Essential Roles of the Dystrophin Manuscript.Pdf
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Supplementary Table 3 Complete List of RNA-Sequencing Analysis of Gene Expression Changed by ≥ Tenfold Between Xenograft and Cells Cultured in 10%O2
Supplementary Table 3 Complete list of RNA-Sequencing analysis of gene expression changed by ≥ tenfold between xenograft and cells cultured in 10%O2 Expr Log2 Ratio Symbol Entrez Gene Name (culture/xenograft) -7.182 PGM5 phosphoglucomutase 5 -6.883 GPBAR1 G protein-coupled bile acid receptor 1 -6.683 CPVL carboxypeptidase, vitellogenic like -6.398 MTMR9LP myotubularin related protein 9-like, pseudogene -6.131 SCN7A sodium voltage-gated channel alpha subunit 7 -6.115 POPDC2 popeye domain containing 2 -6.014 LGI1 leucine rich glioma inactivated 1 -5.86 SCN1A sodium voltage-gated channel alpha subunit 1 -5.713 C6 complement C6 -5.365 ANGPTL1 angiopoietin like 1 -5.327 TNN tenascin N -5.228 DHRS2 dehydrogenase/reductase 2 leucine rich repeat and fibronectin type III domain -5.115 LRFN2 containing 2 -5.076 FOXO6 forkhead box O6 -5.035 ETNPPL ethanolamine-phosphate phospho-lyase -4.993 MYO15A myosin XVA -4.972 IGF1 insulin like growth factor 1 -4.956 DLG2 discs large MAGUK scaffold protein 2 -4.86 SCML4 sex comb on midleg like 4 (Drosophila) Src homology 2 domain containing transforming -4.816 SHD protein D -4.764 PLP1 proteolipid protein 1 -4.764 TSPAN32 tetraspanin 32 -4.713 N4BP3 NEDD4 binding protein 3 -4.705 MYOC myocilin -4.646 CLEC3B C-type lectin domain family 3 member B -4.646 C7 complement C7 -4.62 TGM2 transglutaminase 2 -4.562 COL9A1 collagen type IX alpha 1 chain -4.55 SOSTDC1 sclerostin domain containing 1 -4.55 OGN osteoglycin -4.505 DAPL1 death associated protein like 1 -4.491 C10orf105 chromosome 10 open reading frame 105 -4.491 -

Abstracts from the 50Th European Society of Human Genetics Conference: Electronic Posters
European Journal of Human Genetics (2019) 26:820–1023 https://doi.org/10.1038/s41431-018-0248-6 ABSTRACT Abstracts from the 50th European Society of Human Genetics Conference: Electronic Posters Copenhagen, Denmark, May 27–30, 2017 Published online: 1 October 2018 © European Society of Human Genetics 2018 The ESHG 2017 marks the 50th Anniversary of the first ESHG Conference which took place in Copenhagen in 1967. Additional information about the event may be found on the conference website: https://2017.eshg.org/ Sponsorship: Publication of this supplement is sponsored by the European Society of Human Genetics. All authors were asked to address any potential bias in their abstract and to declare any competing financial interests. These disclosures are listed at the end of each abstract. Contributions of up to EUR 10 000 (ten thousand euros, or equivalent value in kind) per year per company are considered "modest". Contributions above EUR 10 000 per year are considered "significant". 1234567890();,: 1234567890();,: E-P01 Reproductive Genetics/Prenatal and fetal echocardiography. The molecular karyotyping Genetics revealed a gain in 8p11.22-p23.1 region with a size of 27.2 Mb containing 122 OMIM gene and a loss in 8p23.1- E-P01.02 p23.3 region with a size of 6.8 Mb containing 15 OMIM Prenatal diagnosis in a case of 8p inverted gene. The findings were correlated with 8p inverted dupli- duplication deletion syndrome cation deletion syndrome. Conclusion: Our study empha- sizes the importance of using additional molecular O¨. Kırbıyık, K. M. Erdog˘an, O¨.O¨zer Kaya, B. O¨zyılmaz, cytogenetic methods in clinical follow-up of complex Y. -

