Nucleomorph Genome
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

New Zealand's Genetic Diversity
1.13 NEW ZEALAND’S GENETIC DIVERSITY NEW ZEALAND’S GENETIC DIVERSITY Dennis P. Gordon National Institute of Water and Atmospheric Research, Private Bag 14901, Kilbirnie, Wellington 6022, New Zealand ABSTRACT: The known genetic diversity represented by the New Zealand biota is reviewed and summarised, largely based on a recently published New Zealand inventory of biodiversity. All kingdoms and eukaryote phyla are covered, updated to refl ect the latest phylogenetic view of Eukaryota. The total known biota comprises a nominal 57 406 species (c. 48 640 described). Subtraction of the 4889 naturalised-alien species gives a biota of 52 517 native species. A minimum (the status of a number of the unnamed species is uncertain) of 27 380 (52%) of these species are endemic (cf. 26% for Fungi, 38% for all marine species, 46% for marine Animalia, 68% for all Animalia, 78% for vascular plants and 91% for terrestrial Animalia). In passing, examples are given both of the roles of the major taxa in providing ecosystem services and of the use of genetic resources in the New Zealand economy. Key words: Animalia, Chromista, freshwater, Fungi, genetic diversity, marine, New Zealand, Prokaryota, Protozoa, terrestrial. INTRODUCTION Article 10b of the CBD calls for signatories to ‘Adopt The original brief for this chapter was to review New Zealand’s measures relating to the use of biological resources [i.e. genetic genetic resources. The OECD defi nition of genetic resources resources] to avoid or minimize adverse impacts on biological is ‘genetic material of plants, animals or micro-organisms of diversity [e.g. genetic diversity]’ (my parentheses). -

(1 → 4)-Β-D-Glucan Is a Component of Cell Walls in Brown Algae
Insoluble (13), (14)--D-glucan is a component of cell walls in brown algae (Phaeophyceae) and is masked by alginates in tissues Asunción Salmeán, Armando; Duffieux, Delphine; Harholt, Jesper; Qin, Fen; Michel, Gurvan; Czjzek, Mirjam; Willats, William George Tycho; Hervé, Cécile Published in: Scientific Reports DOI: 10.1038/s41598-017-03081-5 Publication date: 2017 Document version Publisher's PDF, also known as Version of record Citation for published version (APA): Asunción Salmeán, A., Duffieux, D., Harholt, J., Qin, F., Michel, G., Czjzek, M., Willats, W. G. T., & Hervé, C. (2017). Insoluble (13), (14)--D-glucan is a component of cell walls in brown algae (Phaeophyceae) and is masked by alginates in tissues. Scientific Reports, 7, [2880]. https://doi.org/10.1038/s41598-017-03081-5 Download date: 23. Sep. 2021 www.nature.com/scientificreports OPEN Insoluble (1 → 3), (1 → 4)-β-D- glucan is a component of cell walls in brown algae (Phaeophyceae) and Received: 28 October 2016 Accepted: 24 April 2017 is masked by alginates in tissues Published: xx xx xxxx Armando A. Salmeán 1, Delphine Duffieux2,3, Jesper Harholt4, Fen Qin4, Gurvan Michel2,3, Mirjam Czjzek2,3, William G. T. Willats1,5 & Cécile Hervé2,3 Brown algae are photosynthetic multicellular marine organisms. They belong to the phylum of Stramenopiles, which are not closely related to land plants and green algae. Brown algae share common evolutionary features with other photosynthetic and multicellular organisms, including a carbohydrate-rich cell-wall. Brown algal cell walls are composed predominantly of the polyanionic polysaccharides alginates and fucose-containing sulfated polysaccharides. These polymers are prevalent over neutral and crystalline components, which are believed to be mostly, if not exclusively, cellulose. -

