An Integrated Approach to Dissecting Oncogene Addiction Implicates a Myb-Coordinated Self-Renewal Program As Essential for Leukemia Maintenance
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Molecular Profile of Tumor-Specific CD8+ T Cell Hypofunction in a Transplantable Murine Cancer Model
Downloaded from http://www.jimmunol.org/ by guest on September 25, 2021 T + is online at: average * The Journal of Immunology , 34 of which you can access for free at: 2016; 197:1477-1488; Prepublished online 1 July from submission to initial decision 4 weeks from acceptance to publication 2016; doi: 10.4049/jimmunol.1600589 http://www.jimmunol.org/content/197/4/1477 Molecular Profile of Tumor-Specific CD8 Cell Hypofunction in a Transplantable Murine Cancer Model Katherine A. Waugh, Sonia M. Leach, Brandon L. Moore, Tullia C. Bruno, Jonathan D. Buhrman and Jill E. Slansky J Immunol cites 95 articles Submit online. Every submission reviewed by practicing scientists ? is published twice each month by Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts http://jimmunol.org/subscription Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html http://www.jimmunol.org/content/suppl/2016/07/01/jimmunol.160058 9.DCSupplemental This article http://www.jimmunol.org/content/197/4/1477.full#ref-list-1 Information about subscribing to The JI No Triage! Fast Publication! Rapid Reviews! 30 days* Why • • • Material References Permissions Email Alerts Subscription Supplementary The Journal of Immunology The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2016 by The American Association of Immunologists, Inc. All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. This information is current as of September 25, 2021. The Journal of Immunology Molecular Profile of Tumor-Specific CD8+ T Cell Hypofunction in a Transplantable Murine Cancer Model Katherine A. -

Chronic Myeloid Leukemia: Mechanisms of Blastic Transformation
Chronic myeloid leukemia: mechanisms of blastic transformation Danilo Perrotti, … , John Goldman, Tomasz Skorski J Clin Invest. 2010;120(7):2254-2264. https://doi.org/10.1172/JCI41246. Science in Medicine The BCR-ABL1 oncoprotein transforms pluripotent HSCs and initiates chronic myeloid leukemia (CML). Patients with early phase (also known as chronic phase [CP]) disease usually respond to treatment with ABL tyrosine kinase inhibitors (TKIs), although some patients who respond initially later become resistant. In most patients, TKIs reduce the leukemia cell load substantially, but the cells from which the leukemia cells are derived during CP (so-called leukemia stem cells [LSCs]) are intrinsically insensitive to TKIs and survive long term. LSCs or their progeny can acquire additional genetic and/or epigenetic changes that cause the leukemia to transform from CP to a more advanced phase, which has been subclassified as either accelerated phase or blastic phase disease. The latter responds poorly to treatment and is usually fatal. Here, we discuss what is known about the molecular mechanisms leading to blastic transformation of CML and propose some novel therapeutic approaches. Find the latest version: https://jci.me/41246/pdf Science in medicine Chronic myeloid leukemia: mechanisms of blastic transformation Danilo Perrotti,1 Catriona Jamieson,2 John Goldman,3 and Tomasz Skorski4 1Department of Molecular Virology, Immunology and Medical Genetics and Comprehensive Cancer Center, The Ohio State University, Columbus, Ohio, USA. 2Division of Hematology-Oncology, Department of Internal Medicine, University of California at San Diego, La Jolla, California, USA. 3Department of Haematology, Imperial College at Hammersmith Hospital, London, United Kingdom. 4Department of Microbiology and Immunology, Temple University, Philadelphia, Pennsylvania, USA. -

Targeting Non-Oncogene Addiction for Cancer Therapy
biomolecules Review Targeting Non-Oncogene Addiction for Cancer Therapy Hae Ryung Chang 1,*,†, Eunyoung Jung 1,†, Soobin Cho 1, Young-Jun Jeon 2 and Yonghwan Kim 1,* 1 Department of Biological Sciences and Research Institute of Women’s Health, Sookmyung Women’s University, Seoul 04310, Korea; [email protected] (E.J.); [email protected] (S.C.) 2 Department of Integrative Biotechnology, Sungkyunkwan University, Suwon 16419, Korea; [email protected] * Correspondence: [email protected] (H.R.C.); [email protected] (Y.K.); Tel.: +82-2-710-9552 (H.R.C.); +82-2-710-9552 (Y.K.) † These authors contributed equally. Abstract: While Next-Generation Sequencing (NGS) and technological advances have been useful in identifying genetic profiles of tumorigenesis, novel target proteins and various clinical biomarkers, cancer continues to be a major global health threat. DNA replication, DNA damage response (DDR) and repair, and cell cycle regulation continue to be essential systems in targeted cancer therapies. Although many genes involved in DDR are known to be tumor suppressor genes, cancer cells are often dependent and addicted to these genes, making them excellent therapeutic targets. In this review, genes implicated in DNA replication, DDR, DNA repair, cell cycle regulation are discussed with reference to peptide or small molecule inhibitors which may prove therapeutic in cancer patients. Additionally, the potential of utilizing novel synthetic lethal genes in these pathways is examined, providing possible new targets for future therapeutics. Specifically, we evaluate the potential of TONSL as a novel gene for targeted therapy. Although it is a scaffold protein with no known enzymatic activity, the strategy used for developing PCNA inhibitors can also be utilized to target TONSL. -

