Overexpression and Activation of Αvβ3 Integrin Differentially Affects Tgfβ2 Signaling in Human Trabecular Meshwork Cells
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Rare Variants in Dynein Heavy Chain Genes in Two Individuals with Situs
bioRxiv preprint doi: https://doi.org/10.1101/2020.03.30.011783; this version posted March 31, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC 4.0 International license. Rare variants in dynein heavy chain genes in two individuals with situs inversus and developmental dyslexia Andrea Bieder1,*, Elisabet Einarsdottir1,2,3, Hans Matsson4,5,6, Harriet E. Nilsson1,7, Jesper Eisfeldt5,8,9, Anca Dragomir10, Martin Paucar11, Tobias Granberg11,12, Tie-Qiang Li13, Anna Lindstrand5,8,14, Juha Kere1,2,15, Isabel Tapia-Páez16,* 1Department of Biosciences and Nutrition, Karolinska Institutet, Huddinge, Sweden 2Molecular Neurology Research Program, University of Helsinki, Helsinki, Finland and Folkhälsan Institute of Genetics, Helsinki, Finland 3Science for Life Laboratory, Department of Gene Technology, KTH-Royal Institute of Technology, Solna, Sweden 4Department of Women´s and Children´s Health, Karolinska Institutet, Solna, Sweden 5Center for Molecular Medicine, Karolinska Institutet, Stockholm, Sweden 6Department of Immunology, Genetics and Pathology, Uppsala University, Uppsala, Sweden 7Department of Biomedical Engineering and Health Systems, School of Engineering Sciences in Chemistry, Biotechnology and Health, KTH Royal Institute of Technology, Huddinge, Sweden 8Department of Molecular Medicine and Surgery, Karolinska Institutet, Stockholm, Sweden 9Science for Life Laboratory, -

LHX2 (NM 004789) Human Tagged ORF Clone Product Data
OriGene Technologies, Inc. 9620 Medical Center Drive, Ste 200 Rockville, MD 20850, US Phone: +1-888-267-4436 [email protected] EU: [email protected] CN: [email protected] Product datasheet for RC210786L4 LHX2 (NM_004789) Human Tagged ORF Clone Product data: Product Type: Expression Plasmids Product Name: LHX2 (NM_004789) Human Tagged ORF Clone Tag: mGFP Symbol: LHX2 Synonyms: hLhx2; LH2 Vector: pLenti-C-mGFP-P2A-Puro (PS100093) E. coli Selection: Chloramphenicol (34 ug/mL) Cell Selection: Puromycin ORF Nucleotide The ORF insert of this clone is exactly the same as(RC210786). Sequence: Restriction Sites: SgfI-MluI Cloning Scheme: ACCN: NM_004789 ORF Size: 1218 bp This product is to be used for laboratory only. Not for diagnostic or therapeutic use. View online » ©2021 OriGene Technologies, Inc., 9620 Medical Center Drive, Ste 200, Rockville, MD 20850, US 1 / 2 LHX2 (NM_004789) Human Tagged ORF Clone – RC210786L4 OTI Disclaimer: The molecular sequence of this clone aligns with the gene accession number as a point of reference only. However, individual transcript sequences of the same gene can differ through naturally occurring variations (e.g. polymorphisms), each with its own valid existence. This clone is substantially in agreement with the reference, but a complete review of all prevailing variants is recommended prior to use. More info OTI Annotation: This clone was engineered to express the complete ORF with an expression tag. Expression varies depending on the nature of the gene. RefSeq: NM_004789.3 RefSeq Size: 2416 bp RefSeq ORF: 1221 bp Locus ID: 9355 UniProt ID: P50458, B3KNJ5 Domains: homeobox, LIM Protein Families: Transcription Factors MW: 44.4 kDa Gene Summary: This gene encodes a protein belonging to a large protein family, members of which carry the LIM domain, a unique cysteine-rich zinc-binding domain. -

