Characterization of Cytosolic Glutathione Peroxidase And
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

The Role of Glutathionperoxidase 4 (GPX4) in Hematopoiesis and Leukemia
The role of glutathionperoxidase 4 (GPX4) in hematopoiesis and leukemia von Kira Célénie Stahnke Inaugral- Dissertation zur Erlangung der Doktorwürde der TierärztliChen Fakultät der Ludwig-Maximilians-Universität MünChen The role of glutathionperoxidase 4 (GPX4) in hematopoiesis and leukemia von Kira Célénie Stahnke aus Dortmund München 2016 Aus dem VeterinärwissensChaftliChen Department der TierärztliChen Fakultät der Ludwig-Maximilians-Universität München Lehrstuhl für Molekulare Tierzucht und Biotechnologie Arbeit angefertigt unter der Leitung von Univ.-Prof. Dr. Eckhard Wolf Angefertigt am Institut für Experimentelle TumorforsChung Universitätsklinikum Ulm Mentor: Prof. Dr. Christian Buske GedruCkt mit Genehmigung der TierärztliChen Fakultät der Ludwig-Maximilians-Universität München Dekan: Univ.-Prof. Dr. JoaChim Braun Berichterstatter: Univ.-Prof. Dr. Eckard Wolf Korreferent/en: Priv.-Doz. Dr. Bianka Schulz Tag der Promotion: 06.02.2016 Table of contents List of Abbreviations..............................................................................2 1 Introduction ........................................................................................6 1.1 ROS and oxidative stress in physiology and pathology..................................................... 6 1.1.1 SourCes of Reactive Oxygen SpeCies..................................................................................................6 1.1.2 PhysiologiCal role of ROS........................................................................................................................8 -

(ER) Membrane Contact Sites (MCS) Uses Toxic Waste to Deliver Messages Edgar Djaha Yoboue1, Roberto Sitia1 and Thomas Simmen2
Yoboue et al. Cell Death and Disease (2018) 9:331 DOI 10.1038/s41419-017-0033-4 Cell Death & Disease REVIEW ARTICLE Open Access Redox crosstalk at endoplasmic reticulum (ER) membrane contact sites (MCS) uses toxic waste to deliver messages Edgar Djaha Yoboue1, Roberto Sitia1 and Thomas Simmen2 Abstract Many cellular redox reactions housed within mitochondria, peroxisomes and the endoplasmic reticulum (ER) generate hydrogen peroxide (H2O2) and other reactive oxygen species (ROS). The contribution of each organelle to the total cellular ROS production is considerable, but varies between cell types and also over time. Redox-regulatory enzymes are thought to assemble at a “redox triangle” formed by mitochondria, peroxisomes and the ER, assembling “redoxosomes” that sense ROS accumulations and redox imbalances. The redoxosome enzymes use ROS, potentially toxic by-products made by some redoxosome members themselves, to transmit inter-compartmental signals via chemical modifications of downstream proteins and lipids. Interestingly, important components of the redoxosome are ER chaperones and oxidoreductases, identifying ER oxidative protein folding as a key ROS producer and controller of the tri-organellar membrane contact sites (MCS) formed at the redox triangle. At these MCS, ROS accumulations could directly facilitate inter-organellar signal transmission, using ROS transporters. In addition, ROS influence the flux 2+ 2+ of Ca ions, since many Ca handling proteins, including inositol 1,4,5 trisphosphate receptors (IP3Rs), SERCA pumps or regulators of the mitochondrial Ca2+ uniporter (MCU) are redox-sensitive. Fine-tuning of these redox and ion signaling pathways might be difficult in older organisms, suggesting a dysfunctional redox triangle may accompany 1234567890 1234567890 the aging process. -

