Rhee Et Al AIDS 2006
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Herpes Simplex Virus Type 1 Oril Is Not Required for Virus Infection In
JOURNAL OF VIROLOGY, Nov. 1987, p. 3528-3535 Vol. 61, No. 11 0022-538X/87/113528-08$02.00/0 Copyright C 1987, American Society for Microbiology Herpes Simplex Virus Type 1 oriL Is Not Required for Virus Replication or for the Establishment and Reactivation of Latent Infection in Mice MARYELLEN POLVINO-BODNAR, PAULO K. ORBERG, AND PRISCILLA A. SCHAFFER* Laboratory of Tumor Virus Genetics, Dana-Farber Cancer Institute, and Department of Microbiology and Molecular Genetics, Harvard Medical School, Boston, Massachusetts 02115 Received 11 May 1987/Accepted 31 July 1987 During the course of experiments designed to isolate deletion mutants of herpes simplex virus type 1 in the gene encoding the major DNA-binding protein, ICP8, a mutant, d61, that grew efficiently in ICP8-expressing Vero cells but not in normal Vero cells was isolated (P. K. Orberg and P. A. Schaffer, J. Virol. 61:1136-1146, 1987). d61 was derived by cotransfection of ICP8-expressing Vero cells with infectious wild-type viral DNA and a plasmid, pDX, that contains an engineered 780-base-pair (bp) deletion in the ICP8 gene, as well as a spontaneous -55-bp deletion in OriL. Gel electrophoresis and Southern blot analysis indicated that d61 DNA carried both deletions present in pDX. The ability of d61 to replicate despite the deletion in OriL suggested that a functional OriL is not essential for virus replication in vitro. Because d61 harbored two mutations, a second mutant, ts+7, with a deletion in oriL-associated sequences and an intact ICP8 gene was constructed. Both d61 and ts+7 replicated efficiently in their respective permissive host cells, although their yields were slightly lower than those of control viruses with intact oriL sequences. -

Foamy Virus Assembly with Emphasis on Pol Encapsidation
Viruses 2013, 5, 886-900; doi:10.3390/v5030886 OPEN ACCESS viruses ISSN 1999-4915 www.mdpi.com/journal/viruses Review Foamy Virus Assembly with Emphasis on Pol Encapsidation Eun-Gyung Lee 1, Carolyn R. Stenbak 2 and Maxine L. Linial 1,* 1 Fred Hutchinson Cancer Research Center, Basic Sciences Division; 1100 Fairview Avenue North, Seattle, WA 98109, USA; E-Mails: [email protected] (EGL); [email protected] (MLL) 2 Seattle University, Biology Department; 901 12th Avenue, Seattle, WA 98122, USA; E-Mail: [email protected] * Authors to whom correspondence should be addressed; E-Mail: [email protected]; Tel.: +1-206- 667-4442; Fax: +1-206-667-5939 Received: 31 January 2013; in revised form: 11 March 2013 / Accepted: 14 March 2013 / Published: 20 March 2013 Abstract: Foamy viruses (FVs) differ from all other genera of retroviruses (orthoretroviruses) in many aspects of viral replication. In this review, we discuss FV assembly, with special emphasis on Pol incorporation. FV assembly takes place intracellularly, near the pericentriolar region, at a site similar to that used by betaretroviruses. The regions of Gag, Pol and genomic RNA required for viral assembly are described. In contrast to orthoretroviral Pol, which is synthesized as a Gag-Pol fusion protein and packaged through Gag-Gag interactions, FV Pol is synthesized from a spliced mRNA lacking all Gag sequences. Thus, encapsidation of FV Pol requires a different mechanism. We detail how WT Pol lacking Gag sequences is incorporated into virus particles. In addition, a mutant in which Pol is expressed as an orthoretroviral-like Gag-Pol fusion protein is discussed. -