The Zinc-Finger Protein CNBP Is Required for Forebrain Formation In
Development 130, 1367-1379 1367 © 2003 The Company of Biologists Ltd doi:10.1242/dev.00349 The zinc-finger protein CNBP is required for forebrain formation in the mouse Wei Chen1,2, Yuqiong Liang1, Wenjie Deng1, Ken Shimizu1, Amir M. Ashique1,2, En Li3 and Yi-Ping Li1,2,* 1Department of Cytokine Biology, The Forsyth Institute, Boston, MA 02115, USA 2Harvard-Forsyth Department of Oral Biology, Harvard School of Dental Medicine, Boston, MA 02115, USA 3Cardiovascular Research Center, Massachusetts General Hospital, Department of Medicine, Harvard Medical School, Charlestown, MA 02129, USA *Author for correspondence (e-mail: [email protected]) Accepted 19 December 2002 SUMMARY Mouse mutants have allowed us to gain significant insight (AME), headfolds and forebrain. In Cnbp–/– embryos, the into axis development. However, much remains to be visceral endoderm remains in the distal tip of the conceptus learned about the cellular and molecular basis of early and the ADE fails to form, whereas the node and notochord forebrain patterning. We describe a lethal mutation mouse form normally. A substantial reduction in cell proliferation strain generated using promoter-trap mutagenesis. The was observed in the anterior regions of Cnbp–/– embryos at mutants exhibit severe forebrain truncation in homozygous gastrulation and neural-fold stages. In these regions, Myc mouse embryos and various craniofacial defects in expression was absent, indicating CNBP targets Myc in heterozygotes. We show that the defects are caused by rostral head formation. Our findings demonstrate that disruption of the gene encoding cellular nucleic acid Cnbp is essential for the forebrain induction and binding protein (CNBP); Cnbp transgenic mice were able specification. -

Sex-Specific Hippocampal 5-Hydroxymethylcytosine Is Disrupted in Response to Acute Stress Ligia A
University of Nebraska - Lincoln DigitalCommons@University of Nebraska - Lincoln Faculty Publications, Department of Statistics Statistics, Department of 2016 Sex-specific hippocampal 5-hydroxymethylcytosine is disrupted in response to acute stress Ligia A. Papale University of Wisconsin, [email protected] Sisi Li University of Wisconsin, [email protected] Andy Madrid University of Wisconsin, [email protected] Qi Zhang University of Nebraska-Lincoln, [email protected] Li Chen Emory University See next page for additional authors Follow this and additional works at: https://digitalcommons.unl.edu/statisticsfacpub Part of the Other Statistics and Probability Commons Papale, Ligia A.; Li, Sisi; Madrid, Andy; Zhang, Qi; Chen, Li; Chopra, Pankaj; Jin, Peng; Keles, Sunduz; and Alisch, Reid S., "Sex- specific hippocampal 5-hydroxymethylcytosine is disrupted in response to acute stress" (2016). Faculty Publications, Department of Statistics. 62. https://digitalcommons.unl.edu/statisticsfacpub/62 This Article is brought to you for free and open access by the Statistics, Department of at DigitalCommons@University of Nebraska - Lincoln. It has been accepted for inclusion in Faculty Publications, Department of Statistics by an authorized administrator of DigitalCommons@University of Nebraska - Lincoln. Authors Ligia A. Papale, Sisi Li, Andy Madrid, Qi Zhang, Li Chen, Pankaj Chopra, Peng Jin, Sunduz Keles, and Reid S. Alisch This article is available at DigitalCommons@University of Nebraska - Lincoln: https://digitalcommons.unl.edu/statisticsfacpub/62 Neurobiology of Disease 96 (2016) 54–66 Contents lists available at ScienceDirect Neurobiology of Disease journal homepage: www.elsevier.com/locate/ynbdi Sex-specific hippocampal 5-hydroxymethylcytosine is disrupted in response to acute stress Ligia A. Papale a,1,SisiLia,c,1, Andy Madrid a,c,QiZhangd,LiChene,PankajChoprae,PengJine, Sündüz Keleş b, Reid S. -