The Apicoplast: a Review of the Derived Plastid of Apicomplexan Parasites
Curr. Issues Mol. Biol. 7: 57-80. Online journalThe Apicoplastat www.cimb.org 57 The Apicoplast: A Review of the Derived Plastid of Apicomplexan Parasites Ross F. Waller1 and Geoffrey I. McFadden2,* way to apicoplast discovery with studies of extra- chromosomal DNAs recovered from isopycnic density 1Botany, University of British Columbia, 3529-6270 gradient fractionation of total Plasmodium DNA. This University Boulevard, Vancouver, BC, V6T 1Z4, Canada group recovered two DNA forms; one a 6kb tandemly 2Plant Cell Biology Research Centre, Botany, University repeated element that was later identifed as the of Melbourne, 3010, Australia mitochondrial genome, and a second, 35kb circle that was supposed to represent the DNA circles previously observed by microscopists (Wilson et al., 1996b; Wilson Abstract and Williamson, 1997). This molecule was also thought The apicoplast is a plastid organelle, homologous to to be mitochondrial DNA, and early sequence data of chloroplasts of plants, that is found in apicomplexan eubacterial-like rRNA genes supported this organellar parasites such as the causative agents of Malaria conclusion. However, as the sequencing effort continued Plasmodium spp. It occurs throughout the Apicomplexa a new conclusion, that was originally embraced with and is an ancient feature of this group acquired by the some awkwardness (“Have malaria parasites three process of endosymbiosis. Like plant chloroplasts, genomes?”, Wilson et al., 1991), began to emerge. apicoplasts are semi-autonomous with their own genome Gradually, evermore convincing character traits of a and expression machinery. In addition, apicoplasts import plastid genome were uncovered, and strong parallels numerous proteins encoded by nuclear genes. These with plastid genomes from non-photosynthetic plants nuclear genes largely derive from the endosymbiont (Epifagus virginiana) and algae (Astasia longa) became through a process of intracellular gene relocation. -

The Taxonomy and Biology of Phytophthora and Pythium
Journal of Bacteriology & Mycology: Open Access Review Article Open Access The taxonomy and biology of Phytophthora and Pythium Abstract Volume 6 Issue 1 - 2018 The genera Phytophthora and Pythium include many economically important species Hon H Ho which have been placed in Kingdom Chromista or Kingdom Straminipila, distinct from Department of Biology, State University of New York, USA Kingdom Fungi. Their taxonomic problems, basic biology and economic importance have been reviewed. Morphologically, both genera are very similar in having coenocytic, hyaline Correspondence: Hon H Ho, Professor of Biology, State and freely branching mycelia, oogonia with usually single oospores but the definitive University of New York, New Paltz, NY 12561, USA, differentiation between them lies in the mode of zoospore differentiation and discharge. Email [email protected] In Phytophthora, the zoospores are differentiated within the sporangium proper and when mature, released in an evanescent vesicle at the sporangial apex, whereas in Pythium, the Received: January 23, 2018 | Published: February 12, 2018 protoplast of a sporangium is transferred usually through an exit tube to a thin vesicle outside the sporangium where zoospores are differentiated and released upon the rupture of the vesicle. Many species of Phytophthora are destructive pathogens of especially dicotyledonous woody trees, shrubs and herbaceous plants whereas Pythium species attacked primarily monocotyledonous herbaceous plants, whereas some cause diseases in fishes, red algae and mammals including humans. However, several mycoparasitic and entomopathogenic species of Pythium have been utilized respectively, to successfully control other plant pathogenic fungi and harmful insects including mosquitoes while the others utilized to produce valuable chemicals for pharmacy and food industry. -

Controlled Sampling of Ribosomally Active Protistan Diversity in Sediment-Surface Layers Identifies Putative Players in the Marine Carbon Sink
The ISME Journal (2020) 14:984–998 https://doi.org/10.1038/s41396-019-0581-y ARTICLE Controlled sampling of ribosomally active protistan diversity in sediment-surface layers identifies putative players in the marine carbon sink 1,2 1 1 3 3 Raquel Rodríguez-Martínez ● Guy Leonard ● David S. Milner ● Sebastian Sudek ● Mike Conway ● 1 1 4,5 6 7 Karen Moore ● Theresa Hudson ● Frédéric Mahé ● Patrick J. Keeling ● Alyson E. Santoro ● 3,8 1,9 Alexandra Z. Worden ● Thomas A. Richards Received: 6 October 2019 / Revised: 4 December 2019 / Accepted: 17 December 2019 / Published online: 9 January 2020 © The Author(s) 2020. This article is published with open access Abstract Marine sediments are one of the largest carbon reservoir on Earth, yet the microbial communities, especially the eukaryotes, that drive these ecosystems are poorly characterised. Here, we report implementation of a sampling system that enables injection of reagents into sediments at depth, allowing for preservation of RNA in situ. Using the RNA templates recovered, we investigate the ‘ribosomally active’ eukaryotic diversity present in sediments close to the water/sediment interface. We 1234567890();,: 1234567890();,: demonstrate that in situ preservation leads to recovery of a significantly altered community profile. Using SSU rRNA amplicon sequencing, we investigated the community structure in these environments, demonstrating a wide diversity and high relative abundance of stramenopiles and alveolates, specifically: Bacillariophyta (diatoms), labyrinthulomycetes and ciliates. The identification of abundant diatom rRNA molecules is consistent with microscopy-based studies, but demonstrates that these algae can also be exported to the sediment as active cells as opposed to dead forms. -

