Meta-Analysis Identifies Seven Susceptibility Loci Involved in the Atopic March
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Contiguous Deletion of the NDP, MAOA, MAOB, and EFHC2 Genes in a Patient with Norrie Disease, Severe Psychomotor Retardation and Myoclonic Epilepsy
ß 2007 Wiley-Liss, Inc. American Journal of Medical Genetics Part A 143A:916–920 (2007) Contiguous Deletion of the NDP, MAOA, MAOB, and EFHC2 Genes in a Patient With Norrie Disease, Severe Psychomotor Retardation and Myoclonic Epilepsy L. Rodriguez-Revenga,1,2 I. Madrigal,1,2 L.S. Alkhalidi,3 L. Armengol,4 E. Gonza´lez,4 C. Badenas,1,2 X. Estivill,1,5 and M. Mila`1,2* 1Biochemistry and Molecular Genetics Department, Hospital Clı´nic, Barcelona, Spain 2IDIBAPS (Institut d’Investigacions Biome`diques August Pi i Sunyer), Barcelona, Spain 3Department of Health and Medical Services, Rashid Hospital, Dubai, United Arab Emirates 4Genes and Disease Programme, Centre for Genomic Regulation (CRG), Barcelona Biomedical Research Park, Barcelona, Spain 5Department of Experimental and Health Sciences, Universitat Pompeu Fabra (UPF), Barcelona, Spain Received 20 February 2006; Accepted 7 September 2006 Norrie disease (ND) is an X-linked disorder, inherited as a Clinical features of the proband include bilateral retinal recessive trait that, therefore, mostly affects males. The gene detachment, microcephaly, severe psychomotor retardation responsible for ND, called NDP, maps to the short arm of without verbal language skills acquired, and epilepsy. The chromosome X (Xp11.4-p11.3). We report here an atypical identification and molecular characterization of this case case of ND, consisting of a patient harboring a large reinforces the idea of a new contiguous gene syndrome that submicroscopic deletion affecting not only the NDP gene would explain the complex phenotype shared by atypical but also the MAOA, MAOB, and EFHC2 genes. Microarray ND patients. ß 2007 Wiley-Liss, Inc. -

WO 2Ull/13162O Al
(12) INTERNATIONAL APPLICATION PUBLISHED UNDER THE PATENT COOPERATION TREATY (PCT) (19) World Intellectual Property Organization International Bureau (10) International Publication Number (43) International Publication Date Χ 1 / A 1 27 October 2011 (27.10.2011) WO 2Ull/13162o Al (51) International Patent Classification: AO, AT, AU, AZ, BA, BB, BG, BH, BR, BW, BY, BZ, C12N 9/02 (2006.01) A61K 38/44 (2006.01) CA, CH, CL, CN, CO, CR, CU, CZ, DE, DK, DM, DO, A61K 38/17 (2006.01) DZ, EC, EE, EG, ES, FI, GB, GD, GE, GH, GM, GT, HN, HR, HU, ID, IL, IN, IS, JP, KE, KG, KM, KN, KP, (21) International Application Number: KR, KZ, LA, LC, LK, LR, LS, LT, LU, LY, MA, MD, PCT/EP20 11/056142 ME, MG, MK, MN, MW, MX, MY, MZ, NA, NG, NI, (22) International Filing Date: NO, NZ, OM, PE, PG, PH, PL, PT, RO, RS, RU, SC, SD, 18 April 201 1 (18.04.201 1) SE, SG, SK, SL, SM, ST, SV, SY, TH, TJ, TM, TN, TR, TT, TZ, UA, UG, US, UZ, VC, VN, ZA, ZM, ZW. (25) Filing Language: English (84) Designated States (unless otherwise indicated, for every (26) Publication Langi English kind of regional protection available): ARIPO (BW, GH, (30) Priority Data: GM, KE, LR, LS, MW, MZ, NA, SD, SL, SZ, TZ, UG, 10160368.6 19 April 2010 (19.04.2010) EP ZM, ZW), Eurasian (AM, AZ, BY, KG, KZ, MD, RU, TJ, TM), European (AL, AT, BE, BG, CH, CY, CZ, DE, DK, (71) Applicants (for all designated States except US): MEDI- EE, ES, FI, FR, GB, GR, HR, HU, IE, IS, FT, LT, LU, ZINISCHE UNIVERSITAT INNSBRUCK [AT/AT]; LV, MC, MK, MT, NL, NO, PL, PT, RO, RS, SE, SI, SK, Christoph-Probst-Platz, Innrain 52, A-6020 Innsbruck SM, TR), OAPI (BF, BJ, CF, CG, CI, CM, GA, GN, GQ, (AT). -