In Thyroid Cancer
Metere et al. Cancer Cell Int (2018) 18:7 https://doi.org/10.1186/s12935-018-0504-4 Cancer Cell International PRIMARY RESEARCH Open Access A possible role for selenoprotein glutathione peroxidase (GPx1) and thioredoxin reductases (TrxR1) in thyroid cancer: our experience in thyroid surgery Alessio Metere1* , Francesca Frezzotti1, Claire Elizabeth Graves2, Massimo Vergine1, Alessandro De Luca1, Donatella Pietraforte3 and Laura Giacomelli1 Abstract Background: Oxidative stress is responsible for some alterations in the chemical structure and, consequently, in the function of proteins, lipids, and DNA. Recent studies have linked oxidative stress to cancers, particularly thyroid cancer, but the mechanisms remain unclear. Here, we further characterize the role of oxidative stress in thyroid cancer by analyzing the expression of two selenium antioxidant molecules, glutathione peroxidase (GPx1) and thioredoxin reductase (TrxR1) in thyroid cancer cells. Methods: Samples of both healthy thyroid tissue and thyroid tumor were taken for analysis after total thyroidectomy. The expression of GPx1 and TrxR1 was revealed by Western blot analysis and quantifed by densitometric analy- ses, while the evaluation of free radicals was performed by Electron Paramagnetic Resonance (EPR)-spin trapping technique. Results: Our results show a decrease in the expression of GPx1 and TrxR1 ( 45.7 and 43.2% respectively, p < 0.01) in the thyroid cancer cells compared to the healthy cells. In addition, the EPR− technique− shows an increase of free radicals in tumor tissue, signifcantly higher than that found in healthy thyroid tissue ( 116.3%, p < 0.01). + Conclusions: Our fndings underscore the relationship between thyroid cancer and oxidative stress, showing the imbalance of the oxidant/antioxidant system in thyroid cancer tissue. -

A Machine Learning Tool for Predicting Protein Function Anna
The Woolf Classifier Building Pipeline: A Machine Learning Tool for Predicting Protein Function Anna Farrell-Sherman Submitted in Partial Fulfillment of the Prerequisite for Honors in Computer Science under the advisement of Eni Mustafaraj and Vanja Klepac-Ceraj May 2019 © 2019 Anna Farrell-Sherman Abstract Proteins are the machinery that allow cells to grow, reproduce, communicate, and create multicellular organisms. For all of their importance, scientists still have a hard time understanding the function of a protein based on its sequence alone. For my honors thesis in computer science, I created a machine learning tool that can predict the function of a protein based solely on its amino acid sequence. My tool gives scientists a structure in which to build a � Nearest Neighbor or random forest classifier to distinguish between proteins that can and cannot perform a given function. Using default Min-Max scaling, and the Matthews Correlation Coefficient for accuracy assessment, the Woolf Pipeline is built with simplified choices to guide users to success. i Acknowledgments There are so many people who made this thesis possible. First, thank you to my wonderful advisors, Eni and Vanja, who never gave up on me, and always pushed me to try my hardest. Thank you also to the other members of my committee, Sohie Lee, Shikha Singh, and Rosanna Hertz for supporting me through this process. To Kevin, Sophie R, Sophie E, and my sister Phoebe, thank you for reading my drafts, and advising me when the going got tough. To all my wonderful friends, who were always there with encouraging words, warm hugs, and many congratulations, I cannot thank you enough. -