Topological Analysis of the Gp41 MPER on Lipid Bilayers Relevant to the Metastable HIV-1 Envelope Prefusion State
Topological analysis of the gp41 MPER on lipid bilayers relevant to the metastable HIV-1 envelope prefusion state Yi Wanga,b, Pavanjeet Kaurc,d, Zhen-Yu J. Sune,1, Mostafa A. Elbahnasawya,b,2, Zahra Hayatic,d, Zhi-Song Qiaoa,b,3, Nhat N. Buic, Camila Chilea,b,4, Mahmoud L. Nasre,5, Gerhard Wagnere, Jia-Huai Wanga,f, Likai Songc, Ellis L. Reinherza,b,6, and Mikyung Kima,g,6 aLaboratory of Immunobiology, Department of Medical Oncology, Dana-Farber Cancer Institute, Boston, MA 02115; bDepartment of Medicine, Harvard Medical School, Boston, MA 02115; cNational High Magnetic Field Laboratory, Florida State University, Tallahassee, FL 32306; dDepartment of Physics, Florida State University, Tallahassee, FL 32306; eDepartment of Biological Chemistry and Molecular Pharmacology, Harvard Medical School, Boston, MA 02115; fDepartment of Cancer Biology, Dana-Farber Cancer Institute, Boston, MA 02215; and gDepartment of Dermatology, Harvard Medical School, Boston, MA 02215 Edited by Peter Palese, Icahn School of Medicine at Mount Sinai, New York, NY, and approved September 23, 2019 (received for review July 18, 2019) The membrane proximal external region (MPER) of HIV-1 envelope immunologically vulnerable epitopes targeted by several of the most glycoprotein (gp) 41 is an attractive vaccine target for elicitation of broadly neutralizing antibodies (bNAbs) developed during the broadly neutralizing antibodies (bNAbs) by vaccination. However, course of natural HIV-1 infection (10–13). Insertion, deletion, current details regarding the quaternary structural organization of and mutations of residues in the MPER defined the functional the MPER within the native prefusion trimer [(gp120/41)3] are elu- importance of the MPER in Env incorporation, viral fusion, and sive and even contradictory, hindering rational MPER immunogen infectivity (14–16). -

(Infected Cell Protein 8) of Herpes Simplex Virus L*
THEJOURNAL OF BIOLOGICAL CHEMISTRY Vol. 262, No. 9, Issue of March 25, pp. 4260-4266, 1987 0 1987 by The American Society of Biological Chemists, Inc Printed in U.S. A. Interaction between theDNA Polymerase and Single-stranded DNA- binding Protein (Infected Cell Protein 8) of Herpes Simplex Virus l* (Received for publication, September 22, 1986) Michael E. O’DonnellS, Per EliasQ,Barbara E. Funnellll, and I. R. Lehman From the Department of Biochemistry, Stanford University School of Medicine, Stanford, California 94305 The herpes virus-encoded DNA replication protein, ICP8 completely inhibits replication of singly primed ssDNA infected cell protein 8 (ICPS), binds specifically to templates by the herpespolymerase. On the other hand, ICP8 single-stranded DNA with a stoichiometry of one ICPS strongly stimulates replication of duplex DNA. It does so, molecule/l2 nucleotides. In the absence of single- however, onlyin the presence of anuclear extract from stranded DNA, it assembles into long filamentous herpes-infected cells. structures. Binding of ICPS inhibits DNA synthesis by the herpes-induced DNA polymerase on singly primed EXPERIMENTALPROCEDURES single-stranded DNA circles. In contrast, ICPS greatly Materials-DNase I and venom phosphodiesterase were obtained stimulates replication of circular duplex DNA by the from Worthington. The syntheticoligodeoxynucleotide (44-mer) was polymerase. Stimulation occurs only in the presence of synthesized by the solid-phase coupling of protected phosphoramidate a nuclear extract from herpes-infected cells. Appear- nucleoside derivatives (11).3H-Labeled #X ssDNA was a gift from ance of the stimulatory activity in nuclear extracts Dr. R. Bryant (this department). pMOB45 (10.5 kilobases) linear coincides closely with the time of appearance of her- duplex plasmid DNA was a gift from Dr. -

Enzyme Activities in Four Different Forms of Human Immunodeficiency
Proc. Nati. Acad. Sci. USA Vol. 88, pp. 45%-4600, June 1991 Biochemistry Enzyme activities in four different forms of human immunodeficiency virus 1 poi gene products (reverse transcriptase/RNase H/crossover linker mutagenesis/baculoirus expression system) YU-WEN HU AND C. YONG KANG Department of Microbiology and Immunology, University of Ottawa, Faculty of Medicine, Ottawa, ON, Canada K1H 8M5 Communicated by Max D. Summers, February 19, 1991 (received for review December 3, 1990) ABSTRACT Five cassettes of the pol gene of human im- activity (10, 11) but no RNase H activity (3, 7, 12, 13). The munodeficiency virus 1 were constructed and inserted under reverse transcriptase isolated from purified HIV-1 has a the control of the polyhedrin gene promoter of Autographa molecular mass of 95 (14) to 110 (15) kDa, as determined by calfornica nuclear polyhedrosis virus by homologous recom- gel filtration and glycerol gradient centrifugation, respec- bination. The first cassette polF contains the full-length pol tively. However, the 98-kDa nonprocessed polyprotein ex- open reading frame; the second cassette pollOO starts with the pressed in Escherichia coli displayed little or no reverse first AUG codon of the pol gene and deletes 103 amino acids transcriptase activity (16). The p66/pSi heterodimer ex- from the amino terminus of the pol gene product; the third pressed in E. coli has an apparent molecular mass of 120-130 cassette po197 deletes the entire protease coding sequence; the kDa and was active (5, 6). In addition, there is a 165-kDa fourth cassette po166 deletes both the protease and endonucle- putative gag-pol precursor polyprotein that also possesses ase/integrase coding sequences; and the fifth cassette polSl reverse transcriptase activity (10). -

