Identification and Validation of a Blood-Based 18-Gene Expression Signature in Colorectal Cancer
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

The Cargo Protein MAP17 (PDZK1IP1) Regulates the Immune Microenvironment
www.impactjournals.com/oncotarget/ Oncotarget, 2017, Vol. 8, (No. 58), pp: 98580-98597 Research Paper The cargo protein MAP17 (PDZK1IP1) regulates the immune microenvironment José M. García-Heredia1,2,3 and Amancio Carnero1,3 1Instituto de Biomedicina de Sevilla, IBIS/Hospital Universitario Virgen del Rocío/Universidad de Sevilla/Consejo Superior de Investigaciones Científicas, Seville, Spain 2Department of Vegetal Biochemistry and Molecular Biology, University of Seville, Seville, Spain 3CIBER de Cáncer, Instituto de Salud Carlos III, Madrid, Spain Correspondence to: Amancio Carnero, email: [email protected] Keywords: MAP17; oncogene; inflammation; cancer; inflammatory diseases Received: July 14, 2017 Accepted: August 25, 2017 Published: October 06, 2017 Copyright: García-Heredia et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License 3.0 (CC BY 3.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited. ABSTRACT Inflammation is a complex defensive response activated after various harmful stimuli allowing the clearance of damaged cells and initiating healing and regenerative processes. Chronic, or pathological, inflammation is also one of the causes of neoplastic transformation and cancer development. MAP17 is a cargo protein that transports membrane proteins from the endoplasmic reticulum. Therefore, its overexpression may be linked to an excess of membrane proteins that may be recognized as an unwanted signal, triggering local inflammation. Therefore, we analyzed whether its overexpression is related to an inflammatory phenotype. In this work, we found a correlation between MAP17 expression and inflammatory phenotype in tumors and in other inflammatory diseases such as Crohn's disease, Barrett's esophagus, COPD or psoriasis. -

Genetic and Genomic Analysis of Hyperlipidemia, Obesity and Diabetes Using (C57BL/6J × TALLYHO/Jngj) F2 Mice
University of Tennessee, Knoxville TRACE: Tennessee Research and Creative Exchange Nutrition Publications and Other Works Nutrition 12-19-2010 Genetic and genomic analysis of hyperlipidemia, obesity and diabetes using (C57BL/6J × TALLYHO/JngJ) F2 mice Taryn P. Stewart Marshall University Hyoung Y. Kim University of Tennessee - Knoxville, [email protected] Arnold M. Saxton University of Tennessee - Knoxville, [email protected] Jung H. Kim Marshall University Follow this and additional works at: https://trace.tennessee.edu/utk_nutrpubs Part of the Animal Sciences Commons, and the Nutrition Commons Recommended Citation BMC Genomics 2010, 11:713 doi:10.1186/1471-2164-11-713 This Article is brought to you for free and open access by the Nutrition at TRACE: Tennessee Research and Creative Exchange. It has been accepted for inclusion in Nutrition Publications and Other Works by an authorized administrator of TRACE: Tennessee Research and Creative Exchange. For more information, please contact [email protected]. Stewart et al. BMC Genomics 2010, 11:713 http://www.biomedcentral.com/1471-2164/11/713 RESEARCH ARTICLE Open Access Genetic and genomic analysis of hyperlipidemia, obesity and diabetes using (C57BL/6J × TALLYHO/JngJ) F2 mice Taryn P Stewart1, Hyoung Yon Kim2, Arnold M Saxton3, Jung Han Kim1* Abstract Background: Type 2 diabetes (T2D) is the most common form of diabetes in humans and is closely associated with dyslipidemia and obesity that magnifies the mortality and morbidity related to T2D. The genetic contribution to human T2D and related metabolic disorders is evident, and mostly follows polygenic inheritance. The TALLYHO/ JngJ (TH) mice are a polygenic model for T2D characterized by obesity, hyperinsulinemia, impaired glucose uptake and tolerance, hyperlipidemia, and hyperglycemia. -