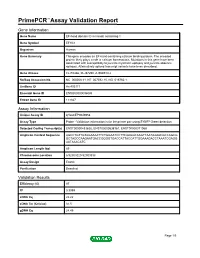
Primepcr™Assay Validation Report
PrimePCR™Assay Validation Report Gene Information Gene Name EF-hand domain (C-terminal) containing 1 Gene Symbol EFHC1 Organism Human Gene Summary This gene encodes an EF-hand-containing calcium binding protein. The encoded protein likely plays a role in calcium homeostasis. Mutations in this gene have been associated with susceptibility to juvenile myoclonic epilepsy and juvenile absence epilepsy. Alternatively spliced transcript variants have been described. Gene Aliases FLJ10466, FLJ37290, dJ304B14.2 RefSeq Accession No. NC_000006.11, NT_007592.15, NG_016760.1 UniGene ID Hs.403171 Ensembl Gene ID ENSG00000096093 Entrez Gene ID 114327 Assay Information Unique Assay ID qHsaCEP0049958 Assay Type Probe - Validation information is for the primer pair using SYBR® Green detection Detected Coding Transcript(s) ENST00000433625, ENST00000538167, ENST00000371068 Amplicon Context Sequence AGCCTGTTGTAGAAAATTCTGGAATCCTTCAAGGCAAGTTAATAAAACGCCAGCG GCTAGCCAAGAATGACCGGGGTGACCATTACCATTGGAAAGACCTAAATCGAGG AATAAACATC Amplicon Length (bp) 89 Chromosome Location 6:52303220-52303338 Assay Design Exonic Purification Desalted Validation Results Efficiency (%) 97 R2 0.9999 cDNA Cq 22.22 cDNA Tm (Celsius) 81.5 gDNA Cq 24.45 Page 1/5 PrimePCR™Assay Validation Report Specificity (%) 100 Information to assist with data interpretation is provided at the end of this report. Page 2/5 PrimePCR™Assay Validation Report EFHC1, Human Amplification Plot Amplification of cDNA generated from 25 ng of universal reference RNA Melt Peak Melt curve analysis of above amplification -

Supplementary Table S4. FGA Co-Expressed Gene List in LUAD
Supplementary Table S4. FGA co-expressed gene list in LUAD tumors Symbol R Locus Description FGG 0.919 4q28 fibrinogen gamma chain FGL1 0.635 8p22 fibrinogen-like 1 SLC7A2 0.536 8p22 solute carrier family 7 (cationic amino acid transporter, y+ system), member 2 DUSP4 0.521 8p12-p11 dual specificity phosphatase 4 HAL 0.51 12q22-q24.1histidine ammonia-lyase PDE4D 0.499 5q12 phosphodiesterase 4D, cAMP-specific FURIN 0.497 15q26.1 furin (paired basic amino acid cleaving enzyme) CPS1 0.49 2q35 carbamoyl-phosphate synthase 1, mitochondrial TESC 0.478 12q24.22 tescalcin INHA 0.465 2q35 inhibin, alpha S100P 0.461 4p16 S100 calcium binding protein P VPS37A 0.447 8p22 vacuolar protein sorting 37 homolog A (S. cerevisiae) SLC16A14 0.447 2q36.3 solute carrier family 16, member 14 PPARGC1A 0.443 4p15.1 peroxisome proliferator-activated receptor gamma, coactivator 1 alpha SIK1 0.435 21q22.3 salt-inducible kinase 1 IRS2 0.434 13q34 insulin receptor substrate 2 RND1 0.433 12q12 Rho family GTPase 1 HGD 0.433 3q13.33 homogentisate 1,2-dioxygenase PTP4A1 0.432 6q12 protein tyrosine phosphatase type IVA, member 1 C8orf4 0.428 8p11.2 chromosome 8 open reading frame 4 DDC 0.427 7p12.2 dopa decarboxylase (aromatic L-amino acid decarboxylase) TACC2 0.427 10q26 transforming, acidic coiled-coil containing protein 2 MUC13 0.422 3q21.2 mucin 13, cell surface associated C5 0.412 9q33-q34 complement component 5 NR4A2 0.412 2q22-q23 nuclear receptor subfamily 4, group A, member 2 EYS 0.411 6q12 eyes shut homolog (Drosophila) GPX2 0.406 14q24.1 glutathione peroxidase -

Detailed Characterization of Human Induced Pluripotent Stem Cells Manufactured for Therapeutic Applications