Horizontal Gene Transfer in Osmotrophs: Playing with Public Goods
ORE Open Research Exeter TITLE Horizontal gene transfer in osmotrophs: playing with public goods. AUTHORS Richards, Thomas A; Talbot, Nicholas J. JOURNAL Nat Rev Microbiol DEPOSITED IN ORE 19 November 2014 This version available at http://hdl.handle.net/10871/15898 COPYRIGHT AND REUSE Open Research Exeter makes this work available in accordance with publisher policies. A NOTE ON VERSIONS The version presented here may differ from the published version. If citing, you are advised to consult the published version for pagination, volume/issue and date of publication PERSPECTIVES however, not unique to fungi. Many bacteria, OPINION for instance, feed in an analogous man ner, and other eukaryotic groups, such as Horizontal gene transfer in hyphochytriomycetes (FIG. 1c) and oomycetes (FIG. 1d) (sometimes collectively termed the pseudofungi26), also feed osmotrophically osmotrophs: playing with public and adopt filamentous growth habits, allow ing invasive growth in heterogeneous sub goods strates. Importantly, these eukaryotes also lost the ability to carry out phagotrophy and Thomas A. Richards and Nicholas J. Talbot became obligately osmotrophic26,27. Osmotrophy has a number of distinct Abstract | Osmotrophic microorganisms, such as fungi and oomycetes, feed advantages as a feeding strategy. External by secreting depolymerizing enzymes to process complex food sources in the digestion of large and complex polymers extracellular environment, and taking up the resulting simple sugars, micronutrients allows greater control over substances that and amino acids. As a consequence of this lifestyle, osmotrophs engage in the are allowed to enter a cell (FIG. 2a), thus acquisition and protection of public goods. In this Opinion article, we propose that minimizing potential routes of infection and intake of harmful substances. -

Horizontal Gene Transfer
Horizontal Gene Transfer (HGT) from Fungi is the RESEARCH NEWS basis for plant pathogenicity in oomycetes It always seemed rather odd that some of which seem to have occurred close to that time. This more detailed study, made oomycetes (Oomycota) were so fungal-like the shift from phagotrophy to osmotrophy possible by increasingly available genome in their behaviour as plant pathogens. and the evolution of the fungal cell wall at sequences, reveals that this phenomenon Now whole-genome comparisons are the base of the monophyletic Fungi clade. is much more extensive than might have starting to reveal just why. Richards et al. Amongst the genes evidently transferred, been imagined. Thus, it is not a matter of (2011) have undertaken a painstaking are one related to features such as the ability oomycetes merely being ‘fungal analogues’ gene-by-gene analysis of the proteomes to break down plant cell walls and take or ‘pseudofungi’, they are actually partly of Hyaloperonospora parasitica and three up sugars, nitrogen and phosphates, and Fungi. species of Phytophthora (P. infestans, P. further ones implicated in overcoming plant It should be noted that HGT is not ramorum, and P. sojae) which reveals defence mechanisms and attacking plant only unidirectional from Fungi into to an extensive pattern of cross-kingdom cells. A schematic diagram of the functional other organisms. Indeed, in another paper horizontal gene transfer (HGT) from Fungi. proteome of oomycetes derived from published this summer, this same group of In the case of P. ramorum, an amazing 7.6 Fungi is provided (their Fig. 2) at which researchers report identifying 323 examples % of the secreted proteome appears to have one can only marvel at the complexity. -

