Introduction to Plant Pathology {Richard N. Strange
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Revisiting the Taxonomy of Allorhizobium Vitis (Ie
bioRxiv preprint doi: https://doi.org/10.1101/2020.12.19.423612; this version posted December 21, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-ND 4.0 International license. Revisiting the taxonomy of Allorhizobium vitis (i.e. Agrobacterium vitis) using genomics - emended description of All. vitis sensu stricto and description of Allorhizobium ampelinum sp. nov. Nemanja Kuzmanović1,*, Enrico Biondi2, Jörg Overmann3, Joanna Puławska4, Susanne Verbarg3, Kornelia Smalla1, Florent Lassalle5,6,* 1Julius Kühn-Institut, Federal Research Centre for Cultivated Plants (JKI), Institute for Epidemiology and Pathogen Diagnostics, Messeweg 11-12, 38104 Braunschweig, Germany 2Alma Mater Studiorum - University of Bologna, Viale G. Fanin, 42, 40127 Bologna, Italy 3Leibniz Institute DSMZ-German Collection of Microorganisms and Cell Cultures, Inhoffenstrasse 7B, 38124 Braunschweig, Germany 4Research Institute of Horticulture, ul. Konstytucji 3 Maja 1/3, 96-100 Skierniewice, Poland 5Imperial College London, St-Mary’s Hospital campus, Department of Infectious Disease Epidemiology, Praed Street, London W2 1NY, UK; Imperial College London, St-Mary’s Hospital campus, MRC Centre for Global Infectious Disease Analysis, Praed Street, London W2 1NY, United Kingdom 6Wellcome Sanger Institute, Pathogens and Microbes Programme, Wellcome Genome Campus, Hinxton, Saffron Walden, CB10 1RQ, United Kingdom *Corresponding authors. Contact: [email protected], [email protected] (N. Kuzmanovid); [email protected] (F. Lassalle) bioRxiv preprint doi: https://doi.org/10.1101/2020.12.19.423612; this version posted December 21, 2020. -

Biocontrol of Crown Gall by Rhizobium Rhizogenes
agronomy Case ReportReport Biocontrol of Crown Gall byby RhizobiumRhizobium rhizogenesrhizogenes:: Challenges in Biopesticide Commercialisation 1 2, Allen Kerr 1 and Gary Bullard 2,** 1 Department of Plant Pathology, University of Adelaide, Adelaide, SA 5064, Australia; [email protected] 1 Department of Plant Pathology, University of Adelaide, Adelaide SA 5064, Australia; [email protected] 2 Bio-Care Technology Pty Ltd., Myocum, NSW 2481, Australia 2 Bio-Care Technology Pty Ltd., Myocum NSW 2481, Australia * Correspondence: [email protected] * Correspondence: [email protected] Received: 18 June 2020 2020;; Accepted: 30 July 2020 2020;; Published: 3 3 August August 2020 2020 Abstract: The biocontrolbiocontrol of crown gall has been practisedpractised in AustraliaAustralia for 4848 years.years. Control is so eefficientfficient thatthat itit is is di difficultfficult to to find find a galleda galled stone stone fruit fruit tree, tree, when when previously, previously, crown crown gall had gall been had abeen major a problem.major problem. This paper This explainspaper explains how it works how andit works why onlyand pathogenswhy only arepathog inhibited.ens are A inhibited. commercial A biopesticidecommercial biopesticide is available inis Australia,available in Canada, Australia, Chile, Canada, New Zealand,Chile, New Turkey, Zealand, the USA, Turkey, South the Africa USA, andSouth Japan. Africa The and challenges Japan. The of commercialisingchallenges of commercialising a biopesticide area biopesticide outlined. Rigid -

