesenchymal Transition 1 Division of Risk Assessment, Biological Safety Research Center, National Institute of Health Sciences, Kawasaki, Japan Abstract 2 Department of Translational Oncology, Epithelial-mesenchymal transition (EMT) is related to malignancy and metastasis FIOC, National Cancer Center Research in cancer. The molecular networks including tyrosine kinases are altered in gastric Institute, Tokyo, Japan cancer (GC). This study aims to reveal the role of ERBBs (erb-b2 receptor tyrosine 3 Department of Pathology, Kobe kinases) in EMT, and generate the molecular network pathway of ERBBs in diffuse- University of Graduate School of type GC and mesenchymal stem cells (MSCs). The expression of ERBB genes was Medicine, Kobe, Japan analyzed in MSCs and diffuse-type GC which has mesenchymal characteristics compared to intestinal-type GC. The signaling and molecular network of ERBB was analyzed using several databases, including cBioPortal for Cancer Genomics, Kyoto *Corresponding author: Encyclopedia of Genes and Genomes (KEGG), BioGRID and VaProS. ERBB2 and Shihori Tanabe, PhD ERBB3 gene expression were up-regulated in diffuse-type GC compared to MSCs. The ERBB3 molecular network includes epidermal growth factor receptor (EGFR), [email protected] cadherin 1 (CDH1), catenin beta 1 (CTNNB1) and EPH receptor A5 (EPHA5). These results demonstrate the importance of the ERBB network in cancer signaling, and revealed a ERBB3 network pathway model in diffuse-type GC and MSCs, and EMT. -

Homo Sapiens, Homo Neanderthalensis and the Denisova Specimen: New Insights on Their Evolutionary Histories Using Whole-Genome Comparisons
Genetics and Molecular Biology, 35, 4 (suppl), 904-911 (2012) Copyright © 2012, Sociedade Brasileira de Genética. Printed in Brazil www.sbg.org.br Research Article Homo sapiens, Homo neanderthalensis and the Denisova specimen: New insights on their evolutionary histories using whole-genome comparisons Vanessa Rodrigues Paixão-Côrtes, Lucas Henrique Viscardi, Francisco Mauro Salzano, Tábita Hünemeier and Maria Cátira Bortolini Departamento de Genética, Instituto de Biociências, Universidade Federal do Rio Grande do Sul, Porto Alegre, RS, Brazil. Abstract After a brief review of the most recent findings in the study of human evolution, an extensive comparison of the com- plete genomes of our nearest relative, the chimpanzee (Pan troglodytes), of extant Homo sapiens, archaic Homo neanderthalensis and the Denisova specimen were made. The focus was on non-synonymous mutations, which consequently had an impact on protein levels and these changes were classified according to degree of effect. A to- tal of 10,447 non-synonymous substitutions were found in which the derived allele is fixed or nearly fixed in humans as compared to chimpanzee. Their most frequent location was on chromosome 21. Their presence was then searched in the two archaic genomes. Mutations in 381 genes would imply radical amino acid changes, with a frac- tion of these related to olfaction and other important physiological processes. Eight new alleles were identified in the Neanderthal and/or Denisova genetic pools. Four others, possibly affecting cognition, occured both in the sapiens and two other archaic genomes. The selective sweep that gave rise to Homo sapiens could, therefore, have initiated before the modern/archaic human divergence. -

Multiple Genomic Copy Number Variants Associated with Periventricular Nodular Heterotopia Indicate Extreme Genetic Heterogeneity
European Journal of Human Genetics (2019) 27:909–918 https://doi.org/10.1038/s41431-019-0335-3 ARTICLE Multiple genomic copy number variants associated with periventricular nodular heterotopia indicate extreme genetic heterogeneity 1 1 1 1 1 1 Elena Cellini ● Annalisa Vetro ● Valerio Conti ● Carla Marini ● Viola Doccini ● Claudia Clementella ● 1 2 2 3,4 Elena Parrini ● Sabrina Giglio ● Matteo Della Monica ● Marco Fichera ● 4 1,5 Sebastiano Antonino Musumeci ● Renzo Guerrini Received: 13 March 2018 / Accepted: 18 December 2018 / Published online: 25 January 2019 © European Society of Human Genetics 2019 Abstract Periventricular nodular heterotopia (PNH) is a brain malformation in which nodules of neurons are ectopically retained along the lateral ventricles. Genetic causes include FLNA abnormalities (classical X-linked PNH), rare variants in ARFGEF2, DCHS1, ERMARD, FAT4, INTS8, MAP1B, MCPH1, and NEDD4L, as well as several chromosomal abnormalities. We performed array-CGH in 106 patients with different malformations of cortical development (MCD) and looked for common 1234567890();,: 1234567890();,: pathways possibly involved in PNH. Forty-two patients, including two parent/proband couples, exhibited PNH associated or not with other brain abnormalities, 44 had polymicrogyria and 20 had rarer MCDs. We found an enrichment of either large rearrangements or cryptic copy number variants (CNVs) in PNH (15/42, 35.7%) vs polymicrogyria (4/44, 9.1%) (i.e., 5.6 times increased risk for PNH of carrying a pathogenic CNV). CNVs in seven genomic regions (2p11.2q12.1, 4p15, 14q11.2q12, 16p13.3, 19q13.33, 20q13.33, 22q11) represented novel, potentially causative, associations with PNH. Through in silico analysis of genes included in imbalances whose breakpoints were clearly detailed, we detected in 9/12 unrelated patients in our series and in 15/24 previously published patients, a significant (P < 0.05) overrepresentation of genes involved in vesicle-mediated transport. -

