Emerging Roles for Non-Selenium Containing ER-Resident Glutathione
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Upregulation of Peroxisome Proliferator-Activated Receptor-Α And
Upregulation of peroxisome proliferator-activated receptor-α and the lipid metabolism pathway promotes carcinogenesis of ampullary cancer Chih-Yang Wang, Ying-Jui Chao, Yi-Ling Chen, Tzu-Wen Wang, Nam Nhut Phan, Hui-Ping Hsu, Yan-Shen Shan, Ming-Derg Lai 1 Supplementary Table 1. Demographics and clinical outcomes of five patients with ampullary cancer Time of Tumor Time to Age Differentia survival/ Sex Staging size Morphology Recurrence recurrence Condition (years) tion expired (cm) (months) (months) T2N0, 51 F 211 Polypoid Unknown No -- Survived 193 stage Ib T2N0, 2.41.5 58 F Mixed Good Yes 14 Expired 17 stage Ib 0.6 T3N0, 4.53.5 68 M Polypoid Good No -- Survived 162 stage IIA 1.2 T3N0, 66 M 110.8 Ulcerative Good Yes 64 Expired 227 stage IIA T3N0, 60 M 21.81 Mixed Moderate Yes 5.6 Expired 16.7 stage IIA 2 Supplementary Table 2. Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analysis of an ampullary cancer microarray using the Database for Annotation, Visualization and Integrated Discovery (DAVID). This table contains only pathways with p values that ranged 0.0001~0.05. KEGG Pathway p value Genes Pentose and 1.50E-04 UGT1A6, CRYL1, UGT1A8, AKR1B1, UGT2B11, UGT2A3, glucuronate UGT2B10, UGT2B7, XYLB interconversions Drug metabolism 1.63E-04 CYP3A4, XDH, UGT1A6, CYP3A5, CES2, CYP3A7, UGT1A8, NAT2, UGT2B11, DPYD, UGT2A3, UGT2B10, UGT2B7 Maturity-onset 2.43E-04 HNF1A, HNF4A, SLC2A2, PKLR, NEUROD1, HNF4G, diabetes of the PDX1, NR5A2, NKX2-2 young Starch and sucrose 6.03E-04 GBA3, UGT1A6, G6PC, UGT1A8, ENPP3, MGAM, SI, metabolism -

Non-Homologous Isofunctional Enzymes: a Systematic Analysis Of
Omelchenko et al. Biology Direct 2010, 5:31 http://www.biology-direct.com/content/5/1/31 RESEARCH Open Access Non-homologousResearch isofunctional enzymes: A systematic analysis of alternative solutions in enzyme evolution Marina V Omelchenko, Michael Y Galperin*, Yuri I Wolf and Eugene V Koonin Abstract Background: Evolutionarily unrelated proteins that catalyze the same biochemical reactions are often referred to as analogous - as opposed to homologous - enzymes. The existence of numerous alternative, non-homologous enzyme isoforms presents an interesting evolutionary problem; it also complicates genome-based reconstruction of the metabolic pathways in a variety of organisms. In 1998, a systematic search for analogous enzymes resulted in the identification of 105 Enzyme Commission (EC) numbers that included two or more proteins without detectable sequence similarity to each other, including 34 EC nodes where proteins were known (or predicted) to have distinct structural folds, indicating independent evolutionary origins. In the past 12 years, many putative non-homologous isofunctional enzymes were identified in newly sequenced genomes. In addition, efforts in structural genomics resulted in a vastly improved structural coverage of proteomes, providing for definitive assessment of (non)homologous relationships between proteins. Results: We report the results of a comprehensive search for non-homologous isofunctional enzymes (NISE) that yielded 185 EC nodes with two or more experimentally characterized - or predicted - structurally unrelated proteins. Of these NISE sets, only 74 were from the original 1998 list. Structural assignments of the NISE show over-representation of proteins with the TIM barrel fold and the nucleotide-binding Rossmann fold. From the functional perspective, the set of NISE is enriched in hydrolases, particularly carbohydrate hydrolases, and in enzymes involved in defense against oxidative stress. -