Whole Exome Sequencing in Families at High Risk for Hodgkin Lymphoma: Identification of a Predisposing Mutation in the KDR Gene
Hodgkin Lymphoma SUPPLEMENTARY APPENDIX Whole exome sequencing in families at high risk for Hodgkin lymphoma: identification of a predisposing mutation in the KDR gene Melissa Rotunno, 1 Mary L. McMaster, 1 Joseph Boland, 2 Sara Bass, 2 Xijun Zhang, 2 Laurie Burdett, 2 Belynda Hicks, 2 Sarangan Ravichandran, 3 Brian T. Luke, 3 Meredith Yeager, 2 Laura Fontaine, 4 Paula L. Hyland, 1 Alisa M. Goldstein, 1 NCI DCEG Cancer Sequencing Working Group, NCI DCEG Cancer Genomics Research Laboratory, Stephen J. Chanock, 5 Neil E. Caporaso, 1 Margaret A. Tucker, 6 and Lynn R. Goldin 1 1Genetic Epidemiology Branch, Division of Cancer Epidemiology and Genetics, National Cancer Institute, NIH, Bethesda, MD; 2Cancer Genomics Research Laboratory, Division of Cancer Epidemiology and Genetics, National Cancer Institute, NIH, Bethesda, MD; 3Ad - vanced Biomedical Computing Center, Leidos Biomedical Research Inc.; Frederick National Laboratory for Cancer Research, Frederick, MD; 4Westat, Inc., Rockville MD; 5Division of Cancer Epidemiology and Genetics, National Cancer Institute, NIH, Bethesda, MD; and 6Human Genetics Program, Division of Cancer Epidemiology and Genetics, National Cancer Institute, NIH, Bethesda, MD, USA ©2016 Ferrata Storti Foundation. This is an open-access paper. doi:10.3324/haematol.2015.135475 Received: August 19, 2015. Accepted: January 7, 2016. Pre-published: June 13, 2016. Correspondence: [email protected] Supplemental Author Information: NCI DCEG Cancer Sequencing Working Group: Mark H. Greene, Allan Hildesheim, Nan Hu, Maria Theresa Landi, Jennifer Loud, Phuong Mai, Lisa Mirabello, Lindsay Morton, Dilys Parry, Anand Pathak, Douglas R. Stewart, Philip R. Taylor, Geoffrey S. Tobias, Xiaohong R. Yang, Guoqin Yu NCI DCEG Cancer Genomics Research Laboratory: Salma Chowdhury, Michael Cullen, Casey Dagnall, Herbert Higson, Amy A. -

Common Genetic Aberrations Associated with Metabolic Interferences in Human Type-2 Diabetes and Acute Myeloid Leukemia: a Bioinformatics Approach
International Journal of Molecular Sciences Article Common Genetic Aberrations Associated with Metabolic Interferences in Human Type-2 Diabetes and Acute Myeloid Leukemia: A Bioinformatics Approach Theodora-Christina Kyriakou 1, Panagiotis Papageorgis 1,2 and Maria-Ioanna Christodoulou 3,* 1 Tumor Microenvironment, Metastasis and Experimental Therapeutics Laboratory, Basic and Translational Cancer Research Center, Department of Life Sciences, European University Cyprus, Nicosia 2404, Cyprus; [email protected] (T.-C.K.); [email protected] (P.P.) 2 European University Cyprus Research Center, Nicosia 2404, Cyprus 3 Tumor Immunology and Biomarkers Laboratory, Basic and Translational Cancer Research Center, Department of Life Sciences, European University Cyprus, Nicosia 2404, Cyprus * Correspondence: [email protected] Abstract: Type-2 diabetes mellitus (T2D) is a chronic metabolic disorder, associated with an increased risk of developing solid tumors and hematological malignancies, including acute myeloid leukemia (AML). However, the genetic background underlying this predisposition remains elusive. We herein aimed at the exploration of the genetic variants, related transcriptomic changes and disturbances in metabolic pathways shared by T2D and AML, utilizing bioinformatics tools and repositories, as well as publicly available clinical datasets. Our approach revealed that rs11709077 and rs1801282, on PPARG, rs11108094 on USP44, rs6685701 on RPS6KA1 and rs7929543 on AC118942.1 comprise Citation: Kyriakou, T.