Aneuploidy: Using Genetic Instability to Preserve a Haploid Genome?
Health Science Campus FINAL APPROVAL OF DISSERTATION Doctor of Philosophy in Biomedical Science (Cancer Biology) Aneuploidy: Using genetic instability to preserve a haploid genome? Submitted by: Ramona Ramdath In partial fulfillment of the requirements for the degree of Doctor of Philosophy in Biomedical Science Examination Committee Signature/Date Major Advisor: David Allison, M.D., Ph.D. Academic James Trempe, Ph.D. Advisory Committee: David Giovanucci, Ph.D. Randall Ruch, Ph.D. Ronald Mellgren, Ph.D. Senior Associate Dean College of Graduate Studies Michael S. Bisesi, Ph.D. Date of Defense: April 10, 2009 Aneuploidy: Using genetic instability to preserve a haploid genome? Ramona Ramdath University of Toledo, Health Science Campus 2009 Dedication I dedicate this dissertation to my grandfather who died of lung cancer two years ago, but who always instilled in us the value and importance of education. And to my mom and sister, both of whom have been pillars of support and stimulating conversations. To my sister, Rehanna, especially- I hope this inspires you to achieve all that you want to in life, academically and otherwise. ii Acknowledgements As we go through these academic journeys, there are so many along the way that make an impact not only on our work, but on our lives as well, and I would like to say a heartfelt thank you to all of those people: My Committee members- Dr. James Trempe, Dr. David Giovanucchi, Dr. Ronald Mellgren and Dr. Randall Ruch for their guidance, suggestions, support and confidence in me. My major advisor- Dr. David Allison, for his constructive criticism and positive reinforcement. -

Supplementary Materials
Supplementary materials Supplementary Table S1: MGNC compound library Ingredien Molecule Caco- Mol ID MW AlogP OB (%) BBB DL FASA- HL t Name Name 2 shengdi MOL012254 campesterol 400.8 7.63 37.58 1.34 0.98 0.7 0.21 20.2 shengdi MOL000519 coniferin 314.4 3.16 31.11 0.42 -0.2 0.3 0.27 74.6 beta- shengdi MOL000359 414.8 8.08 36.91 1.32 0.99 0.8 0.23 20.2 sitosterol pachymic shengdi MOL000289 528.9 6.54 33.63 0.1 -0.6 0.8 0 9.27 acid Poricoic acid shengdi MOL000291 484.7 5.64 30.52 -0.08 -0.9 0.8 0 8.67 B Chrysanthem shengdi MOL004492 585 8.24 38.72 0.51 -1 0.6 0.3 17.5 axanthin 20- shengdi MOL011455 Hexadecano 418.6 1.91 32.7 -0.24 -0.4 0.7 0.29 104 ylingenol huanglian MOL001454 berberine 336.4 3.45 36.86 1.24 0.57 0.8 0.19 6.57 huanglian MOL013352 Obacunone 454.6 2.68 43.29 0.01 -0.4 0.8 0.31 -13 huanglian MOL002894 berberrubine 322.4 3.2 35.74 1.07 0.17 0.7 0.24 6.46 huanglian MOL002897 epiberberine 336.4 3.45 43.09 1.17 0.4 0.8 0.19 6.1 huanglian MOL002903 (R)-Canadine 339.4 3.4 55.37 1.04 0.57 0.8 0.2 6.41 huanglian MOL002904 Berlambine 351.4 2.49 36.68 0.97 0.17 0.8 0.28 7.33 Corchorosid huanglian MOL002907 404.6 1.34 105 -0.91 -1.3 0.8 0.29 6.68 e A_qt Magnogrand huanglian MOL000622 266.4 1.18 63.71 0.02 -0.2 0.2 0.3 3.17 iolide huanglian MOL000762 Palmidin A 510.5 4.52 35.36 -0.38 -1.5 0.7 0.39 33.2 huanglian MOL000785 palmatine 352.4 3.65 64.6 1.33 0.37 0.7 0.13 2.25 huanglian MOL000098 quercetin 302.3 1.5 46.43 0.05 -0.8 0.3 0.38 14.4 huanglian MOL001458 coptisine 320.3 3.25 30.67 1.21 0.32 0.9 0.26 9.33 huanglian MOL002668 Worenine -