(ER) Membrane Contact Sites (MCS) Uses Toxic Waste to Deliver Messages Edgar Djaha Yoboue1, Roberto Sitia1 and Thomas Simmen2
Yoboue et al. Cell Death and Disease (2018) 9:331 DOI 10.1038/s41419-017-0033-4 Cell Death & Disease REVIEW ARTICLE Open Access Redox crosstalk at endoplasmic reticulum (ER) membrane contact sites (MCS) uses toxic waste to deliver messages Edgar Djaha Yoboue1, Roberto Sitia1 and Thomas Simmen2 Abstract Many cellular redox reactions housed within mitochondria, peroxisomes and the endoplasmic reticulum (ER) generate hydrogen peroxide (H2O2) and other reactive oxygen species (ROS). The contribution of each organelle to the total cellular ROS production is considerable, but varies between cell types and also over time. Redox-regulatory enzymes are thought to assemble at a “redox triangle” formed by mitochondria, peroxisomes and the ER, assembling “redoxosomes” that sense ROS accumulations and redox imbalances. The redoxosome enzymes use ROS, potentially toxic by-products made by some redoxosome members themselves, to transmit inter-compartmental signals via chemical modifications of downstream proteins and lipids. Interestingly, important components of the redoxosome are ER chaperones and oxidoreductases, identifying ER oxidative protein folding as a key ROS producer and controller of the tri-organellar membrane contact sites (MCS) formed at the redox triangle. At these MCS, ROS accumulations could directly facilitate inter-organellar signal transmission, using ROS transporters. In addition, ROS influence the flux 2+ 2+ of Ca ions, since many Ca handling proteins, including inositol 1,4,5 trisphosphate receptors (IP3Rs), SERCA pumps or regulators of the mitochondrial Ca2+ uniporter (MCU) are redox-sensitive. Fine-tuning of these redox and ion signaling pathways might be difficult in older organisms, suggesting a dysfunctional redox triangle may accompany 1234567890 1234567890 the aging process. -

Increased Whitening Efficacy and Reduced Cytotoxicity Are Achieved by the Chemical Activation of a Highly Concentrated Hydrogen Peroxide Bleaching Gel
Original Article http://dx.doi.org/10.1590/1678-7757-2018-0453 Increased whitening efficacy and reduced cytotoxicity are achieved by the chemical activation of a highly concentrated hydrogen peroxide bleaching gel Abstract Diana Gabriela SOARES1 Objective: This study was designed for the chemical activation of a 35% hydrogen peroxide (H O ) bleaching gel to increase its whitening effectiveness Natália MARCOMINI² 2 2 and reduce its toxicity. Methodology: First, the bleaching gel - associated or Carla Caroline de Oliveira DUQUE³ not with ferrous sulfate (FS), manganese chloride (MC), peroxidase (PR), Ester Alves Ferreira BORDINI³ or catalase (CT) - was applied (3x 15 min) to enamel/dentin discs adapted Uxua Ortecho ZUTA³ to artificial pulp chambers. Then, odontoblast-like MDPC-23 cells were Fernanda Gonçalves BASSO⁴ exposed for 1 h to the extracts (culture medium + components released Josimeri HEBLING⁵ from the product), for the assessment of viability (MTT assay) and oxidative D Carlos Alberto de Souza COSTA⁴ stress (H2DCFDA). Residual H2O2 and bleaching effectiveness ( E) were also evaluated. Data were analyzed with one-way ANOVA complemented with Tukey’s test (n=8. p<0.05). Results: All chemically activated groups minimized MDPC-23 oxidative stress generation; however, significantly higher cell viability was detected for MC, PR, and CT than for plain 35% H2O2 gel. Nevertheless, FS, MC, PR, and CT reduced the amount of residual H2O2 and increased bleaching effectiveness. Conclusion: Chemical activation of 35% H2O2 gel with MC, PR, and CT minimized residual H2O2 and pulp cell toxicity; but PR duplicated the whitening potential of the bleaching gel after a single 45-minute session. -