Revised Taxonomy of the Family Rhizobiaceae, and Phylogeny of Mesorhizobia Nodulating Glycyrrhiza Spp
Division of Microbiology and Biotechnology Department of Food and Environmental Sciences University of Helsinki Finland Revised taxonomy of the family Rhizobiaceae, and phylogeny of mesorhizobia nodulating Glycyrrhiza spp. Seyed Abdollah Mousavi Academic Dissertation To be presented, with the permission of the Faculty of Agriculture and Forestry of the University of Helsinki, for public examination in lecture hall 3, Viikki building B, Latokartanonkaari 7, on the 20th of May 2016, at 12 o’clock noon. Helsinki 2016 Supervisor: Professor Kristina Lindström Department of Environmental Sciences University of Helsinki, Finland Pre-examiners: Professor Jaakko Hyvönen Department of Biosciences University of Helsinki, Finland Associate Professor Chang Fu Tian State Key Laboratory of Agrobiotechnology College of Biological Sciences China Agricultural University, China Opponent: Professor J. Peter W. Young Department of Biology University of York, England Cover photo by Kristina Lindström Dissertationes Schola Doctoralis Scientiae Circumiectalis, Alimentariae, Biologicae ISSN 2342-5423 (print) ISSN 2342-5431 (online) ISBN 978-951-51-2111-0 (paperback) ISBN 978-951-51-2112-7 (PDF) Electronic version available at http://ethesis.helsinki.fi/ Unigrafia Helsinki 2016 2 ABSTRACT Studies of the taxonomy of bacteria were initiated in the last quarter of the 19th century when bacteria were classified in six genera placed in four tribes based on their morphological appearance. Since then the taxonomy of bacteria has been revolutionized several times. At present, 30 phyla belong to the domain “Bacteria”, which includes over 9600 species. Unlike many eukaryotes, bacteria lack complex morphological characters and practically phylogenetically informative fossils. It is partly due to these reasons that bacterial taxonomy is complicated. -

Richard STRANGE of Rodbourne Cheney
Descendants of Richard STRANGE First Generation 1. Richard Strange 1, died on 19 Jan 1713/14 in Rodbourne Cheney, Wiltshire2. Richard married Bridget1 before 16861. (Bridget died on 4 Jun 1731 in Rodbourne Cheney, Wiltshire2.) Children from this marriage were: + 2 M i. Richard Strange 1, 3 was born in 1696 in Rodbourne Cheney, Wiltshire1, was christened on 11 Jun 1696 in Rodbourne Cheney, Wiltshire1 and died on 3 Jul 1737 in Blunsdon St Andrew, Wiltshire1, 2, 3. 3 F ii. Bridget Strange 1 was born in 1686 in Rodbourne Cheney, Wiltshire1 and was christened on 7 Oct 1686 in Rodbourne Cheney, Wiltshire1. Second Generation (Children) 2. Richard Strange 1, 3 (Richard1) was born in 1696 in Rodbourne Cheney, Wiltshire1, was christened on 11 Jun 1696 in Rodbourne Cheney, Wiltshire1 and died on 3 Jul 1737 in Blunsdon St Andrew, Wiltshire1, 2, 3. Richard married Ann Dyer on 21 Feb 1720/214. (Ann Dyer was born before 17004, died in 1768 in Marston, Wiltshire and was buried on 11 Oct 1768 in Rodbourne Cheney, Wiltshire2.) Children from this marriage were: 4 M i. Richard Strange 1 was born in 1722 in Rodbourne Cheney, Wiltshire1, was christened on 11 Jan 1721/22 in Rodbourne Cheney, Wiltshire1, died in 1740 in London2, 5 and was buried on 16 Aug 1740 in Rodbourne Cheney, Wiltshire2. 5 F ii. Anne Strange 1 was born in 1723 in Rodbourne Cheney, Wiltshire1 and was christened on 22 Mar 1723/24 in Rodbourne Cheney, Wiltshire1. + 6 M iii. Thomas Strange 6, 7 was born in 17257, was christened on 27 Jul 1725 in Rodbourne Cheney, Wiltshire1, died in Oct 1800 in Swindon, Wiltshire7 and was buried on 15 Oct 1800 in Swindon, Wiltshire. -
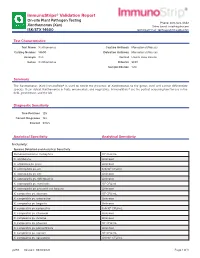
Immunostrips® Validation Report
ImmunoStrips® Validation Report On-site Plant Pathogen Testing Phone: 800-622-4342 Xanthomonas (Xan) Sales Email: [email protected] ISK/STX 14600 Technical Email: [email protected] Test Characteristics Test Name Xanthomonas Capture Antibody Monoclonal (Mouse) Catalog Number 14600 Detection Antibody Monoclonal (Mouse) Acronym Xan Format Lateral Flow Device Genus Xanthomonas Diluents SEB1 Sample Dilution 1:20 Summary The Xanthomonas (Xan) ImmunoStrip® is used to detect the presence of Xanthomonas to the genus level and cannot differentiate species. It can detect Xanthomonas in fruits, ornamentals, and vegetables. ImmunoStrips® are the perfect screening tool for use in the field, greenhouse, and the lab. Diagnostic Sensitivity True Positives 125 Correct Diagnoses 122 Percent 97.6% Analytical Specificity Analytical Sensitivity Inclusivity: Species Detected and Analytical Sensitivity Stenotrophomonas maltophilia 106 CFU/mL X. albilibeans Unknown X. arboricola pv. pruni Unknown X. axonopodis pv. alii 6.9x104 CFU/mL X. axonopodis pv. citri Unknown X. axonopodis pv. diffenbachia Unknown X. axonopodis pv. manihotis 106 CFU/mL X. axonopodis pv. phaseoli var. fuscans Unknown X. campestris pv. aberrans 105 CFU/mL X. campestris pv. armoraciae Unknown X. campestris pv. begonia Unknown X. campestris pv. campestris 6.4x104 CFU/mL X. campestris pv. citromelo Unknown X. campestris pv. incanae Unknown X. campestris pv. phaseoli 105 CFU/mL X. campestris pv. poinsettiicola Unknown X. campestris pv. raphani 105 CFU/mL X. campestris pv. vesicatoria 4.0x105 CFU/mL p293 Revised: 08/18/2021 Page 1 of 3 Species Detected and Analytical Sensitivity X. campestris pv. vitians 105 CFU/mL X. campestris pv. zinnia 107 CFU/mL X. cannabis 1.6x104 CFU/mL X. -