Plastid-Targeting Peptides from the Chlorarachniophyte Bigelowiella Natans
J. Eukaryot. Microbiol., 51(5), 2004 pp. 529±535 q 2004 by the Society of Protozoologists Plastid-Targeting Peptides from the Chlorarachniophyte Bigelowiella natans MATTHEW B. ROGERS,a JOHN M. ARCHIBALD,a,1 MATTHEW A. FIELD,a CATHERINE LI,b BORIS STRIEPENb and PATRICK J. KEELINGa aCanadian Institute for Advanced Research, Department of Botany, University of British Columbia, 3529-6270 University Boulevard, Vancouver, BC, V6T 1Z4, Canada, and bCenter for Tropical & Emerging Global Diseases & Department of Cellular Biology, University of Georgia, 724 Biological Sciences Building, Athens, Georgia 30602, USA ABSTRACT. Chlorarachniophytes are marine amoebo¯agellate protists that have acquired their plastid (chloroplast) through secondary endosymbiosis with a green alga. Like other algae, most of the proteins necessary for plastid function are encoded in the nuclear genome of the secondary host. These proteins are targeted to the organelle using a bipartite leader sequence consisting of a signal peptide (allowing entry in to the endomembrane system) and a chloroplast transit peptide (for transport across the chloroplast envelope mem- branes). We have examined the leader sequences from 45 full-length predicted plastid-targeted proteins from the chlorarachniophyte Bigelowiella natans with the goal of understanding important features of these sequences and possible conserved motifs. The chemical characteristics of these sequences were compared with a set of 10 B. natans endomembrane-targeted proteins and 38 cytosolic or nuclear proteins, which show that the signal peptides are similar to those of most other eukaryotes, while the transit peptides differ from those of other algae in some characteristics. Consistent with this, the leader sequence from one B. -
Complete Nucleotide Sequence of the Chlorarachniophyte Nucleomorph: Nature’S Smallest Nucleus
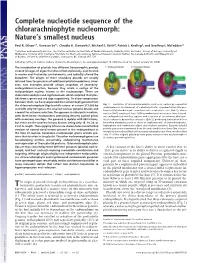
Complete nucleotide sequence of the chlorarachniophyte nucleomorph: Nature’s smallest nucleus Paul R. Gilson*†, Vanessa Su†‡, Claudio H. Slamovits§, Michael E. Reith¶, Patrick J. Keeling§, and Geoffrey I. McFadden‡ʈ *Infection and Immunity Division, The Walter and Eliza Hall Institute of Medical Research, Parkville 3050, Australia; ‡School of Botany, University of Melbourne, Victoria 3010, Australia; ¶Institute for Marine Biosciences, National Research Council, Halifax, NS, Canada B3H 3Z1; and §Department of Botany, University of British Columbia, Vancouver, BC, Canada V6T 1Z4 Edited by Jeffrey D. Palmer, Indiana University, Bloomington, IN, and approved April 19, 2006 (received for review January 26, 2006) The introduction of plastids into different heterotrophic protists created lineages of algae that diversified explosively, proliferated in marine and freshwater environments, and radically altered the biosphere. The origins of these secondary plastids are usually inferred from the presence of additional plastid membranes. How- ever, two examples provide unique snapshots of secondary- endosymbiosis-in-action, because they retain a vestige of the endosymbiont nucleus known as the nucleomorph. These are chlorarachniophytes and cryptomonads, which acquired their plas- tids from a green and red alga respectively. To allow comparisons between them, we have sequenced the nucleomorph genome from the chlorarachniophyte Bigelowiella natans: at a mere 373,000 bp Fig. 1. Evolution of chlorarachniophytes such as B. natans by sequential endosymbioses. Enslavement of a photosynthetic, cyanobacterium-like pro- and with only 331 genes, the smallest nuclear genome known and karyote (Cb) introduces photosynthesis into a eukaryotic host (Euk 1), whose a model for extreme reduction. The genome is eukaryotic in nature, nucleus (Nu1) acquires at least 1,000 cyanobacterial genes over time. -

Seven Gene Phylogeny of Heterokonts
ARTICLE IN PRESS Protist, Vol. 160, 191—204, May 2009 http://www.elsevier.de/protis Published online date 9 February 2009 ORIGINAL PAPER Seven Gene Phylogeny of Heterokonts Ingvild Riisberga,d,1, Russell J.S. Orrb,d,1, Ragnhild Klugeb,c,2, Kamran Shalchian-Tabrizid, Holly A. Bowerse, Vishwanath Patilb,c, Bente Edvardsena,d, and Kjetill S. Jakobsenb,d,3 aMarine Biology, Department of Biology, University of Oslo, P.O. Box 1066, Blindern, NO-0316 Oslo, Norway bCentre for Ecological and Evolutionary Synthesis (CEES),Department of Biology, University of Oslo, P.O. Box 1066, Blindern, NO-0316 Oslo, Norway cDepartment of Plant and Environmental Sciences, P.O. Box 5003, The Norwegian University of Life Sciences, N-1432, A˚ s, Norway dMicrobial Evolution Research Group (MERG), Department of Biology, University of Oslo, P.O. Box 1066, Blindern, NO-0316, Oslo, Norway eCenter of Marine Biotechnology, 701 East Pratt Street, Baltimore, MD 21202, USA Submitted May 23, 2008; Accepted November 15, 2008 Monitoring Editor: Mitchell L. Sogin Nucleotide ssu and lsu rDNA sequences of all major lineages of autotrophic (Ochrophyta) and heterotrophic (Bigyra and Pseudofungi) heterokonts were combined with amino acid sequences from four protein-coding genes (actin, b-tubulin, cox1 and hsp90) in a multigene approach for resolving the relationship between heterokont lineages. Applying these multigene data in Bayesian and maximum likelihood analyses improved the heterokont tree compared to previous rDNA analyses by placing all plastid-lacking heterotrophic heterokonts sister to Ochrophyta with robust support, and divided the heterotrophic heterokonts into the previously recognized phyla, Bigyra and Pseudofungi. Our trees identified the heterotrophic heterokonts Bicosoecida, Blastocystis and Labyrinthulida (Bigyra) as the earliest diverging lineages. -