-C.; common SNPs susceptible to the two diseases and, together with 64 other co-inherited proxy SNPs, Papageorgis, P.; Christodoulou, M.-I. may affect the expression patterns of metabolic genes, such as USP44, METAP2, PPARG, TIMP4 and Common Genetic Aberrations RPS6KA1, in adipose tissue, skeletal muscle, liver, pancreas and whole blood. -

Limb Girdle Muscular Dystrophy in The
Limb Girdle Muscular Dystrophy in the Hutterite Population of North America by Patrick Flosk A Thesis Submitted to the Faculty of Gmduate Studies in Fulhlment of the Requirements for the Degree of Doctor of Philosophy Deparlrnent of Biochemistry and Medical Genetics University of Manitoba WiIuripeg, Manitoba OPatrick Frosk, April 2005 THE UNI\'ERSITY OF MANITOBA FACULTY OF G.RÄDUATE STUDMS COPYRIGHT PERI{ISSION PAGE Limb Girdle Muscular Dystrophy in the Hutterite Population of North Anrerica BY Patrick Frosk A Thesis/Practicum submitted to the Faculfy of Graduate Studies ofThe University of Manitoba in partial fulfìllnlent of the requir.ements of the degree of DOCTOR OF PHILOSOPHY PATRICK FROSK @2005 Permission has been granted to the Library ofrhe university of Manitoba to Iend or sell copies of this thesis/¡rracticum, to the National Library of Canada to microfilm this thesis and to lend or sell copies of the fìlm, and to university Microfilm Inc. to publish a¡l âbstrâct of this thesis/practicun. The author reserves other publication rights, and neither this thesis/practícum nor extensive extracts from it may be printed or othelrvise reproduced \yithout the author's n'ritten permission. Table of Contents Table of Contents i Abstract iv Acknorvledgements vi List of Figures v l List of Tables ix List of Abbreviations x List of Manufacturers x Chapter 1: Introduction I Chapter 2: Revierv of Limb Girdle Muscular Dystrophies J 2,1 General Features of LGMD 3 2.2 Dominant Forms 6 (A) LGMDlA 6 (B) LGMDIB 9 (c) LGMDlC 18 (D) LGMDID 24 (E) LGMD1E -

Supplementary Table 1A. Genes Significantly Altered in A4573 ESFT
Supplementary Table 1A. Genes significantly altered in A4573 ESFT cells following BMI-1knockdown genesymbol genedescription siControl siBMI1 FC Direction P-value AASS aminoadipate-semialdehyde synthase | tetra-peptide repeat homeobox-like6.68 7.24 1.5 Up 0.007 ABCA2 ATP-binding cassette, sub-family A (ABC1), member 2 | neural5.44 proliferation,6.3 differentiation1.8 and Upcontrol, 1 0.006 ABHD4 abhydrolase domain containing 4 7.51 6.69 1.8 Down 0.002 ACACA acetyl-Coenzyme A carboxylase alpha | peroxiredoxin 5 | similar6.2 to High mobility7.26 group2.1 protein UpB1 (High mobility0.009 group protein 1) (HMG-1) (Amphoterin) (Heparin-binding protein p30) | Coenzyme A synthase ACAD9 acyl-Coenzyme A dehydrogenase family, member 9 9.25 8.59 1.6 Down 0.008 ACBD3 acyl-Coenzyme A binding domain containing 3 7.89 8.53 1.6 Up 0.008 ACCN2 amiloride-sensitive cation channel 2, neuronal 5.47 6.28 1.8 Up 0.005 ACIN1 apoptotic chromatin condensation inducer 1 7.15 7.79 1.6 Up 0.008 ACPL2 acid phosphatase-like 2 6.04 7.6 2.9 Up 0.000 ACSL4 acyl-CoA synthetase long-chain family member 4 6.72 5.8 1.9 Down 0.001 ACTA2 actin, alpha 2, smooth muscle, aorta 9.18 8.44 1.7 Down 0.003 ACYP1 acylphosphatase 1, erythrocyte (common) type 7.09 7.66 1.5 Up 0.009 ADA adenosine deaminase 6.34 7.1 1.7 Up 0.009 ADAL adenosine deaminase-like 7.88 6.89 2.0 Down 0.006 ADAMTS1 ADAM metallopeptidase with thrombospondin type 1 motif, 1 6.57 7.65 2.1 Up 0.000 ADARB1 adenosine deaminase, RNA-specific, B1 (RED1 homolog rat) 6.49 7.13 1.6 Up 0.008 ADCY9 adenylate cyclase 9 6.5 7.18 -