Transcription Factors E2A, FOXO1 and FOXP1 Regulate Recombination Activating Gene Expression in Cancer Cells
Transcription Factors E2A, FOXO1 and FOXP1 Regulate Recombination Activating Gene Expression in Cancer Cells Zhengshan Chen1, Yanna Xiao1, Junjun Zhang1, Jing Li1, Yuxuan Liu2, Yingying Zhao2, Changchun Ma1, Jin Luo1, Yamei Qiu1, Guowei Huang1, Christine Korteweg1, Jiang Gu1,2* 1 Department of Pathology, Shantou University Medical College, Shantou, China, 2 Department of Pathology, Peking (Beijing) University Health Science Center, Beijing, China Abstract It has long been accepted that immunoglobulins (Igs) were produced by B lymphoid cells only. Recently Igs have been found to be expressed in various human cancer cells and promote tumor growth. Recombination activating gene 1 (RAG1) and RAG2, which are essential enzymes for initiating variable-diversity-joining segment recombination, have also been found to be expressed in cancer cells. However, the mechanism of RAG activation in these cancer cells has not been elucidated. Here, we investigated the regulatory mechanism of RAG expression in four human cancer cell lines by analyzing transcription factors that induce RAG activation in B cells. By RT-PCR, Western blot and immunofluorescence, we found that transcription factors E2A, FOXO1 and FOXP1 were expressed and localized to the nuclei of these cancer cells. Over- expression of E2A, FOXO1 or Foxp1 increased RAG expression, while RNA interference of E2A, FOXO1 or FOXP1 decreased RAG expression in the cancer cells. Chromatin immunoprecipitation experiments showed acetylation of RAG enhancer (Erag) and E2A, FOXO1 or FOXP1 were bound to Erag in vivo. These results indicate that in these cancer cells the transcription factors E2A, FOXO1 and FOXP1 regulate RAG expression, which initiates Ig gene rearrangement much in the way similar to B lymphocytes. -

1078-0432.CCR-20-1706.Full.Pdf
Author Manuscript Published OnlineFirst on June 29, 2020; DOI: 10.1158/1078-0432.CCR-20-1706 Author manuscripts have been peer reviewed and accepted for publication but have not yet been edited. Gastrointestinal stromal tumor: challenges and opportunities for a new decade César Serrano1,2, Suzanne George3 1Sarcoma Translational Research Laboratory, Vall d’Hebron Institute of Oncology, Barcelona, Spain. 2Department of Medical Oncology, Vall d’Hebron University Hospital, Barcelona, Spain. 3Department of Medical Oncology, Sarcoma Center, Dana-Farber Cancer Institute, Boston, US. Running title: Review on gastrointestinal stromal tumor Key words: Avapritinib; circulating tumor DNA; gastrointestinal stromal tumor; imatinib; regorafenib; ripretinib; sunitinib. Financial support: This work was supported by grants (to C.S.) from SARC Career Development Award, FIS ISCIII (PI19/01271), PERIS 2018 (SLT006/17/221) and FERO Foundation. Conflicts of interest: C.S. has received research grants from Deciphera Pharmaceuticals, Bayer AG and Pfizer, Inc; consulting fees (advisory role) from Deciphera Pharmaceuticals and Blueprint 1 Downloaded from clincancerres.aacrjournals.org on October 1, 2021. © 2020 American Association for Cancer Research. Author Manuscript Published OnlineFirst on June 29, 2020; DOI: 10.1158/1078-0432.CCR-20-1706 Author manuscripts have been peer reviewed and accepted for publication but have not yet been edited. Medicines; payment for lectures from Bayer AG and Blueprint Medicines; and travel grants from Pharmamar, Pfizer, Bayer AG, Novartis and Lilly. S.G. has received research funding to her institution from Blueprint Medicines, Deciphera Pharmaceutical, Bayer AG, Pfizer, Novartis; consulting fees (advisory role) from Blueprint Medicines, Deciphera Pharmaceuticals, Eli Lilly, Bayer AG. Corresponding author: Dr. César Serrano, Vall d’Hebron Institute of Oncology (VHIO), Vall d’Hebron University Hospital. -