Plant Root 11:33-39
www.plantroot.org 33 Original research article Biolistic-mediated plasmid-free transformation for induction of hairy roots Revi einw a tobaccorticle plantsShort report Gulnar Yasybaeva, Zilya Vershinina, Bulat Kuluev, Elena Mikhaylova, Andrey Baymiev and Aleksey Chemeris Institute of Biochemistry and Genetics, Ufa Scientific Centre, Russian Academy of Sciences (IBG USC RAS), pr. Oktyabrya 71, 450054, Ufa, Russia Corresponding author. G. Yasybaeva, E-mail: [email protected], Phone: +7-937-480-9956 Received on December 13, 2016; Accepted on March 8, 2017 Abstract: T-DNA of Ri-plasmid from Agrobacteri- transformed plant tissue, specifically, promotes the um rhizogenes is able to trigger the hairy root syn- appearance of hairy roots. Hairy roots culture (HRC) drome in infected plants. This natural phenomenon can be used for production of various secondary is used to generate hairy root cultures predomi- metabolites such as alkaloids, terpenoids, phenols, nantly only in dicotyledonous plants. We propose a glycosides (Sharifi et al. 2014). Some of them are a new method of hairy roots induction without Agro- source of pharmaceutical raw materials or other bacterium-mediated transformation. The 5461 bp economically valuable compounds. HRCs also can T-DNA region from A. rhizogenes A4 strain with all be used as recombinant proteins producers (Ono and four rol genes was amplified using primers contain- Tian 2011). ing sequences for left and right T-DNA borders on In natural conditions soil bacteria A. rhizogenes their 3’-ends. This amplicon was used for direct infects healthy plants by penetrating through injured transformation of tobacco leaf discs without A. rhi- roots. Thereby simple way of making hairy roots is zogenes and binary vectors. -

2010.-Hungria-MLI.Pdf
Mohammad Saghir Khan l Almas Zaidi Javed Musarrat Editors Microbes for Legume Improvement SpringerWienNewYork Editors Dr. Mohammad Saghir Khan Dr. Almas Zaidi Aligarh Muslim University Aligarh Muslim University Fac. Agricultural Sciences Fac. Agricultural Sciences Dept. Agricultural Microbiology Dept. Agricultural Microbiology 202002 Aligarh 202002 Aligarh India India [email protected] [email protected] Prof. Dr. Javed Musarrat Aligarh Muslim University Fac. Agricultural Sciences Dept. Agricultural Microbiology 202002 Aligarh India [email protected] This work is subject to copyright. All rights are reserved, whether the whole or part of the material is concerned, specifically those of translation, reprinting, re-use of illustrations, broadcasting, reproduction by photocopying machines or similar means, and storage in data banks. Product Liability: The publisher can give no guarantee for all the information contained in this book. The use of registered names, trademarks, etc. in this publication does not imply, even in the absence of a specific statement, that such names are exempt from the relevant protective laws and regulations and therefore free for general use. # 2010 Springer-Verlag/Wien Printed in Germany SpringerWienNewYork is a part of Springer Science+Business Media springer.at Typesetting: SPI, Pondicherry, India Printed on acid-free and chlorine-free bleached paper SPIN: 12711161 With 23 (partly coloured) Figures Library of Congress Control Number: 2010931546 ISBN 978-3-211-99752-9 e-ISBN 978-3-211-99753-6 DOI 10.1007/978-3-211-99753-6 SpringerWienNewYork Preface The farmer folks around the world are facing acute problems in providing plants with required nutrients due to inadequate supply of raw materials, poor storage quality, indiscriminate uses and unaffordable hike in the costs of synthetic chemical fertilizers. -