Stem Cell Rev and Rep DOI 10.1007/s12015-016-9662-8 Detailed Characterization of Human Induced Pluripotent Stem Cells Manufactured for Therapeutic Applications Behnam Ahmadian Baghbaderani 1 & Adhikarla Syama2 & Renuka Sivapatham3 & Ying Pei4 & Odity Mukherjee2 & Thomas Fellner1 & Xianmin Zeng3,4 & Mahendra S. Rao5,6 # The Author(s) 2016. This article is published with open access at Springerlink.com Abstract We have recently described manufacturing of hu- help determine which set of tests will be most useful in mon- man induced pluripotent stem cells (iPSC) master cell banks itoring the cells and establishing criteria for discarding a line. (MCB) generated by a clinically compliant process using cord blood as a starting material (Baghbaderani et al. in Stem Cell Keywords Induced pluripotent stem cells . Embryonic stem Reports, 5(4), 647–659, 2015). In this manuscript, we de- cells . Manufacturing . cGMP . Consent . Markers scribe the detailed characterization of the two iPSC clones generated using this process, including whole genome se- quencing (WGS), microarray, and comparative genomic hy- Introduction bridization (aCGH) single nucleotide polymorphism (SNP) analysis. We compare their profiles with a proposed calibra- Induced pluripotent stem cells (iPSCs) are akin to embryonic tion material and with a reporter subclone and lines made by a stem cells (ESC) [2] in their developmental potential, but dif- similar process from different donors. We believe that iPSCs fer from ESC in the starting cell used and the requirement of a are likely to be used to make multiple clinical products. We set of proteins to induce pluripotency [3]. Although function- further believe that the lines used as input material will be used ally identical, iPSCs may differ from ESC in subtle ways, at different sites and, given their immortal status, will be used including in their epigenetic profile, exposure to the environ- for many years or even decades. -

Elucidating Biological Roles of Novel Murine Genes in Hearing Impairment in Africa
Preprints (www.preprints.org) | NOT PEER-REVIEWED | Posted: 19 September 2019 doi:10.20944/preprints201909.0222.v1 Review Elucidating Biological Roles of Novel Murine Genes in Hearing Impairment in Africa Oluwafemi Gabriel Oluwole,1* Abdoulaye Yal 1,2, Edmond Wonkam1, Noluthando Manyisa1, Jack Morrice1, Gaston K. Mazanda1 and Ambroise Wonkam1* 1Division of Human Genetics, Department of Pathology, Faculty of Health Sciences, University of Cape Town, Observatory, Cape Town, South Africa. 2Department of Neurology, Point G Teaching Hospital, University of Sciences, Techniques and Technology, Bamako, Mali. *Correspondence to: [email protected]; [email protected] Abstract: The prevalence of congenital hearing impairment (HI) is highest in Africa. Estimates evaluated genetic causes to account for 31% of HI cases in Africa, but the identification of associated causative genes mutations have been challenging. In this study, we reviewed the potential roles, in humans, of 38 novel genes identified in a murine study. We gathered information from various genomic annotation databases and performed functional enrichment analysis using online resources i.e. genemania and g.proflier. Results revealed that 27/38 genes are express mostly in the brain, suggesting additional cognitive roles. Indeed, HERC1- R3250X had been associated with intellectual disability in a Moroccan family. A homozygous 216-bp deletion in KLC2 was found in two siblings of Egyptian descent with spastic paraplegia. Up to 27/38 murine genes have link to at least a disease, and the commonest mode of inheritance is autosomal recessive (n=8). Network analysis indicates that 20 other genes have intermediate and biological links to the novel genes, suggesting their possible roles in HI. -