Living Organisms Author Their Read-Write Genomes in Evolution
biology Review Living Organisms Author Their Read-Write Genomes in Evolution James A. Shapiro ID Department of Biochemistry and Molecular Biology, University of Chicago GCIS W123B, 979 E. 57th Street, Chicago, IL 60637, USA; [email protected]; Tel.: +1-773-702-1625 Academic Editor: Andrés Moya Received: 23 August 2017; Accepted: 28 November 2017; Published: 6 December 2017 Abstract: Evolutionary variations generating phenotypic adaptations and novel taxa resulted from complex cellular activities altering genome content and expression: (i) Symbiogenetic cell mergers producing the mitochondrion-bearing ancestor of eukaryotes and chloroplast-bearing ancestors of photosynthetic eukaryotes; (ii) interspecific hybridizations and genome doublings generating new species and adaptive radiations of higher plants and animals; and, (iii) interspecific horizontal DNA transfer encoding virtually all of the cellular functions between organisms and their viruses in all domains of life. Consequently, assuming that evolutionary processes occur in isolated genomes of individual species has become an unrealistic abstraction. Adaptive variations also involved natural genetic engineering of mobile DNA elements to rewire regulatory networks. In the most highly evolved organisms, biological complexity scales with “non-coding” DNA content more closely than with protein-coding capacity. Coincidentally, we have learned how so-called “non-coding” RNAs that are rich in repetitive mobile DNA sequences are key regulators of complex phenotypes. Both biotic and abiotic ecological challenges serve as triggers for episodes of elevated genome change. The intersections of cell activities, biosphere interactions, horizontal DNA transfers, and non-random Read-Write genome modifications by natural genetic engineering provide a rich molecular and biological foundation for understanding how ecological disruptions can stimulate productive, often abrupt, evolutionary transformations. -

The Miniaturized Nuclear Genome of a Eukaryotic Endosymbiont Contains
Proc. Natl. Acad. Sci. USA Vol. 93, pp. 7737-7742, July 1996 Evolution The miniaturized nuclear genome of a eukaryotic endosymbiont contains genes that overlap, genes that are cotranscribed, and the smallest known spliceosomal introns (eukaryotic operons/S13/S4/small nuclear RNP E/clp protease) PAUL R. GILSON* AND GEOFFREY I. MCFADDEN Plant Cell Biology Research Centre, School of Botany, University of Melbourne, Parkville, 3052 Victoria, Australia Communicated by Adrienne E. Clarke, University of Melbourne, Parkville, Victoria, Austrialia, February 12, 1996 (received for review December 1, 1995) ABSTRACT Chlorarachniophyte algae contain a com- (8). The nucleomorph telomere motif (TCTAGGGn) is dif- plex, multi-membraned chloroplast derived from the endo- ferent to that of the host nucleus chromosomes (TTAGGGn) symbiosis of a eukaryotic alga. The vestigial nucleus of the (8), which is consiStent with the nucleomorph being the endosymbiont, called the nucleomorph, contains only three genome of a phylogenetically unrelated endosymbiont. small linear chromosomes with a haploid genome size of 380 Previously, chlorarachniophyte nucleomorph DNA has only kb and is the smallest known eukaryotic genome. Nucleotide been shown to encode eukaryotic rRNAs that are incorpo- sequence data from a subtelomeric fragment of chromosome rated into the ribosomes in the vestigial cytoplasm surrounding III were analyzed as a preliminary investigation of the coding the nucleomorph (2). We earlier speculated the nucleomorph's capacity of this vestigial genome. Several housekeeping genes raison d'etre is to provide proteins for the maintenance of the including U6 small nuclear RNA (snRNA), ribosomal proteins chloroplast (2, 7). To synthesize these chloroplast proteins, the S4 and S13, a core protein of the spliceosome [small nuclear nucleomorph may also have to maintain genes that encode ribonucleoprotein (snRNP) E], and a clp-like protease (clpP) expression, translation and self-replication machinery (2, 7).