Illness-Induced Changes in Thyroid Hormone Metabolism: Focus on the Tissue Level
r e V i e W illness-induced changes in thyroid hormone metabolism: focus on the tissue level J. Kwakkel*, E. Fliers, A. Boelen Department of Endocrinology & Metabolism, Academic Medical Center, University of Amsterdam, the Netherlands, *corresponding author: tel.: +31 (0)20-566 67 01, fax: +31 (0)20-691 76 82, e-mail: [email protected] a b s t r a C t during illness changes in thyroid hormone metabolism ring and the outer (tyrosyl) ring of T4 can be deiodinated, occur, collectively known as the non-thyroidal illness ultimately leading to the formation of 3,3’-di-iodothyronine syndrome (NTIS). NTIS is characterised by low serum (T2) (figure 1). thyroid hormone levels without the expected rise in serum thyroid-stimulating hormone, indicating a major change in thyroid hormone feedback regulation. recent studies n o n - t H yroidal illness syndro M e have made clear that during NTIS differential changes in thyroid hormone metabolism occur in various tissues, the During illness many aspects of thyroid hormone net effect of which may be either activation or inhibition of metabolism change, collectively known as the thyroid hormone action. in this review we discuss systemic non-thyroidal illness syndrome (NTIS). The hallmark of and local changes in thyroid hormone metabolism during NTIS is decreased serum thyroid hormone levels without illness, highlighting their physiological implications in an increase in TSH and TRH expression, indicating terms of disease course. the absence of negative feedback regulation. This may represent a useful adaptation of the body to counteract excessive catabolism observed during illness and can be K e y W o r d s viewed as a part of the acute phase response.4 However, especially during prolonged critical illness in the ICU Deiodinase, inflammation, non-thyroidal illness syndrome, setting NTIS may be maladaptative.5 thyroid hormone figure 1. -
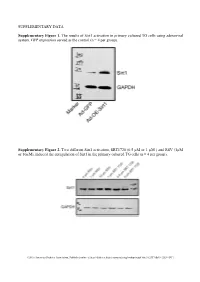
SUPPLEMENTARY DATA Supplementary Figure 1. The
SUPPLEMENTARY DATA Supplementary Figure 1. The results of Sirt1 activation in primary cultured TG cells using adenoviral system. GFP expression served as the control (n = 4 per group). Supplementary Figure 2. Two different Sirt1 activators, SRT1720 (0.5 µM or 1 µM ) and RSV (1µM or 10µM), induced the upregulation of Sirt1 in the primary cultured TG cells (n = 4 per group). ©2016 American Diabetes Association. Published online at http://diabetes.diabetesjournals.org/lookup/suppl/doi:10.2337/db15-1283/-/DC1 SUPPLEMENTARY DATA Supplementary Table 1. Primers used in qPCR Gene Name Primer Sequences Product Size (bp) Sirt1 F: tgccatcatgaagccagaga 241 (NM_001159589) R: aacatcgcagtctccaagga NOX4 F: tgtgcctttattgtgcggag 172 (NM_001285833.1) R: gctgatacactggggcaatg Supplementary Table 2. Antibodies used in Western blot or Immunofluorescence Antibody Company Cat. No Isotype Dilution Sirt1 Santa Cruz * sc-15404 Rabbit IgG 1/200 NF200 Sigma** N5389 Mouse IgG 1/500 Tubulin R&D# MAB1195 Mouse IgG 1/500 NOX4 Abcam† Ab133303 Rabbit IgG 1/500 NOX2 Abcam Ab129068 Rabbit IgG 1/500 phospho-AKT CST‡ #4060 Rabbit IgG 1/500 EGFR CST #4267 Rabbit IgG 1/500 Ki67 Santa Cruz sc-7846 Goat IgG 1/500 * Santa Cruz Biotechnology, Santa Cruz, CA, USA ** Sigma aldrich, Shanghai, China # R&D Systems Inc, Minneapolis, MN, USA † Abcam, Inc., Cambridge, MA, USA ‡ Cell Signaling Technology, Inc., Danvers, MA, USA ©2016 American Diabetes Association. Published online at http://diabetes.diabetesjournals.org/lookup/suppl/doi:10.2337/db15-1283/-/DC1 SUPPLEMENTARY DATA Supplementary -

03-232 Biochemistry Exam II – 2012 -Key Name:______Instructions: This Exam Consists of 100 Points on 6 Pages
03-232 Biochemistry Exam II – 2012 -Key Name:________________________ Instructions: This exam consists of 100 points on 6 pages. Please use the space provided to answer the question or the back of the preceding page. In questions with choices, all your answers will be graded and you will receive the best grade. Allot 1 min/2 points. A B 1. (10 pts) One protein (protein A) binds naphthalene (ligand) by sandwiching it between two tryptophan residues. Another protein Trp L Trp Tyr L Tyr (protein B) also binds naphthalene by sandwiching it between two OH tyrosine residues, however the binding is weaker. The structure of the H protein-ligand complex is shown on the right (side view), the structures N of tryptophan, naphthalene (ligand), and tyrosine are shown as well. i) What are the principal energetic factor(s) that are responsible for binding of naphthalene to these proteins? (4 pts) Tryptophan(Trp) naphthalene (L) Tyrosine (Tyr) ii) Why does the tyrosine containing protein show weaker binding? (4 pts) ii) Which kinetic rate constant, the off-rate (kOFF) or the on-rate (kON), would be most different between the two proteins? In what way would it differ? Why? (2 pts) i) van der waals (H) and the hydrophobic effect (S) (3 pts for one, 4 points for two). ii) van der waals is reduce because the contact between the tyrosine and the ligand will be smaller. There will be a smaller hydrophobic effect as well because the non-polar surface area of tyrosine is smaller than tryptophan. Therefore in protein B, a smaller number of ordered water molecules will be released. -