Chloride Peroxidase Cat
chloride peroxidase Cat. No. EXWM-0491 Lot. No. (See product label) Introduction Description Brings about the chlorination of a range of organic molecules, forming stable C-Cl bonds. Also oxidizes bromide and iodide. Enzymes of this type are either heme-thiolate proteins, or contain vanadate. A secreted enzyme produced by the ascomycetous fungus Caldariomyces fumago (Leptoxyphium fumago) is an example of the heme-thiolate type. It catalyses the production of hypochlorous acid by transferring one oxygen atom from H2O2 to chloride. At a separate site it catalyses the chlorination of activated aliphatic and aromatic substrates, via HClO and derived chlorine species. In the absence of halides, it shows peroxidase (e.g. phenol oxidation) and peroxygenase activities. The latter inserts oxygen from H2O2 into, for example, styrene (side chain epoxidation) and toluene (benzylic hydroxylation), however, these activities are less pronounced than its activity with halides. Has little activity with non-activated substrates such as aromatic rings, ethers or saturated alkanes. The chlorinating peroxidase produced by ascomycetous fungi (e.g. Curvularia inaequalis) is an example of a vanadium chloroperoxidase, and is related to bromide peroxidase (EC 1.11.1.18). It contains vanadate and oxidizes chloride, bromide and iodide into hypohalous acids. In the absence of halides, it peroxygenates organic sulfides and oxidizes ABTS [2,2'-azinobis(3-ethylbenzthiazoline-6-sulfonic acid)] but no phenols. Synonyms chloroperoxidase; CPO; vanadium haloperoxidase Product Information Form Liquid or lyophilized powder EC Number EC 1.11.1.10 CAS No. 9055-20-3 Reaction RH + chloride + H2O2 = RCl + 2 H2O Notes This item requires custom production and lead time is between 5-9 weeks. -
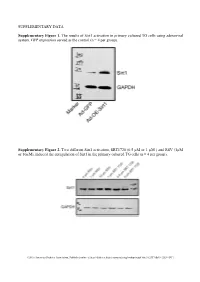
SUPPLEMENTARY DATA Supplementary Figure 1. The
SUPPLEMENTARY DATA Supplementary Figure 1. The results of Sirt1 activation in primary cultured TG cells using adenoviral system. GFP expression served as the control (n = 4 per group). Supplementary Figure 2. Two different Sirt1 activators, SRT1720 (0.5 µM or 1 µM ) and RSV (1µM or 10µM), induced the upregulation of Sirt1 in the primary cultured TG cells (n = 4 per group). ©2016 American Diabetes Association. Published online at http://diabetes.diabetesjournals.org/lookup/suppl/doi:10.2337/db15-1283/-/DC1 SUPPLEMENTARY DATA Supplementary Table 1. Primers used in qPCR Gene Name Primer Sequences Product Size (bp) Sirt1 F: tgccatcatgaagccagaga 241 (NM_001159589) R: aacatcgcagtctccaagga NOX4 F: tgtgcctttattgtgcggag 172 (NM_001285833.1) R: gctgatacactggggcaatg Supplementary Table 2. Antibodies used in Western blot or Immunofluorescence Antibody Company Cat. No Isotype Dilution Sirt1 Santa Cruz * sc-15404 Rabbit IgG 1/200 NF200 Sigma** N5389 Mouse IgG 1/500 Tubulin R&D# MAB1195 Mouse IgG 1/500 NOX4 Abcam† Ab133303 Rabbit IgG 1/500 NOX2 Abcam Ab129068 Rabbit IgG 1/500 phospho-AKT CST‡ #4060 Rabbit IgG 1/500 EGFR CST #4267 Rabbit IgG 1/500 Ki67 Santa Cruz sc-7846 Goat IgG 1/500 * Santa Cruz Biotechnology, Santa Cruz, CA, USA ** Sigma aldrich, Shanghai, China # R&D Systems Inc, Minneapolis, MN, USA † Abcam, Inc., Cambridge, MA, USA ‡ Cell Signaling Technology, Inc., Danvers, MA, USA ©2016 American Diabetes Association. Published online at http://diabetes.diabetesjournals.org/lookup/suppl/doi:10.2337/db15-1283/-/DC1 SUPPLEMENTARY DATA Supplementary -

2011/058582 Al O