TET2 Repression by Androgen Hormone Regulates Global Hydroxymethylation Status and Prostate Cancer Progression
ARTICLE Received 5 Oct 2014 | Accepted 30 Jul 2015 | Published 25 Sep 2015 DOI: 10.1038/ncomms9219 TET2 repression by androgen hormone regulates global hydroxymethylation status and prostate cancer progression Ken-ichi Takayama1,2,*, Aya Misawa1,*, Takashi Suzuki3, Kiyoshi Takagi3, Yoshihide Hayashizaki4,5, Tetsuya Fujimura6, Yukio Homma6, Satoru Takahashi7, Tomohiko Urano1,2 & Satoshi Inoue1,2,8 Modulation of epigenetic patterns has promising efficacy for treating cancer. 5-Hydro- xymethylated cytosine (5-hmC) is an epigenetic mark potentially important in cancer. Here we report that 5-hmC is an epigenetic hallmark of prostate cancer (PCa) progression. A member of the ten–eleven translocation (TET) proteins, which catalyse the oxidation of methylated cytosine (5-mC) to 5-hmC, TET2, is repressed by androgens in PCa. Androgen receptor (AR)-mediated induction of the miR-29 family, which targets TET2, are markedly enhanced in hormone refractory PCa (HRPC) and its high expression predicts poor outcome of PCa patients. Furthermore, decreased expression of miR-29b results in reduced tumour growth and increased TET2 expression in an animal model of HRPC. Interestingly, global 5-hmC modification regulated by miR-29b represses FOXA1 activity. A reduction in 5-hmC activates PCa-related key pathways such as mTOR and AR. Thus, DNA modification directly links the TET2-dependent epigenetic pathway regulated by AR to 5-hmC-mediated tumour progression. 1 Department of Anti-Aging Medicine, Graduate School of Medicine, The University of Tokyo, Bunkyo-ku, Tokyo 113-8655, Japan. 2 Department of Geriatric Medicine, Graduate School of Medicine, The University of Tokyo, Bunkyo-ku, Tokyo 113-8655, Japan. -

FOXP1 Acts Through a Negative Feedback Loop to Suppress FOXO-Induced Apoptosis
Cell Death and Differentiation (2013) 20, 1219–1229 & 2013 Macmillan Publishers Limited All rights reserved 1350-9047/13 www.nature.com/cdd FOXP1 acts through a negative feedback loop to suppress FOXO-induced apoptosis R van Boxtel1,5, C Gomez-Puerto1,6, M Mokry2,6, A Eijkelenboom3, KE van der Vos1, EES Nieuwenhuis2, BMT Burgering3, EW-F Lam4 and PJ Coffer*,1,2 Transcriptional activity of Forkhead box transcription factor class O (FOXO) proteins can result in a variety of cellular outcomes depending on cell type and activating stimulus. These transcription factors are negatively regulated by the phosphoinositol 3-kinase (PI3K)–protein kinase B (PKB) signaling pathway, which is thought to have a pivotal role in regulating survival of tumor cells in a variety of cancers. Recently, it has become clear that FOXO proteins can promote resistance to anti-cancer therapeutics, designed to inhibit PI3K–PKB activity, by inducing the expression of proteins that provide feedback at different levels of this pathway. We questioned whether such a feedback mechanism may also exist directly at the level of FOXO-induced transcription. To identify critical modulators of FOXO transcriptional output, we performed gene expression analyses after conditional activation of key components of the PI3K–PKB–FOXO signaling pathway and identified FOXP1 as a direct FOXO transcriptional target. Using chromatin immunoprecipitation followed by next-generation sequencing, we show that FOXP1 binds enhancers that are pre-occupied by FOXO3. By sequencing the transcriptomes of cells in which FOXO is specifically activated in the absence of FOXP1, we demonstrate that FOXP1 can modulate the expression of a specific subset of FOXO target genes, including inhibiting expression of the pro-apoptotic gene BIK. -