GPX4 at the Crossroads of Lipid Homeostasis and Ferroptosis Giovanni C
REVIEW GPX4 www.proteomics-journal.com GPX4 at the Crossroads of Lipid Homeostasis and Ferroptosis Giovanni C. Forcina and Scott J. Dixon* formation of toxic radicals (e.g., R-O•).[5] Oxygen is necessary for aerobic metabolism but can cause the harmful The eight mammalian GPX proteins fall oxidation of lipids and other macromolecules. Oxidation of cholesterol and into three clades based on amino acid phospholipids containing polyunsaturated fatty acyl chains can lead to lipid sequence similarity: GPX1 and GPX2; peroxidation, membrane damage, and cell death. Lipid hydroperoxides are key GPX3, GPX5, and GPX6; and GPX4, GPX7, and GPX8.[6] GPX1–4 and 6 (in intermediates in the process of lipid peroxidation. The lipid hydroperoxidase humans) are selenoproteins that contain glutathione peroxidase 4 (GPX4) converts lipid hydroperoxides to lipid an essential selenocysteine in the enzyme + alcohols, and this process prevents the iron (Fe2 )-dependent formation of active site, while GPX5, 6 (in mouse and toxic lipid reactive oxygen species (ROS). Inhibition of GPX4 function leads to rats), 7, and 8 use an active site cysteine lipid peroxidation and can result in the induction of ferroptosis, an instead. Unlike other family members, GPX4 (PHGPx) can act as a phospholipid iron-dependent, non-apoptotic form of cell death. This review describes the hydroperoxidase to reduce lipid perox- formation of reactive lipid species, the function of GPX4 in preventing ides to lipid alcohols.[7,8] Thus,GPX4ac- oxidative lipid damage, and the link between GPX4 dysfunction, lipid tivity is essential to maintain lipid home- oxidation, and the induction of ferroptosis. ostasis in the cell, prevent the accumula- tion of toxic lipid ROS and thereby block the onset of an oxidative, iron-dependent, non-apoptotic mode of cell death termed 1. -

Type 3 Lodothyronine Deiodinase: Cloning, in Vitro Expression, and Functional Analysis of the Placental Selenoenzyme
Type 3 lodothyronine deiodinase: cloning, in vitro expression, and functional analysis of the placental selenoenzyme. D Salvatore, … , D L St Germain, P R Larsen J Clin Invest. 1995;96(5):2421-2430. https://doi.org/10.1172/JCI118299. Research Article Type 3 iodothyronine deiodinase (D3) catalyzes the conversion of T4 and T3 to inactive metabolites. It is highly expressed in placenta and thus can regulate circulating fetal thyroid hormone concentrations throughout gestation. We have cloned and expressed a 2.1-kb human placental D3 cDNA which encodes a 32-kD protein with a Km of 1.2 nM for 5 deiodination of T3 and 340 nM for 5' deiodination of reverse T3. The reaction requires DTT and is not inhibited by 6n- propylthiouracil. We quantitated transiently expressed D3 by specifically labeling the protein with bromoacetyl [125I]T3. The Kcat/Km ratio for 5 deiodination of T3 was over 1,000-fold that for 5' deiodination of reverse T3. Human D3 is a selenoenzyme as evidenced by (a) the presence of an in frame UGA codon at position 144, (b) the synthesis of a 32-kD 75Se-labeled protein in D3 cDNA transfected cells, and (c) the presence of a selenocysteine insertion sequence element in the 3' untranslated region of the mRNA which is required for its expression. The D3 selenocysteine insertion sequence element is more potent than that in the type 1 deiodinase or glutathione peroxidase gene, suggesting a high priority for selenocysteine incorporation into this enzyme. The conservation of this enzyme from Xenopus laevis tadpoles to humans implies an essential role for regulation of thyroid hormone inactivation during embryological development. -

Thiol Peroxidases Mediate Specific Genome-Wide Regulation of Gene Expression in Response to Hydrogen Peroxide
Thiol peroxidases mediate specific genome-wide regulation of gene expression in response to hydrogen peroxide Dmitri E. Fomenkoa,1,2, Ahmet Koca,1, Natalia Agishevaa, Michael Jacobsena,b, Alaattin Kayaa,c, Mikalai Malinouskia,c, Julian C. Rutherfordd, Kam-Leung Siue, Dong-Yan Jine, Dennis R. Winged, and Vadim N. Gladysheva,c,2 aDepartment of Biochemistry, University of Nebraska, Lincoln, NE 68588-0664; bDepartment of Life Sciences, Wayne State College, Wayne, NE 68787; dDepartment of Medicine, University of Utah Health Sciences Center, Salt Lake City, UT 84132; eDepartment of Biochemistry, University of Hong Kong, Hong Kong, China; and cDivision of Genetics, Department of Medicine, Brigham and Women’s Hospital and Harvard Medical School, Boston, MA 02115 Edited by Joan Selverstone Valentine, University of California, Los Angeles, CA, and approved December 22, 2010 (received for review July 21, 2010) Hydrogen peroxide is thought to regulate cellular processes by and could withstand significant oxidative stress. It responded to direct oxidation of numerous cellular proteins, whereas antioxi- several redox stimuli by robust transcriptional reprogramming. dants, most notably thiol peroxidases, are thought to reduce However, it was unable to transcriptionally respond to hydrogen peroxides and inhibit H2O2 response. However, thiol peroxidases peroxide. The data suggested that thiol peroxidases transfer have also been implicated in activation of transcription factors oxidative signals from peroxides to target proteins, thus activating and signaling. It remains unclear if these enzymes stimulate or various transcriptional programs. This study revealed a previously inhibit redox regulation and whether this regulation is widespread undescribed function of these proteins, in addition to their roles or limited to a few cellular components. -