DNA·RNA Triple Helix Formation Can Function As a Cis-Acting Regulatory
DNA·RNA triple helix formation can function as a cis-acting regulatory mechanism at the human β-globin locus Zhuo Zhoua, Keith E. Gilesa,b,c, and Gary Felsenfelda,1 aLaboratory of Molecular Biology, National Institute of Diabetes and Digestive and Kidney Diseases, National Institutes of Health, Bethesda, MD 20892; bUniversity of Alabama at Birmingham Stem Cell Institute, University of Alabama at Birmingham, Birmingham, AL 35294; and cDepartment of Biochemistry and Molecular Genetics, University of Alabama at Birmingham, Birmingham, AL 35294 Contributed by Gary Felsenfeld, February 4, 2019 (sent for review January 4, 2019; reviewed by James Douglas Engel and Sergei M. Mirkin) We have identified regulatory mechanisms in which an RNA tran- of the criteria necessary to establish the presence of a triplex script forms a DNA duplex·RNA triple helix with a gene or one of its structure, we first describe and characterize triplex formation at regulatory elements, suggesting potential auto-regulatory mecha- the FAU gene in human erythroid K562 cells. FAU encodes a nisms in vivo. We describe an interaction at the human β-globin protein that is a fusion containing fubi, a ubiquitin-like protein, locus, in which an RNA segment embedded in the second intron of and ribosomal protein S30. Although fubi function is unknown, the β-globin gene forms a DNA·RNA triplex with the HS2 sequence posttranslational processing produces S30, a component of the within the β-globin locus control region, a major regulator of glo- 40S ribosome. We used this system to refine methods necessary bin expression. We show in human K562 cells that the triplex is to detect triplex formation and to distinguish it from R-loop stable in vivo. -

Meta-Analysis of Nasopharyngeal Carcinoma
BMC Genomics BioMed Central Research article Open Access Meta-analysis of nasopharyngeal carcinoma microarray data explores mechanism of EBV-regulated neoplastic transformation Xia Chen†1,2, Shuang Liang†1, WenLing Zheng1,3, ZhiJun Liao1, Tao Shang1 and WenLi Ma*1 Address: 1Institute of Genetic Engineering, Southern Medical University, Guangzhou, PR China, 2Xiangya Pingkuang associated hospital, Pingxiang, Jiangxi, PR China and 3Southern Genomics Research Center, Guangzhou, Guangdong, PR China Email: Xia Chen - [email protected]; Shuang Liang - [email protected]; WenLing Zheng - [email protected]; ZhiJun Liao - [email protected]; Tao Shang - [email protected]; WenLi Ma* - [email protected] * Corresponding author †Equal contributors Published: 7 July 2008 Received: 16 February 2008 Accepted: 7 July 2008 BMC Genomics 2008, 9:322 doi:10.1186/1471-2164-9-322 This article is available from: http://www.biomedcentral.com/1471-2164/9/322 © 2008 Chen et al; licensee BioMed Central Ltd. This is an Open Access article distributed under the terms of the Creative Commons Attribution License (http://creativecommons.org/licenses/by/2.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. Abstract Background: Epstein-Barr virus (EBV) presumably plays an important role in the pathogenesis of nasopharyngeal carcinoma (NPC), but the molecular mechanism of EBV-dependent neoplastic transformation is not well understood. The combination of bioinformatics with evidences from biological experiments paved a new way to gain more insights into the molecular mechanism of cancer. Results: We profiled gene expression using a meta-analysis approach. Two sets of meta-genes were obtained. Meta-A genes were identified by finding those commonly activated/deactivated upon EBV infection/reactivation. -

