High-Resolution Analysis Identifies High Frequency of KIR-A
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Human and Mouse CD Marker Handbook Human and Mouse CD Marker Key Markers - Human Key Markers - Mouse
Welcome to More Choice CD Marker Handbook For more information, please visit: Human bdbiosciences.com/eu/go/humancdmarkers Mouse bdbiosciences.com/eu/go/mousecdmarkers Human and Mouse CD Marker Handbook Human and Mouse CD Marker Key Markers - Human Key Markers - Mouse CD3 CD3 CD (cluster of differentiation) molecules are cell surface markers T Cell CD4 CD4 useful for the identification and characterization of leukocytes. The CD CD8 CD8 nomenclature was developed and is maintained through the HLDA (Human Leukocyte Differentiation Antigens) workshop started in 1982. CD45R/B220 CD19 CD19 The goal is to provide standardization of monoclonal antibodies to B Cell CD20 CD22 (B cell activation marker) human antigens across laboratories. To characterize or “workshop” the antibodies, multiple laboratories carry out blind analyses of antibodies. These results independently validate antibody specificity. CD11c CD11c Dendritic Cell CD123 CD123 While the CD nomenclature has been developed for use with human antigens, it is applied to corresponding mouse antigens as well as antigens from other species. However, the mouse and other species NK Cell CD56 CD335 (NKp46) antibodies are not tested by HLDA. Human CD markers were reviewed by the HLDA. New CD markers Stem Cell/ CD34 CD34 were established at the HLDA9 meeting held in Barcelona in 2010. For Precursor hematopoetic stem cell only hematopoetic stem cell only additional information and CD markers please visit www.hcdm.org. Macrophage/ CD14 CD11b/ Mac-1 Monocyte CD33 Ly-71 (F4/80) CD66b Granulocyte CD66b Gr-1/Ly6G Ly6C CD41 CD41 CD61 (Integrin b3) CD61 Platelet CD9 CD62 CD62P (activated platelets) CD235a CD235a Erythrocyte Ter-119 CD146 MECA-32 CD106 CD146 Endothelial Cell CD31 CD62E (activated endothelial cells) Epithelial Cell CD236 CD326 (EPCAM1) For Research Use Only. -

Expression Tissue
The Journal of Immunology Killer Cell Ig-Like Receptor and Leukocyte Ig-Like Receptor Transgenic Mice Exhibit Tissue- and Cell-Specific Transgene Expression1 Danny Belkin,* Michaela Torkar,* Chiwen Chang,* Roland Barten,* Mauro Tolaini,† Anja Haude,* Rachel Allen,* Michael J. Wilson,* Dimitris Kioussis,† and John Trowsdale2* To generate an experimental model for exploring the function, expression pattern, and developmental regulation of human Ig-like activating and inhibitory receptors, we have generated transgenic mice using two human genomic clones: 52N12 (a 150-Kb clone encompassing the leukocyte Ig-like receptor (LILR)B1 (ILT2), LILRB4 (ILT3), and LILRA1 (LIR6) genes) and 1060P11 (a 160-Kb clone that contains ten killer cell Ig-like receptor (KIR) genes). Both the KIR and LILR families are encoded within the leukocyte receptor complex, and are involved in immune modulation. We have also produced a novel mAb to LILRA1 to facilitate expression studies. The LILR transgenes were expressed in a similar, but not identical, pattern to that observed in humans: LILRB1 was expressed in B cells, most NK cells, and a small number of T cells; LILRB4 was expressed in a B cell subset; and LILRA1 was found on a ring of cells surrounding B cell areas on spleen sections, consistent with other data showing monocyte/macrophage expression. KIR transgenic mice showed KIR2DL2 expression on a subset of NK cells and T cells, similar to the pattern seen in humans, and expression of KIR2DL4, KIR3DS1, and KIR2DL5 by splenic NK cells. These observations indicate that linked regulatory elements within the genomic clones are sufficient to allow appropriate expression of KIRs in mice, and illustrate that the presence of the natural ligands for these receptors, in the form of human MHC class I proteins, is not necessary for the expression of the KIRs observed in these mice. -
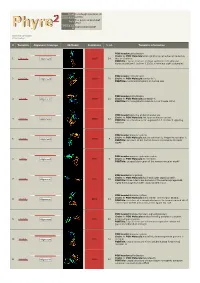
Phyre 2 Results for A1YIY0
Email [email protected] Description A1YIY0 Tue Jul 30 13:19:15 BST Date 2013 Unique Job 1035bc4b501530df ID Detailed template information # Template Alignment Coverage 3D Model Confidence % i.d. Template Information PDB header:cell adhesion Chain: A: PDB Molecule:down syndrome cell adhesion molecule 1 c3dmkA_ Alignment 100.0 14 (dscam) isoform PDBTitle: crystal structure of down syndrome cell adhesion molecule (dscam)2 isoform 1.30.30, n-terminal eight ig domains PDB header:cell adhesion 2 c2om5A_ Alignment 100.0 20 Chain: A: PDB Molecule:contactin 2; PDBTitle: n-terminal fragment of human tax1 PDB header:cell adhesion 3 c3jxaA_ Alignment 100.0 21 Chain: A: PDB Molecule:contactin 4; PDBTitle: immunoglobulin domains 1-4 of mouse cntn4 PDB header:signaling protein/transferase Chain: A: PDB Molecule:tek tyrosine kinase variant; 4 c4k0vA_ 100.0 12 Alignment PDBTitle: structural basis for angiopoietin-1 mediated signaling initiation PDB header:immune system Chain: A: PDB Molecule:natural cytotoxicity triggering receptor 1; 5 c1p6fA_ 100.0 9 Alignment PDBTitle: structure of the human natural cytotoxicity receptor nkp46 PDB header:immune system/receptor 6 c1ollA_ Alignment 99.9 9 Chain: A: PDB Molecule:nk receptor; PDBTitle: extracellular region of the human receptor nkp46 PDB header:viral protein Chain: A: PDB Molecule:hoc head outer capsid protein; 7 c3shsA_ 99.9 18 Alignment PDBTitle: three n-terminal domains of the bacteriophage rb49 highly immunogenic2 outer capsid protein (hoc) PDB header:immune system Chain: E: PDB Molecule:natural -