Genomic Diversity in Rotavirus and the Current Scenario of Rotavirus Vaccines: a Brief Overview
Central Archives of Paediatrics and Developmental Pathology Bringing Excellence in Open Access Review Article *Corresponding author Vijaya Lakshmi Nag, Department of Microbiology, All India Institute of Medical Sciences, Jodhpur, India, Tel: Genomic Diversity in Rotavirus 918003996874; Email: Submitted: 15 October 2017 and the Current Scenario of Accepted: 01 November 2017 Published: 03 November 2017 Copyright Rotavirus Vaccines: A Brief © 2017 Nag et al. Overview OPEN ACCESS Keywords Vijaya Lakshmi Nag* and Kumar Saurabh • Rotavirus infection Department of Microbiology, All India Institute of Medical Sciences, India • Genomic diversity • Rotavirus vaccines Abstract Rotavirus associated infection still dominates the list of diarrheal illnesses in paediatric population despite the availability of effective Rotavirus vaccines. Frequent genetic reassortment in the genome of rotavirus leads to emergence of novel genotypes in different geographic areas worldwide. This review aims to give a brief overview of the genetic diversity prevailing in rotavirus worldwide with a focus on the vaccines available for rotavirus infection. ABBREVIATIONS may also be used. Electron microscopy and polyacrylamide gel electrophoresis are also useful. Molecular techniques like nested RV: Rotavirus Infection; RNA: Ribonucleic Acid; EIA: polymerase chain reaction (PCR), multiplex PCR and Reverse Enzyme Immunoassay; ELISA: Enzyme Linked Immunosorbent transcriptase PCR (RT-PCR) in stool samples can additionally Assay; IgA: Immunoglobulin A; IgG: Immunoglobulin G; PCR: help in the genotypic studies [5-7]. The genetic sequencing Polymerase Chain Reaction; RT-PCR: Reverse Transcriptase PCR; methods like microarray and real time PCR (RT-qPCR) are very VP: Viral Structural Protein; NSP: Non-Structural Protein; PAGE: sensitive methods for diagnosis [8]. Polyacrylamide Gel Electrophoresis; RRV: Rhesus Rotavirus, UIP: Universal Immunization Programme; WHO: World Health Vaccine development and implementation has seen lots of Organization. -

Lentivirus and Lentiviral Vectors Fact Sheet
Lentivirus and Lentiviral Vectors Family: Retroviridae Genus: Lentivirus Enveloped Size: ~ 80 - 120 nm in diameter Genome: Two copies of positive-sense ssRNA inside a conical capsid Risk Group: 2 Lentivirus Characteristics Lentivirus (lente-, latin for “slow”) is a group of retroviruses characterized for a long incubation period. They are classified into five serogroups according to the vertebrate hosts they infect: bovine, equine, feline, ovine/caprine and primate. Some examples of lentiviruses are Human (HIV), Simian (SIV) and Feline (FIV) Immunodeficiency Viruses. Lentiviruses can deliver large amounts of genetic information into the DNA of host cells and can integrate in both dividing and non- dividing cells. The viral genome is passed onto daughter cells during division, making it one of the most efficient gene delivery vectors. Most lentiviral vectors are based on the Human Immunodeficiency Virus (HIV), which will be used as a model of lentiviral vector in this fact sheet. Structure of the HIV Virus The structure of HIV is different from that of other retroviruses. HIV is roughly spherical with a diameter of ~120 nm. HIV is composed of two copies of positive ssRNA that code for nine genes enclosed by a conical capsid containing 2,000 copies of the p24 protein. The ssRNA is tightly bound to nucleocapsid proteins, p7, and enzymes needed for the development of the virion: reverse transcriptase (RT), proteases (PR), ribonuclease and integrase (IN). A matrix composed of p17 surrounds the capsid ensuring the integrity of the virion. This, in turn, is surrounded by an envelope composed of two layers of phospholipids taken from the membrane of a human cell when a newly formed virus particle buds from the cell. -