Hereditary Duplication of Proximal Chromosome 1Q (Q11q22) in A
1of4 ELECTRONIC LETTER J Med Genet: first published as 10.1136/jmg.39.12.e79 on 1 December 2002. Downloaded from Hereditary duplication of proximal chromosome 1q (q11q22) in a patient with T lymphoblastic lymphoma/leukaemia: a family study using G banding and comparative genomic hybridisation N P H Chan,MHLNg,SHCheng, V Lee, K S Tsang,TTLau,CKLi ............................................................................................................................. J Med Genet 2002;39:e79(http://www.jmedgenet.com/cgi/content/full/39/12/e79) berrations of the long arm of chromosome 1 have been Key points linked to many diseases, both in acquired somatic neo- Aplastic conditions and constitutional genetic disorders. These abnormalities include structural rearrangements and • We report a case of inherited proximal duplication of various deletions or amplifications affecting whole chromo- chromosome 1q in an 11 year old boy presenting with some arms or specific regions. Gain of chromosome 1q has T lymphoblastic lymphoma/leukaemia. The duplication been reported in many solid tumours, including invasive car- of chromosome 1 (q11q22) was identified using G cinomas of the breast, cervical cancers, renal cell carcinoma, banding and confirmed with comparative genomic sarcomas, and hepatocellular carcinomas.1–6 Complete or hybridisation. partial trisomy of 1q has been reported in cases of acute • Cytogenetic studies performed in his family members myeloid leukaemia, myelodysplastic syndrome, myeloprolif- showed the same chromosomal aberration in his mother erative diseases, Fanconi anaemia, and B lineage lymphomas, and sister, but not in his older brother who had a normal where involvement of 1q21 is frequently observed.7–14 karyotype. No dysmorphic syndromes, congenital mal- formations, or psychomotor retardation were observed CASE REPORT in either the index patient or his mother or sister. -

Placental Transcriptome Adaptations to Maternal Nutrient Restriction in Sheep
International Journal of Molecular Sciences Article Placental Transcriptome Adaptations to Maternal Nutrient Restriction in Sheep Chelsie B. Steinhauser 1,†, Colleen A. Lambo 2,†, Katharine Askelson 1, Gregory W. Burns 3, Susanta K. Behura 4,5, Thomas E. Spencer 4, Fuller W. Bazer 1 and Michael Carey Satterfield 1,* 1 Department of Animal Science, Texas A & M University, College Station, TX 77843, USA; [email protected] (C.B.S.); [email protected] (K.A.); [email protected] (F.W.B.) 2 Department of Veterinary Physiology and Pharmacology, Texas A & M University, College Station, TX 77843, USA; [email protected] 3 Department of Obstetrics, Gynecology and Reproductive Biology, Michigan State University, Grand Rapids, MI 49503, USA; [email protected] 4 Division of Animal Sciences, University of Missouri, Columbia, MO 65211, USA; [email protected] (S.K.B.); [email protected] (T.E.S.) 5 Institute for Data Science and Informatics, University of Missouri, Columbia, MO 65211, USA * Correspondence: csatterfi[email protected]; Tel.: +1-979-845-6448 † These authors contributed equally to this work. Abstract: Placental development is modified in response to maternal nutrient restriction (NR), resulting in a spectrum of fetal growth rates. Pregnant sheep carrying singleton fetuses and fed either 100% (n = 8) or 50% (NR; n = 28) of their National Research Council (NRC) recommended intake from days 35–135 of pregnancy were used to elucidate placentome transcriptome alterations at both Citation: Steinhauser, C.B.; Lambo, day 70 and day 135. NR fetuses were further designated into upper (NR NonSGA; n = 7) and lower C.A.; Askelson, K.; Burns, G.W.; quartiles (NR SGA; n = 7) based on day 135 fetal weight. -

Gene Expression Analysis of Mevalonate Kinase Deficiency
International Journal of Environmental Research and Public Health Article Gene Expression Analysis of Mevalonate Kinase Deficiency Affected Children Identifies Molecular Signatures Related to Hematopoiesis Simona Pisanti * , Marianna Citro , Mario Abate , Mariella Caputo and Rosanna Martinelli * Department of Medicine, Surgery and Dentistry ‘Scuola Medica Salernitana’, University of Salerno, Via Salvatore Allende, 84081 Baronissi (SA), Italy; [email protected] (M.C.); [email protected] (M.A.); [email protected] (M.C.) * Correspondence: [email protected] (S.P.); [email protected] (R.M.) Abstract: Mevalonate kinase deficiency (MKD) is a rare autoinflammatory genetic disorder charac- terized by recurrent fever attacks and systemic inflammation with potentially severe complications. Although it is recognized that the lack of protein prenylation consequent to mevalonate pathway blockade drives IL1β hypersecretion, and hence autoinflammation, MKD pathogenesis and the molecular mechanisms underlaying most of its clinical manifestations are still largely unknown. In this study, we performed a comprehensive bioinformatic analysis of a microarray dataset of MKD patients, using gene ontology and Ingenuity Pathway Analysis (IPA) tools, in order to identify the most significant differentially expressed genes and infer their predicted relationships into biological processes, pathways, and networks. We found that hematopoiesis linked biological functions and pathways are predominant in the gene ontology of differentially expressed genes in MKD, in line with the observed clinical feature of anemia. We also provided novel information about the molecular Citation: Pisanti, S.; Citro, M.; Abate, M.; Caputo, M.; Martinelli, R. mechanisms at the basis of the hematological abnormalities observed, that are linked to the chronic Gene Expression Analysis of inflammation and to defective prenylation. -