Supplementary Table 1: Adhesion Genes Data Set
Supplementary Table 1: Adhesion genes data set PROBE Entrez Gene ID Celera Gene ID Gene_Symbol Gene_Name 160832 1 hCG201364.3 A1BG alpha-1-B glycoprotein 223658 1 hCG201364.3 A1BG alpha-1-B glycoprotein 212988 102 hCG40040.3 ADAM10 ADAM metallopeptidase domain 10 133411 4185 hCG28232.2 ADAM11 ADAM metallopeptidase domain 11 110695 8038 hCG40937.4 ADAM12 ADAM metallopeptidase domain 12 (meltrin alpha) 195222 8038 hCG40937.4 ADAM12 ADAM metallopeptidase domain 12 (meltrin alpha) 165344 8751 hCG20021.3 ADAM15 ADAM metallopeptidase domain 15 (metargidin) 189065 6868 null ADAM17 ADAM metallopeptidase domain 17 (tumor necrosis factor, alpha, converting enzyme) 108119 8728 hCG15398.4 ADAM19 ADAM metallopeptidase domain 19 (meltrin beta) 117763 8748 hCG20675.3 ADAM20 ADAM metallopeptidase domain 20 126448 8747 hCG1785634.2 ADAM21 ADAM metallopeptidase domain 21 208981 8747 hCG1785634.2|hCG2042897 ADAM21 ADAM metallopeptidase domain 21 180903 53616 hCG17212.4 ADAM22 ADAM metallopeptidase domain 22 177272 8745 hCG1811623.1 ADAM23 ADAM metallopeptidase domain 23 102384 10863 hCG1818505.1 ADAM28 ADAM metallopeptidase domain 28 119968 11086 hCG1786734.2 ADAM29 ADAM metallopeptidase domain 29 205542 11085 hCG1997196.1 ADAM30 ADAM metallopeptidase domain 30 148417 80332 hCG39255.4 ADAM33 ADAM metallopeptidase domain 33 140492 8756 hCG1789002.2 ADAM7 ADAM metallopeptidase domain 7 122603 101 hCG1816947.1 ADAM8 ADAM metallopeptidase domain 8 183965 8754 hCG1996391 ADAM9 ADAM metallopeptidase domain 9 (meltrin gamma) 129974 27299 hCG15447.3 ADAMDEC1 ADAM-like, -