Mechanism of Hepatitis B Virus Cccdna Formation
viruses Review Mechanism of Hepatitis B Virus cccDNA Formation Lei Wei and Alexander Ploss * 110 Lewis Thomas Laboratory, Department of Molecular Biology, Princeton University, Washington Road, Princeton, NJ 08544, USA; [email protected] * Correspondence: [email protected]; Tel.: +1-609-258-7128 Abstract: Hepatitis B virus (HBV) remains a major medical problem affecting at least 257 million chronically infected patients who are at risk of developing serious, frequently fatal liver diseases. HBV is a small, partially double-stranded DNA virus that goes through an intricate replication cycle in its native cellular environment: human hepatocytes. A critical step in the viral life-cycle is the conversion of relaxed circular DNA (rcDNA) into covalently closed circular DNA (cccDNA), the latter being the major template for HBV gene transcription. For this conversion, HBV relies on multiple host factors, as enzymes capable of catalyzing the relevant reactions are not encoded in the viral genome. Combinations of genetic and biochemical approaches have produced findings that provide a more holistic picture of the complex mechanism of HBV cccDNA formation. Here, we review some of these studies that have helped to provide a comprehensive picture of rcDNA to cccDNA conversion. Mechanistic insights into this critical step for HBV persistence hold the key for devising new therapies that will lead not only to viral suppression but to a cure. Keywords: hepatitis B virus; HBV; viral replication; cccDNA biogenesis; rcDNA; DNA repair Citation: Wei, L.; Ploss, A. 1. Overview of HBV Life Cycle and cccDNA Biogenesis Mechanism of Hepatitis B Virus cccDNA Formation. Viruses 2021, 13, The hepatotropic HBV belongs to the Hepadnaviridae family and is a blood-borne 1463. -

Capsid Coding Sequence Is Required for Efficient Replication of Human
JOURNAL OF VIROLOGY, Mar. 1996, p. 1941–1952 Vol. 70, No. 3 0022-538X/96/$04.0010 Copyright q 1996, American Society for Microbiology Capsid Coding Sequence Is Required for Efficient Replication of Human Rhinovirus 14 RNA 1 1,2 KEVIN L. MCKNIGHT AND STANLEY M. LEMON * Departments of Microbiology and Immunology1 and Medicine,2 The University of North Carolina at Chapel Hill, Chapel Hill, North Carolina 27599-7030 Received 24 August 1995/Accepted 1 December 1995 Mechanisms by which the plus-sense RNA genomes of picornaviruses are replicated remain poorly defined, but existing models do not suggest a role for sequences encoding the capsid proteins. However, candidate RNA replicons (DP1bgal and DP1Luc), representing the sequence of human rhinovirus 14 virus (HRV-14) with reporter protein sequences (b-galactosidase or luciferase, respectively) replacing most of the P1 capsid-coding region, failed to replicate in transfected H1-HeLa cells despite efficient primary cleavage of the polyprotein. To determine which P1 sequences might be required for RNA replication, HRV-14 mutants in which segments of the P1 region were removed in frame from the genome were constructed. Mutants with deletions involving the 5*-proximal 1,489 nucleotides of the P1 region replicated efficiently, while those with deletions involving the 3* 1,079 nucleotides did not. Reintroduction of the 3* P1 sequence into the nonreplicating DP1Luc construct resulted in a new candidate replicon, DP1Luc/VP3, which replicated well and expressed luciferase efficiently. Capsid proteins provided in trans by helper virus failed to rescue the nonreplicating DP1Luc genome but were able to package the larger-than-genome-length DP1Luc/VP3 replicon. -