Exome Sequencing Reveals Independent SGCD Deletions Causing Limb Girdle Muscular Dystrophy in Boston Terriers Cox Et Al
Exome sequencing reveals independent SGCD deletions causing limb girdle muscular dystrophy in Boston terriers Cox et al. Cox et al. Skeletal Muscle (2017) 7:15 DOI 10.1186/s13395-017-0131-0 Cox et al. Skeletal Muscle (2017) 7:15 DOI 10.1186/s13395-017-0131-0 RESEARCH Open Access Exome sequencing reveals independent SGCD deletions causing limb girdle muscular dystrophy in Boston terriers Melissa L. Cox1†, Jacquelyn M. Evans2†, Alexander G. Davis2, Ling T. Guo3, Jennifer R. Levy4,5, Alison N. Starr-Moss2, Elina Salmela6,7, Marjo K. Hytönen6,7, Hannes Lohi6,7, Kevin P. Campbell4,5, Leigh Anne Clark2* and G. Diane Shelton3* Abstract Background: Limb-girdle muscular dystrophies (LGMDs) are a heterogeneous group of inherited autosomal myopathies that preferentially affect voluntary muscles of the shoulders and hips. LGMD has been clinically described in several breeds of dogs, but the responsible mutations are unknown. The clinical presentation in dogs is characterized by marked muscle weakness and atrophy in the shoulder and hips during puppyhood. Methods: Following clinical evaluation, the identification of the dystrophic histological phenotype on muscle histology, and demonstration of the absence of sarcoglycan-sarcospan complex by immunostaining, whole exome sequencing was performed on five Boston terriers: one affected dog and its three family members and one unrelated affected dog. Results: Within sarcoglycan-δ (SGCD), a two base pair deletion segregating with LGMD in the family was discovered, and a deletion encompassing exons 7 and 8 was found in the unrelated dog. Both mutations are predicted to cause an absence of SGCD protein, confirmed by immunohistochemistry. The mutations are private to each family. -

In Vivo and in Vitro Analysis of Dll1 and Pax6 Function in the Adult Mouse Pancreas
TECHNISCHE UNIVERSITÄT MÜNCHEN Lehrstuhl für Experimentelle Genetik In vivo and in vitro analysis of Dll1 and Pax6 function in the adult mouse pancreas Davide Cavanna Vollständiger Abdruck der von der Fakultät Wissenschaftszentrum Weihenstephan für Ernährung, Landnutzung und Umwelt der Technischen Universität München zur Erlangung des akademischen Grades eines Doktors der Naturwissenschaften genehmigten Dissertation. Vorsitzender: Univ.-Prof. Dr. D. Langosch Prüfer der Dissertation: 1. Univ.-Prof. Dr. M. Hrabé de Angelis 2. Univ.-Prof. A. Schnieke, Ph.D. Die Dissertation wurde am 03.07.2013 bei der Technischen Universität München eingereicht und durch die Fakultät Wissenschaftszentrum Weihenstephan für Ernährung, Landnutzung und Umwelt am 10.12.2013 angenommen. I. Table of contents I. TABLE OF CONTENTS .................................................................................................. I II. FIGURES AND TABLES ................................................................................................ V III. ABBREVIATIONS ................................................................................................. VIII IV. PUBLICATIONS, TALKS, AND POSTERS ................................................................... XI V. ACKNOWLEDGMENTS .............................................................................................. XII VI. AFFIRMATION ..................................................................................................... XIV 1. SUMMARY/ZUSAMMENFASSUNG ............................................................................ -