(12) INTERNATIONAL APPLICATION PUBLISHED UNDER THE PATENT COOPERATION TREATY (PCT) (19) World Intellectual Property Organization International Bureau (10) International Publication Number (43) International Publication Date i 1 m 19 May 2011 (19.05.2011) 2011/058582 Al (51) International Patent Classification: Road, Sholinganallur, Chennai 600 119 (IN). CHEN- C07C 259/06 (2006.01) A61P 31/00 (2006.01) NIAPPAN, Vinoth Kumar [IN/IN]; Orchid Research C07D 277/46 (2006.01) A61K 31/426 (2006.01) Laboratories Ltd., R & D Centre: Plot No: 476/14, Old C07D 277/48 (2006.01) A61K 31/55 (2006.01) Mahabalipuram Road, Sholinganallur, Chennai 600 119 C07D 487/08 (2006.01) (IN). GANESAN, Karthikeyan [IN/IN]; Orchid Re search Laboratories Ltd., R & D Centre: Plot No: 476/14, (21) International Application Number: Old Mahabalipuram Road, Sholinganallur, Chennai 600 PCT/IN20 10/000738 119 (IN). NARAYANAN, Shridhar [IN/IN]; Orchid Re (22) International Filing Date: search Laboratories Ltd., R & D Centre: Plot No: 476/14, 12 November 2010 (12.1 1.2010) Old Mahabalipuram Road, Sholinganallur, Chennai 600 119 (IN). (25) Filing Language: English (74) Agent: UDAYAMPALAYAM PALANISAMY, (26) Publication Language: English Senthilkumar; Orchid Chemicals & Pharmaceuticals (30) Priority Data: LTD., R & D Centre: Plot No: 476/14, Old Mahabalipu 2810/CHE/2009 16 November 2009 (16. 11.2009) IN ram Road, Sholinganallur, Chennai 600 119 (IN). (71) Applicant (for all designated States except US): OR¬ (81) Designated States (unless otherwise indicated, for every CHID RESEARCH LABORATORIES LTD. [IN/IN]; kind of national protection available): AE, AG, AL, AM, Orchid Towers, 313, Valluvar Kottam High Road, AO, AT, AU, AZ, BA, BB, BG, BH, BR, BW, BY, BZ, Nungambakkam, Chennai 600 034 (IN). -

GPX4 at the Crossroads of Lipid Homeostasis and Ferroptosis Giovanni C
REVIEW GPX4 www.proteomics-journal.com GPX4 at the Crossroads of Lipid Homeostasis and Ferroptosis Giovanni C. Forcina and Scott J. Dixon* formation of toxic radicals (e.g., R-O•).[5] Oxygen is necessary for aerobic metabolism but can cause the harmful The eight mammalian GPX proteins fall oxidation of lipids and other macromolecules. Oxidation of cholesterol and into three clades based on amino acid phospholipids containing polyunsaturated fatty acyl chains can lead to lipid sequence similarity: GPX1 and GPX2; peroxidation, membrane damage, and cell death. Lipid hydroperoxides are key GPX3, GPX5, and GPX6; and GPX4, GPX7, and GPX8.[6] GPX1–4 and 6 (in intermediates in the process of lipid peroxidation. The lipid hydroperoxidase humans) are selenoproteins that contain glutathione peroxidase 4 (GPX4) converts lipid hydroperoxides to lipid an essential selenocysteine in the enzyme + alcohols, and this process prevents the iron (Fe2 )-dependent formation of active site, while GPX5, 6 (in mouse and toxic lipid reactive oxygen species (ROS). Inhibition of GPX4 function leads to rats), 7, and 8 use an active site cysteine lipid peroxidation and can result in the induction of ferroptosis, an instead. Unlike other family members, GPX4 (PHGPx) can act as a phospholipid iron-dependent, non-apoptotic form of cell death. This review describes the hydroperoxidase to reduce lipid perox- formation of reactive lipid species, the function of GPX4 in preventing ides to lipid alcohols.[7,8] Thus,GPX4ac- oxidative lipid damage, and the link between GPX4 dysfunction, lipid tivity is essential to maintain lipid home- oxidation, and the induction of ferroptosis. ostasis in the cell, prevent the accumula- tion of toxic lipid ROS and thereby block the onset of an oxidative, iron-dependent, non-apoptotic mode of cell death termed 1. -

Role of Oxidative Stress and Nrf2/KEAP1 Signaling in Colorectal Cancer: Mechanisms and Therapeutic Perspectives with Phytochemicals