Anti-Aging: Senolytics Or Gerostatics (Unconventional View)
www.oncotarget.com Oncotarget, 2021, Vol. 12, (No. 18), pp: 1821-1835 Review Anti-aging: senolytics or gerostatics (unconventional view) Mikhail V. Blagosklonny1 1Roswell Park Cancer Institute, Buffalo, NY 14263, USA Correspondence to: Mikhail V. Blagosklonny, email: [email protected], [email protected] Keywords: geroscience; senolytics; hyperfunction theory; aging; sirolimus Received: June 07, 2021 Accepted: July 05, 2021 Published: August 31, 2021 Copyright: © 2021 Blagosklonny. This is an open access article distributed under the terms of the Creative Commons Attribution License (CC BY 3.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited. ABSTRACT Senolytics are basically anti-cancer drugs, repurposed to kill senescent cells selectively. It is even more difficult to selectively kill senescent cells than to kill cancer cells. Based on lessons of cancer therapy, here I suggest how to exploit oncogene-addiction and to combine drugs to achieve selectivity. However, even if selective senolytic combinations will be developed, there is little evidence that a few senescent cells are responsible for organismal aging. I also discuss gerostatics, such as rapamycin and other rapalogs, pan-mTOR inhibitors, dual PI3K/mTOR inhibitors, which inhibit growth- and aging-promoting pathways. Unlike senolytics, gerostatics do not kill cells but slow down cellular geroconversion to senescence. Numerous studies demonstrated that inhibition of the mTOR pathways by any means (genetic, pharmacological and dietary) extends lifespan. Currently, only two studies demonstrated that senolytics (fisetin and a combination Dasatinib plus Quercetin) extend lifespan in mice. These senolytics slightly inhibit the mTOR pathway. Thus, life extension by these senolytics can be explained by their slight rapamycin-like (gerostatic) effects. -
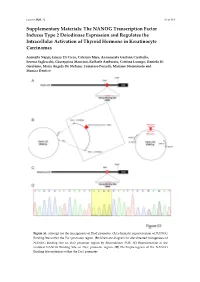
The NANOG Transcription Factor Induces Type 2 Deiodinase Expression and Regulates the Intracellular Activation of Thyroid Hormone in Keratinocyte Carcinomas
Cancers 2020, 12 S1 of S18 Supplementary Materials: The NANOG Transcription Factor Induces Type 2 Deiodinase Expression and Regulates the Intracellular Activation of Thyroid Hormone in Keratinocyte Carcinomas Annarita Nappi, Emery Di Cicco, Caterina Miro, Annunziata Gaetana Cicatiello, Serena Sagliocchi, Giuseppina Mancino, Raffaele Ambrosio, Cristina Luongo, Daniela Di Girolamo, Maria Angela De Stefano, Tommaso Porcelli, Mariano Stornaiuolo and Monica Dentice Figure S1. Strategy for the mutagenesis of Dio2 promoter. (A) Schematic representation of NANOG Binding Site within the Dio2 promoter region. (B) Schematic diagram for site‐directed mutagenesis of NANOG Binding Site on Dio2 promoter region by Recombinant PCR. (C) Representation of the mutated NANOG Binding Site on Dio2 promoter region. (D) Electropherogram of the NANOG Binding Site mutation within the Dio2 promoter. Cancers 2020, 12 S2 of S18 Figure S2. Strategy for the silencing of NANOG expression. (A) Cloning strategies for the generation of NANOG shRNA expression vectors. (B) Electropherograms of the NANOG shRNA sequences cloned into pcDNA3.1 vector. (C) Validation of effective NANOG down-modulation by two different NANOG shRNA vectors was assessed by Western Blot analysis of NANOG expression in BCC cells. (D) Quantification of NANOG protein levels versus Tubulin levels in the same experiment as in C is represented by histograms. Cancers 2020, 12 S3 of S18 Figure S3. The CD34+ cells are characterized by the expression of typical epithelial stemness genes. The mRNA levels of a panel of indicated stemness markers of epidermis were measured by Real Time PCR in the same experiment indicated in figure 3F and G. Cancers 2020, 12 S4 of S18 Figure S4. -