The Effect of Mechanical Force on Gene Expression of Human Bladder Smooth Muscle Cells" (2012)
Philadelphia College of Osteopathic Medicine DigitalCommons@PCOM PCOM Biomedical Studies Student Scholarship Student Dissertations, Theses and Papers 6-2012 The ffecE t of Mechanical Force on Gene Expression of Human Bladder Smooth Muscle Cells Christopher A. Callan Philadelphia College of Osteopathic Medicine, [email protected] Follow this and additional works at: http://digitalcommons.pcom.edu/biomed Part of the Molecular Genetics Commons, and the Urogenital System Commons Recommended Citation Callan, Christopher A., "The Effect of Mechanical Force on Gene Expression of Human Bladder Smooth Muscle Cells" (2012). PCOM Biomedical Studies Student Scholarship. Paper 36. This Thesis is brought to you for free and open access by the Student Dissertations, Theses and Papers at DigitalCommons@PCOM. It has been accepted for inclusion in PCOM Biomedical Studies Student Scholarship by an authorized administrator of DigitalCommons@PCOM. For more information, please contact [email protected]. PHILADELPHIA COLLEGE OF OSTEOPATHIC MEDICINE Philadelphia, Pennsylvania The Effect Of Mechanical Force on Gene Expression of Human Bladder Smooth Muscle Cells A thesis submitted in partial fulfillnment of the requirements for the degree of MASTER OF BIOMEDICAL SCIENCE by Christopher A. Callan June 2012 We approve the thesis of Christopher A. Callan _______________________________________________________________________ Edward J. Macarak, Ph.D. Date Chairman; Department of Anatomy and Cell Biology University of Pennsylvania School of Dental Medicine Thesis -

Figure S1. Gene Ontology Classification of Abeliophyllum Distichum Leaves Extract-Induced Degs
Figure S1. Gene ontology classification of Abeliophyllum distichum leaves extract-induced DEGs. The results are summarized in three main categories: Biological process, Cellular component and Molecular function. Figure S2. KEGG pathway enrichment analysis using Abeliophyllum distichum leaves extract-DEGs (A). Venn diagram analysis of DEGs involved in PI3K/Akt signaling pathway and Rap1 signaling pathway (B). Figure S3. The expression (A) and protein levels (B) of Akt3 in AL-treated SK-MEL2 cells. Values with different superscripted letters are significantly different (p < 0.05). Table S1. Abeliophyllum distichum leaves extract-induced DEGs. log2 Fold Gene name Gene description Change A2ML1 alpha-2-macroglobulin-like protein 1 isoform 2 [Homo sapiens] 3.45 A4GALT lactosylceramide 4-alpha-galactosyltransferase [Homo sapiens] −1.64 ABCB4 phosphatidylcholine translocator ABCB4 isoform A [Homo sapiens] −1.43 ABCB5 ATP-binding cassette sub-family B member 5 isoform 1 [Homo sapiens] −2.99 ABHD17C alpha/beta hydrolase domain-containing protein 17C [Homo sapiens] −1.62 ABLIM2 actin-binding LIM protein 2 isoform 1 [Homo sapiens] −2.53 ABTB2 ankyrin repeat and BTB/POZ domain-containing protein 2 [Homo sapiens] −1.48 ACACA acetyl-CoA carboxylase 1 isoform 1 [Homo sapiens] −1.76 ACACB acetyl-CoA carboxylase 2 precursor [Homo sapiens] −2.03 ACSM1 acyl-coenzyme A synthetase ACSM1, mitochondrial [Homo sapiens] −3.05 disintegrin and metalloproteinase domain-containing protein 19 preproprotein [Homo ADAM19 −1.65 sapiens] disintegrin and metalloproteinase -