MAP17's Up-Regulation, a Crosspoint in Cancer and Inflammatory
García-Heredia and Carnero Molecular Cancer (2018) 17:80 https://doi.org/10.1186/s12943-018-0828-7 REVIEW Open Access Dr. Jekyll and Mr. Hyde: MAP17’s up- regulation, a crosspoint in cancer and inflammatory diseases José M. García-Heredia1,2,3 and Amancio Carnero1,3* Abstract Inflammation is a common defensive response that is activated after different harmful stimuli. This chronic, or pathological, inflammation is also one of the causes of neoplastic transformation and cancer development. MAP17 is a small protein localized to membranes with a restricted pattern of expression in adult tissues. However, its expression is common in destabilized cells, as it is overexpressed both in inflammatory diseases and in cancer. MAP17 is overexpressed in most, if not all, carcinomas and in many tumors of mesenchymal origin, and correlates with higher grade and poorly differentiated tumors. This overexpression drives deep changes in cell homeostasis including increased oxidative stress, deregulation of signaling pathways and increased growth rates. Importantly, MAP17 is associated in tumors with inflammatory cells infiltration, not only in cancer but in various inflammatory diseases such as Barret’s esophagus, lupus, Crohn’s, psoriasis and COPD. Furthermore, MAP17 also modifies the expression of genes connected to inflammation, showing a clear induction of the inflammatory profile. Since MAP17 appears highly correlated with the infiltration of inflammatory cells in cancer, is MAP17 overexpression an important cellular event connecting tumorigenesis and inflammation? Keywords: MAP17, Cancer, Inflammatory diseases Background process ends when the activated cells undergo apoptosis in Inflammatory response is a common defensive process acti- a highly regulated process that finishes after pathogens and vated after different harmful stimuli, constituting a highly cell debris have been phagocytized [9]. -