Role of Estrogen Receptor in Breast Cancer Cell Gene Expression
4046 MOLECULAR MEDICINE REPORTS 13: 4046-4050, 2016 Role of estrogen receptor in breast cancer cell gene expression YABING ZHENG1, XIYING SHAO1, YUAN HUANG1, LEI SHI1, BO CHEN2, XIAOJIA WANG1, HONGJIAN YANG3, ZHANHONG CHEN1 and XIPING ZHANG3 Departments of 1Medical Oncology (Breast), 2Pathology and 3Breast Surgery, Zhejiang Cancer Hospital, Hangzhou, Zhejiang 310022, P.R. China Received April 28, 2015; Accepted February 23, 2016 DOI: 10.3892/mmr.2016.5018 Abstract. The aim of the present study was to establish the Europe in 2012, and the number of breast cancer-associated underlying regulatory mechanism of estrogen receptor (ER) mortalities is 131,000 (6). Furthermore, breast cancer is in breast cancer cell gene expression. A gene expression the most common cause of cancer-associated mortality in profile accession( no. GSE11324) was downloaded from the females. Therefore, it is essential to understand its molecular Gene Expression Omnibus (GEO) database. Differentially mechanism and develop more effective therapeutic methods expressed genes (DEGs) from an estrogen treatment group and for breast cancer treatment. a control group were identified. Chromatin immunoprecipita- The estrogen receptor (ER) is critical in determining the tion with high-throughput sequencing data (series GSE25710) phenotype of human breast cancers and is one of the most was obtained from the GEO for the ER binding sites, and important therapeutic targets (7). Furthermore, certain studies binding and expression target analysis was performed. A total have suggested that activation of ER is responsible for various of 3,122 DEGs were obtained and ER was demonstrated to biological processes, including cell growth and differentia- exhibit inhibition and activation roles during the regulation tion, and programmed cell death (8,9). -
Oncogene Addiction As a Rationale for Targeted Anti-Cancer Therapy in Hepatocellular Carcinoma
호암상 수상 기념특강 Oncogene addiction as a rationale for targeted anti-cancer therapy in hepatocellular carcinoma Dae-Ghon Kim Division of Gastroenterology and Hepatology, Department of Internal Medicine, Chonbuk National University Medical School and Hospital, Jeonju, Jeonbuk, Korea Abstract The concept of oncogene addiction was first introduced by Bernard Weinstein in 2000, with particular reference to the observation that some cyclin D-overexpressing cancers reverse their malignant phenotype upon cyclin-D deple- tion by means of RNA interference. It postulates that some tumours rely on one single dominant oncogene for growth and survival, so that inhibition of this specific oncogene is sufficient to halt the neoplastic phenotype. A large amount of evidence has proven the pervasive power of this notion, both in basic research and in therapeutic applications. Application of this concept to the clinical setting has achieved variable success in some various cancer types, including chronic myelogeneous leukaemia harbouring the BCR-ABL translocation, Erb2 overexpressing breast cancer, and non-small cell lung cancer harbouring a subset of EGFR mutations (Table1). However, in the face of such a considerable body of knowledge, the intimate molecular mechanisms mediating this phenomenon remain elusive. At the clinical level, successful translation of the oncogene addiction model into the rational and effective design of targeted therapeutics against individual oncoproteins still faces major obstacles, mainly due to the emergence of escape mechanisms and drug resistance. Sorafenib, a tyrosine kinase inhibitor (TKI), has demon- strated clinical efficacy in patients with HCC. Studies in patients with lung, breast, or colorectal cancers indicated that the genetic heterogeneity of cancer cells within a tumor affect its response to therapeutics designed to target specific molecules. -

Emerging Insights of Tumor Heterogeneity and Drug Resistance Mechanisms in Lung Cancer Targeted Therapy Zuan-Fu Lim1,2,3 and Patrick C
Lim and Ma Journal of Hematology & Oncology (2019) 12:134 https://doi.org/10.1186/s13045-019-0818-2 REVIEW Open Access Emerging insights of tumor heterogeneity and drug resistance mechanisms in lung cancer targeted therapy Zuan-Fu Lim1,2,3 and Patrick C. Ma3* Abstract The biggest hurdle to targeted cancer therapy is the inevitable emergence of drug resistance. Tumor cells employ different mechanisms to resist the targeting agent. Most commonly in EGFR-mutant non-small cell lung cancer, secondary resistance mutations on the target kinase domain emerge to diminish the binding affinity of first- and second-generation inhibitors. Other alternative resistance mechanisms include activating complementary bypass pathways and phenotypic transformation. Sequential monotherapies promise to temporarily address the problem of acquired drug resistance, but evidently are limited by the tumor cells’ ability to adapt and evolve new resistance mechanisms to persist in the drug environment. Recent studies have nominated a model of drug resistance and tumor progression under targeted therapy as a result of a small subpopulation of cells being able to endure the drug (minimal residual disease cells) and eventually develop further mutations that allow them to regrow and become the dominant population in the therapy-resistant tumor. This subpopulation of cells appears to have developed through a subclonal event, resulting in driver mutations different from the driver mutation that is tumor- initiating in the most common ancestor. As such, an understanding of intratumoral heterogeneity—the driving force behind minimal residual disease—is vital for the identification of resistance drivers that results from branching evolution. Currently available methods allow for a more comprehensive and holistic analysis of tumor heterogeneity in that issues associated with spatial and temporal heterogeneity can now be properly addressed.