Micrornas As New Regulators of Neutrophil Extracellular Trap Formation
International Journal of Molecular Sciences Review MicroRNAs as New Regulators of Neutrophil Extracellular Trap Formation Sonia Águila † , Ascensión M. de los Reyes-García †, María P. Fernández-Pérez, Laura Reguilón-Gallego, Laura Zapata-Martínez, Inmaculada Ruiz-Lorente, Vicente Vicente , Rocío González-Conejero *,‡ and Constantino Martínez *,‡ Department of Hematology and Medical Oncology, Morales Meseguer University Hospital, Centro Regional de Hemodonación, Universidad de Murcia, IMIB, C/Ronda de Garay S/N, 30003 Murcia, Spain; [email protected] (S.Á.); [email protected] (A.M.d.l.R.-G.); [email protected] (M.P.F.-P.); [email protected] (L.R.-G.); [email protected] (L.Z.-M.); [email protected] (I.R.-L.); [email protected] (V.V.) * Correspondence: [email protected] (R.G.-C.); [email protected] (C.M.); Tel.: +34-968341990 (R.G.-C. & C.M.); Fax: +34-968261914 (R.G.-C. & C.M.) † These authors contributed equally to this work. ‡ These authors shared senior authorship. Abstract: Neutrophil extracellular traps (NETs) are formed after neutrophils expelled their chromatin content in order to primarily capture and eliminate pathogens. However, given their characteristics due in part to DNA and different granular proteins, NETs may induce a procoagulant response linking inflammation and thrombosis. Unraveling NET formation molecular mechanisms as well Citation: Águila, S.; de los as the intracellular elements that regulate them is relevant not only for basic knowledge but also Reyes-García, A.M.; Fernández-Pérez, to design diagnostic and therapeutic tools that may prevent their deleterious effects observed in M.P.; Reguilón-Gallego, L.; several inflammatory pathologies (e.g., cardiovascular and autoimmune diseases, cancer). -

Discovery of Oxidative Enzymes for Food Engineering. Tyrosinase and Sulfhydryl Oxi- Dase
Dissertation VTT PUBLICATIONS 763 1,0 0,5 Activity 0,0 2 4 6 8 10 pH Greta Faccio Discovery of oxidative enzymes for food engineering Tyrosinase and sulfhydryl oxidase VTT PUBLICATIONS 763 Discovery of oxidative enzymes for food engineering Tyrosinase and sulfhydryl oxidase Greta Faccio Faculty of Biological and Environmental Sciences Department of Biosciences – Division of Genetics ACADEMIC DISSERTATION University of Helsinki Helsinki, Finland To be presented for public examination with the permission of the Faculty of Biological and Environmental Sciences of the University of Helsinki in Auditorium XII at the University of Helsinki, Main Building, Fabianinkatu 33, on the 31st of May 2011 at 12 o’clock noon. ISBN 978-951-38-7736-1 (soft back ed.) ISSN 1235-0621 (soft back ed.) ISBN 978-951-38-7737-8 (URL: http://www.vtt.fi/publications/index.jsp) ISSN 1455-0849 (URL: http://www.vtt.fi/publications/index.jsp) Copyright © VTT 2011 JULKAISIJA – UTGIVARE – PUBLISHER VTT, Vuorimiehentie 5, PL 1000, 02044 VTT puh. vaihde 020 722 111, faksi 020 722 4374 VTT, Bergsmansvägen 5, PB 1000, 02044 VTT tel. växel 020 722 111, fax 020 722 4374 VTT Technical Research Centre of Finland, Vuorimiehentie 5, P.O. Box 1000, FI-02044 VTT, Finland phone internat. +358 20 722 111, fax + 358 20 722 4374 Edita Prima Oy, Helsinki 2011 2 Greta Faccio. Discovery of oxidative enzymes for food engineering. Tyrosinase and sulfhydryl oxi- dase. Espoo 2011. VTT Publications 763. 101 p. + app. 67 p. Keywords genome mining, heterologous expression, Trichoderma reesei, Aspergillus oryzae, sulfhydryl oxidase, tyrosinase, catechol oxidase, wheat dough, ascorbic acid Abstract Enzymes offer many advantages in industrial processes, such as high specificity, mild treatment conditions and low energy requirements.