The Interaction Between Prpf8 and Dzip1 and Its Role in Mitral Valve Prolapse
Medical University of South Carolina MEDICA MUSC Theses and Dissertations 2019 The Interaction Between Prpf8 and Dzip1 and its Role in Mitral Valve Prolapse Janiece Sharmell Glover Medical University of South Carolina Follow this and additional works at: https://medica-musc.researchcommons.org/theses Recommended Citation Glover, Janiece Sharmell, "The Interaction Between Prpf8 and Dzip1 and its Role in Mitral Valve Prolapse" (2019). MUSC Theses and Dissertations. 210. https://medica-musc.researchcommons.org/theses/210 This Thesis is brought to you for free and open access by MEDICA. It has been accepted for inclusion in MUSC Theses and Dissertations by an authorized administrator of MEDICA. For more information, please contact [email protected]. The Interaction Between Prpf8 and Dzip1 and its Role in Mitral Valve Prolapse by Janiece Sharmell Glover A thesis submitted to the faculty of the Medical University of South Carolina in partial fulfillment of the requirements for the degree of Master of Science in Biomedical Sciences in the College of Graduate Studies. Department of Regenerative Medicine and Cell Biology 2019 Approved by: _____________________________________ Chairman, Russell (Chip) Norris _____________________________________ Laura Kasman _____________________________________ Antonis Kourtidis _____________________________________ Robin Muise-Helmericks Copyright Page Dedication This thesis is dedicated to my beloved family: my father, Fred, my mother, Marilyn and my sister Brittany, who fairly early on instilled in me that I could achieve anything I put my mind to! I would have never completed this journey without your constant support, love and prayers. Thank you for believing in me and feeding me words of encouragement in times of need. iii Acknowledgements First and foremost, I want to give all the honor and glory to my Lord and Savior, Jesus Christ. -

Roles of the Hippo Pathway in Lung Development and Tumorigenesis
IJC International Journal of Cancer Roles of the Hippo pathway in lung development and tumorigenesis Benjamin Yeung†, Jihang Yu† and Xiaolong Yang Department of Pathology and Molecular Medicine, Queen’s University, Kingston, Ontario, Canada Lung cancer is the most commonly diagnosed cancer and accounts for one fifth of all cancer deaths worldwide. Although sig- nificant progress has been made toward our understanding of the causes of lung cancer, the 5-year survival is still lower than 15%. Therefore, there is an urgent need for novel lung cancer biomarkers and drug targets. The Hippo signaling pathway is an emerging signaling pathway that regulates various biological processes. Recently, increasing evidence suggests that the Hippo pathway may play important roles in not only lung development but also lung tumorigenesis. In this review article, we will summarize the most recent advances and predict future directions on this new cancer research field. 5,7,16–25 According to the World Health Organization (2014), lung mammals. The core components of this pathway are Mini Review cancer is the most common cancer worldwide, killing 1.6 composed of two serine/threonine (S/T) kinases Mst1 and its million people annually. Surprisingly, despite the advance- homolog Mst2 (mammalian homolog of Drosophila Hippo) ment of surgical, radio- and chemotherapies over the years, and LATS1 and its homolog LATS2, two adaptor/scaffold the 5-year survival rate of lung cancer remains <15%.1 While proteins hMOB1 and WW45 and two WW domain- many tumor suppressor genes (p53, Rb, etc.), oncogenes (K- containing transcriptional co-activators TAZ and its paralog ras, EGFR, HER2, etc.) or signaling pathways (MAPK, PI3K, YAP (Fig.