Ldb1 Is Essential for Development of Nkx2.1 Lineage Derived Gabaergic and Cholinergic Neurons in the Telencephalon
Developmental Biology 385 (2014) 94–106 Contents lists available at ScienceDirect Developmental Biology journal homepage: www.elsevier.com/locate/developmentalbiology Ldb1 is essential for development of Nkx2.1 lineage derived GABAergic and cholinergic neurons in the telencephalon Yangu Zhao a,n,1, Pierre Flandin b,2, Daniel Vogt b, Alexander Blood a, Edit Hermesz a,3, Heiner Westphal a, John L. R. Rubenstein b,nn a Program on Genomics of Differentiation, Eunice Kennedy Shriver National Institute of Child Health and Human Development, NIH, Bethesda, MD 20892, USA b Department of Psychiatry and the Nina Ireland Laboratory of Developmental Neurobiology, University of California San Francisco, San Francisco, CA 94143, USA article info abstract Article history: The progenitor zones of the embryonic mouse ventral telencephalon give rise to GABAergic and cholinergic Received 1 June 2013 neurons. We have shown previously that two LIM-homeodomain (LIM-HD) transcription factors, Lhx6 and Received in revised form Lhx8, that are downstream of Nkx2.1, are critical for the development of telencephalic GABAergic and 8 October 2013 cholinergic neurons. Here we investigate the role of Ldb1, a nuclear protein that binds directly to all LIM-HD Accepted 9 October 2013 factors, in the development of these ventral telencephalon derived neurons. We show that Ldb1 is expressed Available online 21 October 2013 in the Nkx2.1 cell lineage during embryonic development and in mature neurons. Conditional deletion of Ldb1 Keywords: causes defects in the expression of a series of genes in the ventral telencephalon and severe impairment in the Differentiation tangential migration of cortical interneurons from the ventral telencephalon. -

List of Genes Associated with Sudden Cardiac Death (Scdgseta) Gene
List of genes associated with sudden cardiac death (SCDgseta) mRNA expression in normal human heart Entrez_I Gene symbol Gene name Uniprot ID Uniprot name fromb D GTEx BioGPS SAGE c d e ATP-binding cassette subfamily B ABCB1 P08183 MDR1_HUMAN 5243 √ √ member 1 ATP-binding cassette subfamily C ABCC9 O60706 ABCC9_HUMAN 10060 √ √ member 9 ACE Angiotensin I–converting enzyme P12821 ACE_HUMAN 1636 √ √ ACE2 Angiotensin I–converting enzyme 2 Q9BYF1 ACE2_HUMAN 59272 √ √ Acetylcholinesterase (Cartwright ACHE P22303 ACES_HUMAN 43 √ √ blood group) ACTC1 Actin, alpha, cardiac muscle 1 P68032 ACTC_HUMAN 70 √ √ ACTN2 Actinin alpha 2 P35609 ACTN2_HUMAN 88 √ √ √ ACTN4 Actinin alpha 4 O43707 ACTN4_HUMAN 81 √ √ √ ADRA2B Adrenoceptor alpha 2B P18089 ADA2B_HUMAN 151 √ √ AGT Angiotensinogen P01019 ANGT_HUMAN 183 √ √ √ AGTR1 Angiotensin II receptor type 1 P30556 AGTR1_HUMAN 185 √ √ AGTR2 Angiotensin II receptor type 2 P50052 AGTR2_HUMAN 186 √ √ AKAP9 A-kinase anchoring protein 9 Q99996 AKAP9_HUMAN 10142 √ √ √ ANK2/ANKB/ANKYRI Ankyrin 2 Q01484 ANK2_HUMAN 287 √ √ √ N B ANKRD1 Ankyrin repeat domain 1 Q15327 ANKR1_HUMAN 27063 √ √ √ ANKRD9 Ankyrin repeat domain 9 Q96BM1 ANKR9_HUMAN 122416 √ √ ARHGAP24 Rho GTPase–activating protein 24 Q8N264 RHG24_HUMAN 83478 √ √ ATPase Na+/K+–transporting ATP1B1 P05026 AT1B1_HUMAN 481 √ √ √ subunit beta 1 ATPase sarcoplasmic/endoplasmic ATP2A2 P16615 AT2A2_HUMAN 488 √ √ √ reticulum Ca2+ transporting 2 AZIN1 Antizyme inhibitor 1 O14977 AZIN1_HUMAN 51582 √ √ √ UDP-GlcNAc: betaGal B3GNT7 beta-1,3-N-acetylglucosaminyltransfe Q8NFL0 -