Mouse Tmtc2 Conditional Knockout Project (CRISPR/Cas9)
https://www.alphaknockout.com Mouse Tmtc2 Conditional Knockout Project (CRISPR/Cas9) Objective: To create a Tmtc2 conditional knockout Mouse model (C57BL/6J) by CRISPR/Cas-mediated genome engineering. Strategy summary: The Tmtc2 gene (NCBI Reference Sequence: NM_177368 ; Ensembl: ENSMUSG00000036019 ) is located on Mouse chromosome 10. 12 exons are identified, with the ATG start codon in exon 1 and the TGA stop codon in exon 12 (Transcript: ENSMUST00000061506). Exon 2 will be selected as conditional knockout region (cKO region). Deletion of this region should result in the loss of function of the Mouse Tmtc2 gene. To engineer the targeting vector, homologous arms and cKO region will be generated by PCR using BAC clone RP23-34F6 as template. Cas9, gRNA and targeting vector will be co-injected into fertilized eggs for cKO Mouse production. The pups will be genotyped by PCR followed by sequencing analysis. Note: Exon 2 starts from about 3.35% of the coding region. The knockout of Exon 2 will result in frameshift of the gene. The size of intron 1 for 5'-loxP site insertion: 159879 bp, and the size of intron 2 for 3'-loxP site insertion: 42438 bp. The size of effective cKO region: ~1071 bp. The cKO region does not have any other known gene. Page 1 of 8 https://www.alphaknockout.com Overview of the Targeting Strategy Wildtype allele gRNA region 5' gRNA region 3' 1 2 12 Targeting vector Targeted allele Constitutive KO allele (After Cre recombination) Legends Exon of mouse Tmtc2 Homology arm cKO region loxP site Page 2 of 8 https://www.alphaknockout.com Overview of the Dot Plot Window size: 10 bp Forward Reverse Complement Sequence 12 Note: The sequence of homologous arms and cKO region is aligned with itself to determine if there are tandem repeats. -

Novel Gene Discovery in Primary Ciliary Dyskinesia
Novel Gene Discovery in Primary Ciliary Dyskinesia Mahmoud Raafat Fassad Genetics and Genomic Medicine Programme Great Ormond Street Institute of Child Health University College London A thesis submitted in conformity with the requirements for the degree of Doctor of Philosophy University College London 1 Declaration I, Mahmoud Raafat Fassad, confirm that the work presented in this thesis is my own. Where information has been derived from other sources, I confirm that this has been indicated in the thesis. 2 Abstract Primary Ciliary Dyskinesia (PCD) is one of the ‘ciliopathies’, genetic disorders affecting either cilia structure or function. PCD is a rare recessive disease caused by defective motile cilia. Affected individuals manifest with neonatal respiratory distress, chronic wet cough, upper respiratory tract problems, progressive lung disease resulting in bronchiectasis, laterality problems including heart defects and adult infertility. Early diagnosis and management are essential for better respiratory disease prognosis. PCD is a highly genetically heterogeneous disorder with causal mutations identified in 36 genes that account for the disease in about 70% of PCD cases, suggesting that additional genes remain to be discovered. Targeted next generation sequencing was used for genetic screening of a cohort of patients with confirmed or suggestive PCD diagnosis. The use of multi-gene panel sequencing yielded a high diagnostic output (> 70%) with mutations identified in known PCD genes. Over half of these mutations were novel alleles, expanding the mutation spectrum in PCD genes. The inclusion of patients from various ethnic backgrounds revealed a striking impact of ethnicity on the composition of disease alleles uncovering a significant genetic stratification of PCD in different populations. -