antioxidants Review Role of Oxidative Stress and Nrf2/KEAP1 Signaling in Colorectal Cancer: Mechanisms and Therapeutic Perspectives with Phytochemicals Da-Young Lee, Moon-Young Song and Eun-Hee Kim * College of Pharmacy and Institute of Pharmaceutical Sciences, CHA University, Seongnam 13488, Korea; [email protected] (D.-Y.L.); [email protected] (M.-Y.S.) * Correspondence: [email protected]; Tel.: +82-31-881-7179 Abstract: Colorectal cancer still has a high incidence and mortality rate, according to a report from the American Cancer Society. Colorectal cancer has a high prevalence in patients with inflammatory bowel disease. Oxidative stress, including reactive oxygen species (ROS) and lipid peroxidation, has been known to cause inflammatory diseases and malignant disorders. In particular, the nuclear factor erythroid 2-related factor 2 (Nrf2)/Kelch-like ECH-related protein 1 (KEAP1) pathway is well known to protect cells from oxidative stress and inflammation. Nrf2 was first found in the homolog of the hematopoietic transcription factor p45 NF-E2, and the transcription factor Nrf2 is a member of the Cap ‘N’ Collar family. KEAP1 is well known as a negative regulator that rapidly degrades Nrf2 through the proteasome system. A range of evidence has shown that consumption of phytochemicals has a preventive or inhibitory effect on cancer progression or proliferation, depending on the stage of colorectal cancer. Therefore, the discovery of phytochemicals regulating the Nrf2/KEAP1 axis and Citation: Lee, D.-Y.; Song, M.-Y.; verification of their efficacy have attracted scientific attention. In this review, we summarize the role Kim, E.-H. Role of Oxidative Stress of oxidative stress and the Nrf2/KEAP1 signaling pathway in colorectal cancer, and the possible and Nrf2/KEAP1 Signaling in utility of phytochemicals with respect to the regulation of the Nrf2/KEAP1 axis in colorectal cancer. -

Chloroperoxidase Generation of Singlet a Molecular Oxygen
Proc. Natl. Acad. Sci. USA Vol. 80, pp. 5195-5197, September 1983 Biochemistry Chloroperoxidase generation of singlet A molecular oxygen observed directly by spectroscopy in the 1- to 1.6-pm region [(0,0) transition/chloride peroxidase/hydrogen peroxide/chloride ion] AHSAN U. KHAN*, PETER GEBAUERt, AND LOWELL P. HAGERt *Institute of Molecular Biophysics and Department of Chemistry, Florida State University, Tallahassee, Florida 32306; and tDepartment of Biochemistry, University of Illinois at Urbana-Champaign, Urbana, Illinois 61801 Communicated by-Michael Kasha, May 9; 1983 ABSTRACT The (0,o) 'Ag -* 3 singlet molecular oxygen ,.um region and based on a lead sulfide detector and the other, chemiluminescence emission from a biological reaction system-, a more sensitive- germanium photodiode detector, operating chloroperoxidase (EC 1.11.1.10) acting on hydrogen peroxide at between 1.00 and 1.60 Am. The advent of this instrumentation low pH (phosphate buffer, pH 1.85) with C¢1 as a cosubstirate, was (1-3) has opened up a window on the near-infrared spectral re- recorded with an ultrasensitive IR spectrometer. The strong gion so that luminescence studies of reactions suspected of chemiluminescence emission peak observed at 1.30 ,um provides generating intermediates with (possibly very weak) near-in- clear evidence of the enzymatic generation of (excited) singlet ox- frared emission can now be monitored directly, as has been done ygen from peroxide in this system. successfully with photosensitization by dye (1) and superoxide anion disproportionation (3). The present work used-the liquid The observation of 'Ag -- -' singlet molecular oxygen chemi- nitrogen-cooled photovoltaic intrinsic germanium detector, which luminescence emission generated by the chloroperoxidase- is described elsewhere (3). -

New Automatically Built Profiles for a Better Understanding of the Peroxidase Superfamily Evolution