Punk · Film RARE PERIODICALS RARE
We specialize in RARE JOURNALS, PERIODICALS and MAGAZINES Please ask for our Catalogues and come to visit us at: rare PERIODIcAlS http://antiq.benjamins.com music · pop · beat · PUNk · fIlM RARE PERIODICALS Search from our Website for Unusual, Rare, Obscure - complete sets and special issues of journals, in the best possible condition. Avant Garde Art Documentation Concrete Art Fluxus Visual Poetry Small Press Publications Little Magazines Artist Periodicals De-Luxe editions CAT. Beat Periodicals 296 Underground and Counterculture and much more Catalogue No. 296 (2016) JOHN BENJAMINS ANTIQUARIAT Visiting address: Klaprozenweg 75G · 1033 NN Amsterdam · The Netherlands Postal address: P.O. BOX 36224 · 1020 ME Amsterdam · The Netherlands tel +31 20 630 4747 · fax +31 20 673 9773 · [email protected] JOHN BENJAMINS ANTIQUARIAT B.V. AMSTERDAM cat.296.cover.indd 1 05/10/2016 12:39:06 antiquarian PERIODIcAlS MUSIC · POP · BEAT · PUNK · FILM Cover illustrations: DOWN BEAT ROLLING STONE [#19111] page 13 [#18885] page 62 BOSTON ROCK FLIPSIDE [#18939] page 7 [#18941] page 18 MAXIMUM ROCKNROLL HEAVEN [#16254] page 36 [#18606] page 24 Conditions of sale see inside back-cover Catalogue No. 296 (2016) JOHN BENJAMINS ANTIQUARIAT B.V. AMSTERDAM 111111111111111 [#18466] DE L’AME POUR L’AME. The Patti Smith Fan Club Journal Numbers 5 and 6 (out of 8 published). October 1977 [With Related Ephemera]. - July 1978. [Richmond Center, WI]: (The Patti Smith Fan Club), (1978). Both first editions. 4to., 28x21,5 cm. side-stapled wraps. Photo-offset duplicated. Both fine, in original mailing envelopes (both opened a bit rough but otherwise good condition). EUR 1,200.00 Fanzine published in Wisconsin by Nanalee Berry with help from Patti’s mom Beverly. -

Gall-ID: Tools for Genotyping Gall-Causing Phytopathogenic Bacteria
Gall-ID: tools for genotyping gall-causing phytopathogenic bacteria Edward W. Davis II1,2,*, Alexandra J. Weisberg1,*, Javier F. Tabima1, Niklaus J. Grunwald1,2,3,4 and Jeff H. Chang1,2,3 1 Department of Botany and Plant Pathology, Oregon State University, Corvallis, OR, United States 2 Molecular and Cellular Biology Program, Oregon State University, Corvallis, OR, United States 3 Center for Genome Research and Biocomputing, Oregon State University, Corvallis, OR, United States 4 Horticultural Crops Research Laboratory, USDA-ARS, Corvallis, OR, United States * These authors contributed equally to this work. ABSTRACT Understanding the population structure and genetic diversity of plant pathogens, as well as the effect of agricultural practices on pathogen evolution, is important for disease management. Developments in molecular methods have contributed to increase the resolution for accurate pathogen identification, but those based on analysis of DNA sequences can be less straightforward to use. To address this, we developed Gall-ID, a web-based platform that uses DNA sequence information from 16S rDNA, multilocus sequence analysis and whole genome sequences to group disease-associated bacteria to their taxonomic units. Gall-ID was developed with a particular focus on gall-forming bacteria belonging to Agrobacterium, Pseudomonas savastanoi, Pantoea agglomerans, and Rhodococcus. Members of these groups of bacteria cause growth deformation of plants, and some are capable of infecting many species of field, orchard, and nursery crops. Gall-ID also enables the use of high-throughput sequencing reads to search for evidence for homologs of characterized virulence genes, and provides downloadable software pipelines for automating multilocus sequence analysis, analyzing genome sequences for average nucleotide identity, and constructing core genome phylogenies. -