Discovery of an O-Mannosylation Pathway Selectively Serving Cadherins and Protocadherins
Discovery of an O-mannosylation pathway selectively serving cadherins and protocadherins Ida Signe Bohse Larsena,1, Yoshiki Narimatsua,1, Hiren Jitendra Joshia,1, Lina Siukstaitea, Oliver J. Harrisonb, Julia Braschb, Kerry M. Goodmanb, Lars Hansena, Lawrence Shapirob,c,d, Barry Honigb,c,d,e, Sergey Y. Vakhrusheva, Henrik Clausena, and Adnan Halima,2,3 aDepartment of Cellular and Molecular Medicine, Faculty of Health Sciences, Copenhagen Center for Glycomics, University of Copenhagen, DK-2200 Copenhagen, Denmark; bDepartment of Biochemistry and Molecular Biophysics, Columbia University, New York, NY 10032; cZuckerman Mind Brain Behavior Institute, Columbia University, New York, NY 10032; dDepartment of Systems Biology, Columbia University, New York, NY 10032; and eHoward Hughes Medical Institute, Columbia University, New York, NY 10032 Edited by Stuart A. Kornfeld, Washington University School of Medicine, St. Louis, MO, and approved September 6, 2017 (received for review May 22, 2017) The cadherin (cdh) superfamily of adhesion molecules carry muscular dystrophies that have been designated α-dystroglyca- O-linked mannose (O-Man) glycans at highly conserved sites nopathies because deficient O-Man glycosylation of α-DG dis- localized to specific β-strands of their extracellular cdh (EC) domains. rupts the interaction between the dystrophin glycoprotein complex These O-Man glycans do not appear to be elongated like O-Man and the ECM (7–9). Several studies have also implicated de- glycans found on α-dystroglycan (α-DG), and we recently demon- ficiency of POMT2 with E-cdh dysfunction (10–12), although di- strated that initiation of cdh/protocadherin (pcdh) O-Man glycosyl- rect evidence for a role in glycosylation of cdhs and pcdhs is ation is not dependent on the evolutionary conserved POMT1/ missing. -

Novel Extensions of Label Propagation for Biomarker Discovery in Genomic Data
NOVEL EXTENSIONS OF LABEL PROPAGATION FOR BIOMARKER DISCOVERY IN GENOMIC DATA by Matthew E. Stokes B.S. Systems and Control Engineering, Case Western Reserve University, 2008 M.S. Intelligent Systems Program / Biomedical Informatics, University of Pittsburgh 2011 Submitted to the Graduate Faculty of the Kenneth P. Dietrich School of Arts and Sciences in fulfillment of the requirements for the degree of Doctor of Philosophy University of Pittsburgh 2014 UNIVERSITY OF PITTSBURGH DIETRICH SCHOOL OF ARTS AND SCIENCES This dissertation proposal was presented by Matthew E. Stokes on July 17, 2014 and approved by M. Michael Barmada, PhD, Department of Human Genetics Gregory F. Cooper, MD, PhD, Department of Biomedical Informatics and the Intelligent Systems Program Milos Hauskrecht, PhD, Department of Computer Science and the Intelligent Systems Program Dissertation Advisor: Shyam Visweswaran, MD, PhD, Department of Biomedical Informatics and the Intelligent Systems Program ii NOVEL EXTENSIONS OF LABEL PORPAGATION FOR BIOMARKER DISCOVERY IN GENOMIC DATA Matthew E. Stokes, M.S University of Pittsburgh, 2014 Copyright © by Matthew E. Stokes 2014 iii NOVEL EXTENSIONS OF LABEL PROPAGATION FOR BIOMARKER DISCOVERY IN GENOMIC DATA Matthew E. Stokes, PhD University of Pittsburgh, 2014 One primary goal of analyzing genomic data is the identification of biomarkers which may be causative of, correlated with, or otherwise biologically relevant to disease phenotypes. In this work, I implement and extend a multivariate feature ranking algorithm called label propagation (LP) for biomarker discovery in genome-wide single-nucleotide polymorphism (SNP) data. This graph-based algorithm utilizes an iterative propagation method to efficiently compute the strength of association between a SNP and a phenotype. -