Studies on HIV-1 Virion Infectivity Factor
Studies on HIV-L Virion Infectivity Factor Feng Feng M.Sc. Infectious Diseases Laboratories Institute of Medical and Veterinary Science Adelaide South Australia School of Molecular & Biomedical Science University of Adelaide Adelaide South Australia ¡k ¡l.rl. {. t {. {. ¡1. ¡ß {. tk * *,1. tß * * * * {. A thesis submitted to the University of Adelaide infuffilment of the requirements far the degree of Doctor of, Philosophy October,2004 Contents VI Declaration of Originality.... .................. IX X Publications Related to this Study xll Chapter L. IntrOduCtiOn... .................. ... .... o. ........ 1 1.1 Historical Background of HMAIDS 1..2 Overview of Human Immunodeficiency Virus............ 4 1.2.1 Classification of HIV 4 1.2.2 }J.IV virion structure 5 1.2.3 Genetic organisation of HIV 5 6 1 .2.4 IF'IV replication CYcle. 1.2.4.t Viral attachment and viral fusion 7 1.2.4.2 Reverse transcription. .' . I 1.2.4.3 Viral DNA integration.. '... 9 I.2.4.4 Transcription and translation l0 1.2.4.5 Assembly of virus and viral budding. 1l 1.2.5 HIV gene expression. t2 1.2.5.1HIV proviral genome transcription - overview ... ' l2 I.2.5.2 Regulation of transcription' ' l3 1.2.6 I{IV encoded proteins and their functions. l5 I.2.6,1 The major structure proteins. 15 I.2.6.2 Regulatory proteinsiaccessary proteins. .' l8 1.2.t HIV-1 Virion infectivity factor (ViÐ'.. 24 1.2.7.1HIV-l Vif localisation and biological activity' 24 112.7 .2 Vif is required for efficient reverse transcription. .. 25 I.2.7.3 Vif interacts with viral proteins and RNA. -
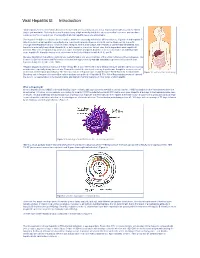
Viral Hepatitis B: Introduction
Viral Hepatitis B: Introduction “Viral hepatitis," refers to infections that affect the liver and are caused by viruses. It is a major public health issue in the United States and worldwide. Not only does viral hepatitis carry a high morbidity, but it also stresses medical resources and can have severe economic consequences. The majority of all viral hepatitis cases are preventable. Viral hepatitis includes five distinct disease entities, which are caused by at least five different viruses. Hepatitis A and hepatitis B (infectious and serum hepatitis, respectively) are considered separate diseases and both can be diagnosed by a specific serologic test. Hepatitis C and E comprise a third category, each a distinct type, with Hepatitis C parenterally transmitted, and hepatitis E enterically transmitted. Hepatitis D, or delta hepatitis, is another distinct virus that is dependent upon hepatitis B infection. This form of hepatitis may occur as a super-infectionin a hepatitis B carrier or as a co-infection in an individual with acute hepatitis B. Hepatitis viruses most often found in the United States include A, B, C, and D. Because fatality from hepatitis is relatively low, mortality figures are a poor indicator of the actual incidence of these diseases. The Centers for Disease Control and Prevention estimated that approximately 400,000–600,000 people were infected with viral hepatitis during the decade of the 1990s. Hepatitis plagued mankind as early as the fifth century BC. It was referenced in early biblical literature and described as occurring in outbreaks, especially during times of war. Toward the end of the nineteenth century, hepatitis was thought to occur as a result of infection of the hepatic parenchyma. -

The Sars-Cov-2 Viroporin E and Its Interaction with Amantadine an Analysis
Journal of Pharmaceutics & Pharmacology Special Issue Repurposing of Adamantanes for the Potential Prevention or Treatment of COVID-19 Editor: Dr. Butterworth Roger F Professor of Medicine, University of Montreal, Canada Avens Publishing Group Inviting Innovations Open Access Review Article J Pharmaceu Pharmacol September 2020 Vol.:8, Issue:1 © All rights are reserved by Aranda-Abreu GE. AvensJournal Publishing of Group Inviting Innovations The Sars-Cov-2 Viroporin E and Pharmaceutics & its Interaction with Amantadine Pharmacology Aranda-Abreu GE*, Hernández-Aguilar ME, an Analysis Herrera-Covarrubias D and Rojas-Durán F Universidad Veracruzana/Centro de Investigaciones Cerebrales, México Abstract Address for Correspondence This article discusses the function of the SARS-Cov-2 virus E-channel Aranda-Abreu GE, Universidad Veracruzana/Centro de Investigaciones and its interaction with amantadine.In this analysis it is proposed that Cerebrales, Xalapa, Veracruz, México; E-mail: [email protected] amantadine is able to enter the E-channel of the coronavirus, inhibiting the viral content to enter the cell avoiding viral replication. Therefore, Submission: 14 August 2020 amantadine is a drug that can help decrease the symptoms of Accepted: 15 September 2020 coronavirus. It is also mentioned how amantadine may be acting with Published: 21 September 2020 the Spike protein, through a molecular docking study. Copyright: © 2020 Aranda-Abreu GE, et al. This is an open access article distributed under the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any Introduction medium, provided the original work is properly cited. At the end of 2019, a new virus called COVID-19 was spreading in an alarming way in Wuhan City in China.