Karen Mantler by Karen Mantler
Karen Mantler by Karen Mantler I was conceived by Carla Bley and Michael Mantler at the Newport Jazz Festival in 1965. Born in 1966, I was immediately swept into the musician's life on the road. After having checked me at the coatroom of the Berlin Jazz Festival, to the horror of the press, my parents realized that I was going to have to learn to play an instrument in order to be useful. But since I was still just a baby and they couldn't leave me alone, they had to bring me on stage with them and keep me under the piano. This is probably why I feel most at home on the stage. In 1971, when I was four, my mother let me have a part in Escalator Over The Hill and the next year I sang on another of her records, Tropic Appetites. By 1977 I had learned to play the glockenspiel, and I joined the Carla Bley Band. I toured Europe and the States with her several times and played on her Musique Mecanique album. After playing at Carnegie Hall in 1980, where I tried to steal the show by pretending to be Carla Bley, my mother fired me, telling me "get your own band". I realized that I was going to have to learn a more complicated instrument. After trying drums, bass, and flute, which I always lost interest in, I settled on the clarinet. I joined my elementary school band and quickly rose to the head of the clarinet section. The band director let me take the first improvised solo in the history of the Phoenicia (a small town near Woodstock, NY) elementary school. -

Mr Lonely by Richard Strange
Mr Lonely by Richard Strange As we boarded the flight from London to Inverness, it seemed perhaps the filming of Harmony Korine’s new film, “Mr Lonely”, had already begun. A 6’2” bristly-chinned, barrel-chested “stewardess” welcomed us aboard the aircraft and announced, in a booming baritone voice, “Good Morning, My name is Sarah. Welcome aboard.” As it transpired, Sarah was a wise-cracking testament to the flight company’s equal opportunities programme. (S)he kept us amused for the whole 2-hour flight, even through the bumpiest of landings in a blustery Scots squall. I had been a fan of Harmony Korine’s since his 1997 directorial debut, the disturbingly quirky Gummo. However, I had no idea that the fidgety, punky livewire I spent two hours chatting with at a London party was Korine. I never thought to ask him what work he did- we were laughing too much, exchanging ever-more outrageous stories, and comparing near-death experiences. Only when I was leaving the party did we exchange phone numbers. A week later, as I was about to fly to Los Angeles for a 3-month run of the Tom Waits/Robert Wilson/William Burroughs theatrical collaboration The Black Rider, Korine called me and announced, “Hey. I want you to play Abe Lincoln in my new movie. We film in the Highlands of Scotland, June through August. Do the dates work?” I told him they worked, and had “his people speak to my people”. And so in June I boarded the low-cost flight heading north. -

Michael Stevens' the Road to Interzone
“The scholarship surrounding the life and work of William Burroughs is in the midst of a renaissance. Students of Burroughs are turning away from myths, legends, and sensationalistic biographical detail in order to delve deeply into textual analysis, archival research, and explorations of literary and artistic history. Michael Stevens’ The Road to Interzone is an important part of this changing landscape. In a manner similar to Ralph Maud’s Charles Olson’s Reading, The Road to Interzone places the life and literature of “el Hombre Invisible” into sharper focus by listing and commenting on, in obsessive detail, the breadth of literary material Burroughs read, referred to, researched, and reviewed. Stevens reveals Burroughs to be a man of letters and of great learning, while simultaneously shedding light on the personal obsessions, pet theories, childhood favorites, and guilty pleasures, which make Burroughs such a unique and fascinating figure. Stevens’ book provides a wealth of new and important information for those deeply interested in Burroughs and will no doubt prove essential to future scholarship. Like Oliver Harris’ The Secret of Fascination and Robert Sobieszek’s Ports of Entry before it, The Road to Interzone is an indispensable addition to the canon of Burroughs Studies.” -Jed Birmingham “Michael Stevens has created a new kind of biography out of love for William S. Burroughs and love of books. Author worship and bibliophilia become one at the point of obsession, which of course is the point where they become interesting. Burroughs’ reading was intense and far flung, and Stevens has sleuthed out a portrait of that reading--the books Burroughs lent his name to in the form of introductions and blurbs, the books in his various libraries, the books he refers to, the books that found their way into his writing, and much more! Along with lively notes from Stevens, we have Burroughs throughout--his opinions, perceptions, the ‘grain of his voice.’ That in itself makes Stevens’ book a notable achievement.