A Regulator of Innate Immune Responses
(19) TZZ ¥_T (11) EP 2 942 357 A1 (12) EUROPEAN PATENT APPLICATION (43) Date of publication: (51) Int Cl.: 11.11.2015 Bulletin 2015/46 C07K 14/47 (2006.01) A61K 38/00 (2006.01) C12N 15/113 (2010.01) (21) Application number: 15169327.2 (22) Date of filing: 04.08.2009 (84) Designated Contracting States: (72) Inventor: Barber, Glen N. AT BE BG CH CY CZ DE DK EE ES FI FR GB GR Palmetto Bay, FL 33157 (US) HR HU IE IS IT LI LT LU LV MC MK MT NL NO PL PT RO SE SI SK SM TR (74) Representative: Inspicos A/S Kogle Allé 2 (30) Priority: 04.08.2008 US 129975 P P.O. Box 45 2970 Hørsholm (DK) (62) Document number(s) of the earlier application(s) in accordance with Art. 76 EPC: Remarks: 09805473.7 / 2 324 044 This application was filed on 27-05-2015 as a divisional application to the application mentioned (71) Applicant: Barber, Glen N. under INID code 62. Palmetto Bay, FL 33157 (US) (54) STING (STIMULATOR OF INTEFERON GENES), A REGULATOR OF INNATE IMMUNE RESPONSES (57) Novel molecules termed STING which include STING compositions are useful for the treatment of an nucleic acids, polynucleotides, oligonucleotides, pep- immune-related disorder, including treating and prevent- tides, mutants, variants and active fragments thereof, ing infection by modulating immunity. modulate innate and adaptive immunity in a subject. EP 2 942 357 A1 Printed by Jouve, 75001 PARIS (FR) EP 2 942 357 A1 Description RELATED APPLICATIONS 5 [0001] This application claims priority under 35 USC § 119 to U.S. -

TMTC2 (G-15): Sc-169651
SAN TA C RUZ BI OTEC HNOL OG Y, INC . TMTC2 (G-15): sc-169651 BACKGROUND SOURCE The tetratricopeptide repeat (TPR) motif is a degenerate, 34 amino acid TMTC2 (G-15) is an affinity purified goat polyclonal antibody raised against sequence found in many proteins and acts to mediate protein-protein inter - a peptide mapping within an internal region of TMTC2 of human origin. actions in various pathways. At the sequence level, there can be up to 16 tandem TPR repeats, each of which has a helix-turn-helix shape that stacks PRODUCT on other TPR repeats to achieve ligand binding specificity. TMTC2 (trans - Each vial contains 200 µg IgG in 1.0 ml of PBS with < 0.1% sodium azide membrane and tetratricopeptide repeat containing 2) is an 836 amino acid and 0.1% gelatin. multi-pass membrane protein that contains ten TPR repeats and belongs to the TMTC family. The gene encoding TMTC2 maps to human chromosome 12, Blocking peptide available for competition studies, sc-169651 P, (100 µg which encodes over 1,100 genes and comprises approximately 4.5% of the peptide in 0.5 ml PBS containing < 0.1% sodium azide and 0.2% BSA). human genome. Chromosome 12 is associated with a variety of diseases and afflictions, including hypochondrogenesis, achondrogenesis, Kniest dysplasia, APPLICATIONS Noonan syndrome and trisomy 12p, which causes facial developmental TMTC2 (G-15) is recommended for detection of TMTC2 of mouse, rat and defects and seizure disorders. human origin by Western Blotting (starting dilution 1:200, dilution range 1:100-1:1000), immunofluorescence (starting dilution 1:50, dilution range REFERENCES 1:50-1:500) and solid phase ELISA (starting dilution 1:30, dilution range 1.