Agricultural University of Athens
ΓΕΩΠΟΝΙΚΟ ΠΑΝΕΠΙΣΤΗΜΙΟ ΑΘΗΝΩΝ ΣΧΟΛΗ ΕΠΙΣΤΗΜΩΝ ΤΩΝ ΖΩΩΝ ΤΜΗΜΑ ΕΠΙΣΤΗΜΗΣ ΖΩΙΚΗΣ ΠΑΡΑΓΩΓΗΣ ΕΡΓΑΣΤΗΡΙΟ ΓΕΝΙΚΗΣ ΚΑΙ ΕΙΔΙΚΗΣ ΖΩΟΤΕΧΝΙΑΣ ΔΙΔΑΚΤΟΡΙΚΗ ΔΙΑΤΡΙΒΗ Εντοπισμός γονιδιωματικών περιοχών και δικτύων γονιδίων που επηρεάζουν παραγωγικές και αναπαραγωγικές ιδιότητες σε πληθυσμούς κρεοπαραγωγικών ορνιθίων ΕΙΡΗΝΗ Κ. ΤΑΡΣΑΝΗ ΕΠΙΒΛΕΠΩΝ ΚΑΘΗΓΗΤΗΣ: ΑΝΤΩΝΙΟΣ ΚΟΜΙΝΑΚΗΣ ΑΘΗΝΑ 2020 ΔΙΔΑΚΤΟΡΙΚΗ ΔΙΑΤΡΙΒΗ Εντοπισμός γονιδιωματικών περιοχών και δικτύων γονιδίων που επηρεάζουν παραγωγικές και αναπαραγωγικές ιδιότητες σε πληθυσμούς κρεοπαραγωγικών ορνιθίων Genome-wide association analysis and gene network analysis for (re)production traits in commercial broilers ΕΙΡΗΝΗ Κ. ΤΑΡΣΑΝΗ ΕΠΙΒΛΕΠΩΝ ΚΑΘΗΓΗΤΗΣ: ΑΝΤΩΝΙΟΣ ΚΟΜΙΝΑΚΗΣ Τριμελής Επιτροπή: Aντώνιος Κομινάκης (Αν. Καθ. ΓΠΑ) Ανδρέας Κράνης (Eρευν. B, Παν. Εδιμβούργου) Αριάδνη Χάγερ (Επ. Καθ. ΓΠΑ) Επταμελής εξεταστική επιτροπή: Aντώνιος Κομινάκης (Αν. Καθ. ΓΠΑ) Ανδρέας Κράνης (Eρευν. B, Παν. Εδιμβούργου) Αριάδνη Χάγερ (Επ. Καθ. ΓΠΑ) Πηνελόπη Μπεμπέλη (Καθ. ΓΠΑ) Δημήτριος Βλαχάκης (Επ. Καθ. ΓΠΑ) Ευάγγελος Ζωίδης (Επ.Καθ. ΓΠΑ) Γεώργιος Θεοδώρου (Επ.Καθ. ΓΠΑ) 2 Εντοπισμός γονιδιωματικών περιοχών και δικτύων γονιδίων που επηρεάζουν παραγωγικές και αναπαραγωγικές ιδιότητες σε πληθυσμούς κρεοπαραγωγικών ορνιθίων Περίληψη Σκοπός της παρούσας διδακτορικής διατριβής ήταν ο εντοπισμός γενετικών δεικτών και υποψηφίων γονιδίων που εμπλέκονται στο γενετικό έλεγχο δύο τυπικών πολυγονιδιακών ιδιοτήτων σε κρεοπαραγωγικά ορνίθια. Μία ιδιότητα σχετίζεται με την ανάπτυξη (σωματικό βάρος στις 35 ημέρες, ΣΒ) και η άλλη με την αναπαραγωγική -
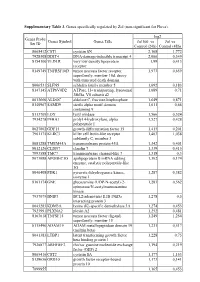
Supplementary Table 3. Genes Specifically Regulated by Zol (Non-Significant for Fluva)
Supplementary Table 3. Genes specifically regulated by Zol (non-significant for Fluva). log2 Genes Probe Genes Symbol Genes Title Zol100 vs Zol vs Set ID Control (24h) Control (48h) 8065412 CST1 cystatin SN 2,168 1,772 7928308 DDIT4 DNA-damage-inducible transcript 4 2,066 0,349 8154100 VLDLR very low density lipoprotein 1,99 0,413 receptor 8149749 TNFRSF10D tumor necrosis factor receptor 1,973 0,659 superfamily, member 10d, decoy with truncated death domain 8006531 SLFN5 schlafen family member 5 1,692 0,183 8147145 ATP6V0D2 ATPase, H+ transporting, lysosomal 1,689 0,71 38kDa, V0 subunit d2 8013660 ALDOC aldolase C, fructose-bisphosphate 1,649 0,871 8140967 SAMD9 sterile alpha motif domain 1,611 0,66 containing 9 8113709 LOX lysyl oxidase 1,566 0,524 7934278 P4HA1 prolyl 4-hydroxylase, alpha 1,527 0,428 polypeptide I 8027002 GDF15 growth differentiation factor 15 1,415 0,201 7961175 KLRC3 killer cell lectin-like receptor 1,403 1,038 subfamily C, member 3 8081288 TMEM45A transmembrane protein 45A 1,342 0,401 8012126 CLDN7 claudin 7 1,339 0,415 7993588 TMC7 transmembrane channel-like 7 1,318 0,3 8073088 APOBEC3G apolipoprotein B mRNA editing 1,302 0,174 enzyme, catalytic polypeptide-like 3G 8046408 PDK1 pyruvate dehydrogenase kinase, 1,287 0,382 isozyme 1 8161174 GNE glucosamine (UDP-N-acetyl)-2- 1,283 0,562 epimerase/N-acetylmannosamine kinase 7937079 BNIP3 BCL2/adenovirus E1B 19kDa 1,278 0,5 interacting protein 3 8043283 KDM3A lysine (K)-specific demethylase 3A 1,274 0,453 7923991 PLXNA2 plexin A2 1,252 0,481 8163618 TNFSF15 tumor necrosis