University of Geneva Practical training report submitted for the Master Degree in Proteomics and Bioinformatics New automatically built profiles for a better understanding of the peroxidase superfamily evolution presented by Dominique Koua Supervisors: Dr Christophe DUNAND Dr Nicolas HULO Laboratory of Plant Physiology, Dr Christian J.A. SIGRIST University of Geneva Swiss Institute of Bioinformatics PROSITE group. Geneva, April, 18th 2008 Abstract Motivation: Peroxidases (EC 1.11.1.x), which are encoded by small or large multigenic families, are involved in several important physiological and developmental processes. These proteins are extremely widespread and present in almost all living organisms. An important number of haem and non-haem peroxidase sequences are annotated and classified in the peroxidase database PeroxiBase (http://peroxibase.isb-sib.ch). PeroxiBase contains about 5800 peroxidase sequences classified as haem peroxidases and non-haem peroxidases and distributed between thirteen superfamilies and fifty subfamilies, (Passardi et al., 2007). However, only a few classification tools are available for the characterisation of peroxidase sequences: InterPro motifs, PRINTS and specifically designed PROSITE profiles. However, these PROSITE profiles are very global and do not allow the differenciation between very close subfamily sequences nor do they allow the prediction of specific cellular localisations. Due to the rapid growth in the number of available sequences, there is a need for continual updates and corrections of peroxidase protein sequences as well as for new tools that facilitate acquisition and classification of existing and new sequences. Currently, the PROSITE generalised profile building manner and their usage do not allow the differentiation of sequences from subfamilies showing a high degree of similarity. -

Microspore Embryogenesis: Targeting the Determinant Factors of Stress- Induced Cell Reprogramming for Crop Improvement
1 2 3 4 Microspore embryogenesis: targeting the determinant factors of stress- 5 induced cell reprogramming for crop improvement 6 7 8 Pilar S. Testillano 9 10 Pollen Biotechnolgy of Crop Plants group. Biological Research Center, CIB-CSIC. 11 Ramiro de Maeztu 9, 28040 Madrid. Spain 12 E-mail: [email protected] 13 Phone: +34-918373112 (Ext: 4366) 14 15 Date of submission: October 31, 2018 16 Number of figures: 5. 17 Word count: 6312. 18 19 Running title: 20 Determinant factors of stress-induced microspore embryogenesis in crops 1 21 ABSTRACT 22 23 Under stress, isolated microspores are reprogrammed in vitro towards embryogenesis, 24 producing doubled haploid plants, useful biotechnological tools in plant breeding as a 25 source of new genetic variability, fixed in homozygous plants in only one generation. 26 Stress-induced cell death and low rates of cell reprogramming are major factors that 27 reduce the process yield. Knowledge gained in recent years has revealed that 28 microspore embryogenesis initiation and progression involve a complex network of 29 factors, whose roles are not yet well understood. Autophagy and cell-death proteases are 30 crucial players in the response to stress, while cell reprogramming and totipotency 31 acquisition are regulated by hormonal and epigenetic mechanisms. Auxin biosynthesis, 32 transport and action are required for microspore embryogenesis. Initial stages involve 33 DNA hypomethylation, H3K9 demethylation, and H3/H4 acetylation. Cell wall 34 remodelling, with pectin de-methylesterification and AGP expression, is necessary for 35 embryo development. We will review recent findings regarding the determinant factors 36 underlying stress-induced microspore embryogenesis, focusing on the role of 37 autophagy, cell death, auxin, chromatin modifications, and cell wall.