Targeted Shrna Screening Identified Critical Roles of Pleckstrin-2 in Erythropoiesis
Hematopoiesis SUPPLEMENTARY APPENDIX Targeted shRNA screening identified critical roles of pleckstrin-2 in erythropoiesis Baobing Zhao,1* Ganesan Keerthivasan,1* Yang Mei,1 Jing Yang,1 James McElherne,1 Piu Wong,2 John G. Doench,3 Gang Feng,4 David E. Root,3 and Peng Ji1 1Department of Pathology, Feinberg School of Medicine, Northwestern University, Chicago, IL; 2Whitehead Institute for Biomedical Research, Cambridge, MA; 3Broad Institute of Harvard University and the Massachusetts Institute of Technology, Cambridge, MA; and 4Biomedical Informatics Center, Northwestern University, Chicago, IL, USA *BZ and GK contributed equally to this work. ©2014 Ferrata Storti Foundation. This is an open-access paper. doi:10.3324/haematol.2014.105809 Manuscript received on February 12, 2014. Manuscript accepted on April 14, 2014. Correspondence: [email protected] Supplemental Methods Purification and culture of fetal liver cells for targeted screening Fetal liver cells were isolated from E13.5 C57BL/6 embryos and mechanically dissociated by pipetting in PBS containing 10% fetal bovine serum (GIBCO). Single-cell suspensions were prepared by passing the dissociated cells through 40 µm cell strainers (BD Biosciences). Total fetal liver cells were labeled with biotin-conjugated anti-TER119 antibody (1:100) (eBioscience), and TER119 negative cells were purified using EasySep column free cell isolation system according to the manufacturer’s instructions (Stem Cell Technologies). Purified cells were seeded in round bottom 96 well plates at a cell density of 1x103 cells/200µl media/well. The purified cells were infected by distinct lentiviruses encoding shRNAs against selected genes in each well of the 96-well plate. The infection strategies were schematically illustrated in Figure 1A. -
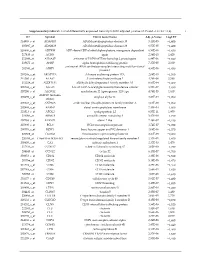
Supplementary Table S1. List of Differentially Expressed
Supplementary table S1. List of differentially expressed transcripts (FDR adjusted p‐value < 0.05 and −1.4 ≤ FC ≥1.4). 1 ID Symbol Entrez Gene Name Adj. p‐Value Log2 FC 214895_s_at ADAM10 ADAM metallopeptidase domain 10 3,11E‐05 −1,400 205997_at ADAM28 ADAM metallopeptidase domain 28 6,57E‐05 −1,400 220606_s_at ADPRM ADP‐ribose/CDP‐alcohol diphosphatase, manganese dependent 6,50E‐06 −1,430 217410_at AGRN agrin 2,34E‐10 1,420 212980_at AHSA2P activator of HSP90 ATPase homolog 2, pseudogene 6,44E‐06 −1,920 219672_at AHSP alpha hemoglobin stabilizing protein 7,27E‐05 2,330 aminoacyl tRNA synthetase complex interacting multifunctional 202541_at AIMP1 4,91E‐06 −1,830 protein 1 210269_s_at AKAP17A A‐kinase anchoring protein 17A 2,64E‐10 −1,560 211560_s_at ALAS2 5ʹ‐aminolevulinate synthase 2 4,28E‐06 3,560 212224_at ALDH1A1 aldehyde dehydrogenase 1 family member A1 8,93E‐04 −1,400 205583_s_at ALG13 ALG13 UDP‐N‐acetylglucosaminyltransferase subunit 9,50E‐07 −1,430 207206_s_at ALOX12 arachidonate 12‐lipoxygenase, 12S type 4,76E‐05 1,630 AMY1C (includes 208498_s_at amylase alpha 1C 3,83E‐05 −1,700 others) 201043_s_at ANP32A acidic nuclear phosphoprotein 32 family member A 5,61E‐09 −1,760 202888_s_at ANPEP alanyl aminopeptidase, membrane 7,40E‐04 −1,600 221013_s_at APOL2 apolipoprotein L2 6,57E‐11 1,600 219094_at ARMC8 armadillo repeat containing 8 3,47E‐08 −1,710 207798_s_at ATXN2L ataxin 2 like 2,16E‐07 −1,410 215990_s_at BCL6 BCL6 transcription repressor 1,74E‐07 −1,700 200776_s_at BZW1 basic leucine zipper and W2 domains 1 1,09E‐06 −1,570 222309_at -

1 Novel Expression Signatures Identified by Transcriptional Analysis
ARD Online First, published on October 7, 2009 as 10.1136/ard.2009.108043 Ann Rheum Dis: first published as 10.1136/ard.2009.108043 on 7 October 2009. Downloaded from Novel expression signatures identified by transcriptional analysis of separated leukocyte subsets in SLE and vasculitis 1Paul A Lyons, 1Eoin F McKinney, 1Tim F Rayner, 1Alexander Hatton, 1Hayley B Woffendin, 1Maria Koukoulaki, 2Thomas C Freeman, 1David RW Jayne, 1Afzal N Chaudhry, and 1Kenneth GC Smith. 1Cambridge Institute for Medical Research and Department of Medicine, Addenbrooke’s Hospital, Hills Road, Cambridge, CB2 0XY, UK 2Roslin Institute, University of Edinburgh, Roslin, Midlothian, EH25 9PS, UK Correspondence should be addressed to Dr Paul Lyons or Prof Kenneth Smith, Department of Medicine, Cambridge Institute for Medical Research, Addenbrooke’s Hospital, Hills Road, Cambridge, CB2 0XY, UK. Telephone: +44 1223 762642, Fax: +44 1223 762640, E-mail: [email protected] or [email protected] Key words: Gene expression, autoimmune disease, SLE, vasculitis Word count: 2,906 The Corresponding Author has the right to grant on behalf of all authors and does grant on behalf of all authors, an exclusive licence (or non-exclusive for government employees) on a worldwide basis to the BMJ Publishing Group Ltd and its Licensees to permit this article (if accepted) to be published in Annals of the Rheumatic Diseases and any other BMJPGL products to exploit all subsidiary rights, as set out in their licence (http://ard.bmj.com/ifora/licence.pdf). http://ard.bmj.com/ on September 29, 2021 by guest. Protected copyright. 1 Copyright Article author (or their employer) 2009.