Review Article Gas7 Is Required for Mesenchymal Stem Cell-Derived Bone Development
View metadata, citation and similar papers at core.ac.uk brought to you by CORE provided by Crossref Hindawi Publishing Corporation Stem Cells International Volume 2013, Article ID 137010, 6 pages http://dx.doi.org/10.1155/2013/137010 Review Article Gas7 Is Required for Mesenchymal Stem Cell-Derived Bone Development Chuck C.-K. Chao, Feng-Chun Hung, and Jack J. Chao Department of Biochemistry and Molecular Biology and Institute of Biomedical Sciences, College of Medicine, Chang Gung University, Taoyuan 333, Taiwan Correspondence should be addressed to Chuck C.-K. Chao; [email protected] Received 20 January 2013; Accepted 12 May 2013 Academic Editor: Gael¨ Y. Rochefort Copyright © 2013 Chuck C.-K. Chao et al. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. Mesenchymal stem cells (MSCs) can differentiate into osteoblasts and lead to bone formation in the body. Osteoblast differentiation and bone development are regulated by a network of molecular signals and transcription factors induced by several proteins, including BMP2, osterix, and Runx2. We recently observed that the growth-arrest-specific 7 gene (Gas7) is upregulated during differentiation of human MSCs into osteoblasts. Downregulation of Gas7 using short-hairpin RNA decreased the expression of Runx2, a master regulator of osteogenesis, and its target genes (alkaline phosphatase, type I collagen, osteocalcin, and osteopontin). In addition, knockdown of Gas7 decreased the mineralization of dexamethasone-treated MSCs in culture. Conversely, ectopic expression of Gas7 induced Runx2-dependent transcriptional activity and gene expression leading to osteoblast differentiation and matrix mineralization. -

Diet-Induced Obesity Mediated by the JNK/DIO2 Signal Transduction Pathway
Downloaded from genesdev.cshlp.org on September 24, 2021 - Published by Cold Spring Harbor Laboratory Press Diet-induced obesity mediated by the JNK/DIO2 signal transduction pathway Santiago Vernia,1 Julie Cavanagh-Kyros,1,2 Tamera Barrett,1,2 Dae Young Jung,1 Jason K. Kim,1,3 and Roger J. Davis1,2,4 1Program in Molecular Medicine, University of Massachusetts Medical School, Worcester, Massachusetts 01605, USA; 2Howard Hughes Medical Institute, Worcester, Massachusetts 01605, USA; 3Department of Medicine, Division of Endocrinology, Metabolism, and Diabetes, University of Massachusetts Medical School, Worcester, Massachusetts 01605, USA The cJun N-terminal kinase (JNK) signaling pathway is a key mediator of metabolic stress responses caused by consuming a high-fat diet, including the development of obesity. To test the role of JNK, we examined diet- induced obesity in mice with targeted ablation of Jnk genes in the anterior pituitary gland. These mice exhibited an increase in the pituitary expression of thyroid-stimulating hormone (TSH), an increase in the blood concentration of thyroid hormone (T4), increased energy expenditure, and markedly reduced obesity compared with control mice. The increased amount of pituitary TSH was caused by reduced expression of type 2 iodothyronine deiodinase (Dio2), a gene that is required for T4-mediated negative feedback regulation of TSH expression. These data establish a molecular mechanism that accounts for the regulation of energy expenditure and the development of obesity by the JNK signaling pathway. [Keywords: DIO2; JNK; obesity; pituitary gland; thyroid hormone] Supplemental material is available for this article. Received June 5, 2013; revised version accepted October 3, 2013. -
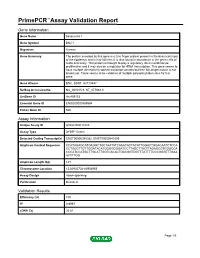
Primepcr™Assay Validation Report
PrimePCR™Assay Validation Report Gene Information Gene Name basonuclin 1 Gene Symbol BNC1 Organism Human Gene Summary The protein encoded by this gene is a zinc finger protein present in the basal cell layer of the epidermis and in hair follicles. It is also found in abundance in the germ cells of testis and ovary. This protein is thought to play a regulatory role in keratinocyte proliferation and it may also be a regulator for rRNA transcription. This gene seems to have multiple alternatively spliced transcript variants but their full-length nature is not known yet. There seems to be evidence of multiple polyadenylation sites for this gene. Gene Aliases BNC, BSN1, HsT19447 RefSeq Accession No. NC_000015.9, NT_077661.3 UniGene ID Hs.459153 Ensembl Gene ID ENSG00000169594 Entrez Gene ID 646 Assay Information Unique Assay ID qHsaCID0017223 Assay Type SYBR® Green Detected Coding Transcript(s) ENST00000345382, ENST00000541809 Amplicon Context Sequence CCATAGAGCATGAGGCTGCTAATATCAAACACTACATTGGACTGGACAATCTCCA CCTGGCTTGTTGGATACATGGGGGGGATCCTTAGCTTACTTAGAGCGTGGGCCA CCCATCCATGCTTGCATTGGTCACACTGACGGTGGTTTATTTTCCCGGGTTTGAA ACTTTGG Amplicon Length (bp) 141 Chromosome Location 15:83935724-83936955 Assay Design Intron-spanning Purification Desalted Validation Results Efficiency (%) 100 R2 0.9997 cDNA Cq 26.81 Page 1/5 PrimePCR™Assay Validation Report cDNA Tm (Celsius) 84.5 gDNA Cq 39.42 Specificity (%) 100 Information to assist with data interpretation is provided at the end of this report. Page 2/5 PrimePCR™Assay Validation Report BNC1, Human Amplification -

Supplementary Table 1: Adhesion Genes Data Set
Supplementary Table 1: Adhesion genes data set PROBE Entrez Gene ID Celera Gene ID Gene_Symbol Gene_Name 160832 1 hCG201364.3 A1BG alpha-1-B glycoprotein 223658 1 hCG201364.3 A1BG alpha-1-B glycoprotein 212988 102 hCG40040.3 ADAM10 ADAM metallopeptidase domain 10 133411 4185 hCG28232.2 ADAM11 ADAM metallopeptidase domain 11 110695 8038 hCG40937.4 ADAM12 ADAM metallopeptidase domain 12 (meltrin alpha) 195222 8038 hCG40937.4 ADAM12 ADAM metallopeptidase domain 12 (meltrin alpha) 165344 8751 hCG20021.3 ADAM15 ADAM metallopeptidase domain 15 (metargidin) 189065 6868 null ADAM17 ADAM metallopeptidase domain 17 (tumor necrosis factor, alpha, converting enzyme) 108119 8728 hCG15398.4 ADAM19 ADAM metallopeptidase domain 19 (meltrin beta) 117763 8748 hCG20675.3 ADAM20 ADAM metallopeptidase domain 20 126448 8747 hCG1785634.2 ADAM21 ADAM metallopeptidase domain 21 208981 8747 hCG1785634.2|hCG2042897 ADAM21 ADAM metallopeptidase domain 21 180903 53616 hCG17212.4 ADAM22 ADAM metallopeptidase domain 22 177272 8745 hCG1811623.1 ADAM23 ADAM metallopeptidase domain 23 102384 10863 hCG1818505.1 ADAM28 ADAM metallopeptidase domain 28 119968 11086 hCG1786734.2 ADAM29 ADAM metallopeptidase domain 29 205542 11085 hCG1997196.1 ADAM30 ADAM metallopeptidase domain 30 148417 80332 hCG39255.4 ADAM33 ADAM metallopeptidase domain 33 140492 8756 hCG1789002.2 ADAM7 ADAM metallopeptidase domain 7 122603 101 hCG1816947.1 ADAM8 ADAM metallopeptidase domain 8 183965 8754 hCG1996391 ADAM9 ADAM metallopeptidase domain 9 (meltrin gamma) 129974 27299 hCG15447.3 ADAMDEC1 ADAM-like,