Flow Reagents Single Color Antibodies CD Chart
CD CHART CD N° Alternative Name CD N° Alternative Name CD N° Alternative Name Beckman Coulter Clone Beckman Coulter Clone Beckman Coulter Clone T Cells B Cells Granulocytes NK Cells Macrophages/Monocytes Platelets Erythrocytes Stem Cells Dendritic Cells Endothelial Cells Epithelial Cells T Cells B Cells Granulocytes NK Cells Macrophages/Monocytes Platelets Erythrocytes Stem Cells Dendritic Cells Endothelial Cells Epithelial Cells T Cells B Cells Granulocytes NK Cells Macrophages/Monocytes Platelets Erythrocytes Stem Cells Dendritic Cells Endothelial Cells Epithelial Cells CD1a T6, R4, HTA1 Act p n n p n n S l CD99 MIC2 gene product, E2 p p p CD223 LAG-3 (Lymphocyte activation gene 3) Act n Act p n CD1b R1 Act p n n p n n S CD99R restricted CD99 p p CD224 GGT (γ-glutamyl transferase) p p p p p p CD1c R7, M241 Act S n n p n n S l CD100 SEMA4D (semaphorin 4D) p Low p p p n n CD225 Leu13, interferon induced transmembrane protein 1 (IFITM1). p p p p p CD1d R3 Act S n n Low n n S Intest CD101 V7, P126 Act n p n p n n p CD226 DNAM-1, PTA-1 Act n Act Act Act n p n CD1e R2 n n n n S CD102 ICAM-2 (intercellular adhesion molecule-2) p p n p Folli p CD227 MUC1, mucin 1, episialin, PUM, PEM, EMA, DF3, H23 Act p CD2 T11; Tp50; sheep red blood cell (SRBC) receptor; LFA-2 p S n p n n l CD103 HML-1 (human mucosal lymphocytes antigen 1), integrin aE chain S n n n n n n n l CD228 Melanotransferrin (MT), p97 p p CD3 T3, CD3 complex p n n n n n n n n n l CD104 integrin b4 chain; TSP-1180 n n n n n n n p p CD229 Ly9, T-lymphocyte surface antigen p p n p n -

A Structural Analysis of Mide Chants
A Structural Analysis of Mide Chants GEORGE FULFORD McMaster University Introduction In this paper I shall investigate the relationship between words and im agery in seven song scrolls used by members of an Ojibwa religious society known as the Midewiwin. These texts were collected in the late 1880s by W.J. Hoffman for the Bureau of American Ethnology and subsequently published in their seventh Annual Report (Hoffman 1891).1 All the pictographs which I shall discuss were inscribed on birch bark and used by members of the Midewiwin to record chants used in their ceremonies. According to Hoffman (1891:192) these chants consisted of only a few words or short phrases. They were sung by single individuals — never in chorus — and were repeated over and over again, usually to the accompaniment of a wooden kettle drum. In a previous study (Fulford 1989) I analyzed patterns of structural variation among these pictographs and outlined how three complex symbols — the otter, bear and bird — evolved from clan emblems into pictographic markers. The focus of my earlier study was purely iconographic; in this paper I shall explore the verbal structure of Midewiwin chants in order to show some of the ways in which they were pictographically encoded. For the sake of convenience, I have limited my discussion to song scrolls sharing the otter symbol. Six of the seven scrolls that I shall examine contain this marking device. Although one (designated Scroll C in the appendix) lacks an otter, it displays many other formal similarities with Hoffman published transcriptions and translations of 23 songs performed at the White Earth Reservation in northwestern Minnesota. -

5892 Cisco Category: Standards Track August 2010 ISSN: 2070-1721
Internet Engineering Task Force (IETF) P. Faltstrom, Ed. Request for Comments: 5892 Cisco Category: Standards Track August 2010 ISSN: 2070-1721 The Unicode Code Points and Internationalized Domain Names for Applications (IDNA) Abstract This document specifies rules for deciding whether a code point, considered in isolation or in context, is a candidate for inclusion in an Internationalized Domain Name (IDN). It is part of the specification of Internationalizing Domain Names in Applications 2008 (IDNA2008). Status of This Memo This is an Internet Standards Track document. This document is a product of the Internet Engineering Task Force (IETF). It represents the consensus of the IETF community. It has received public review and has been approved for publication by the Internet Engineering Steering Group (IESG). Further information on Internet Standards is available in Section 2 of RFC 5741. Information about the current status of this document, any errata, and how to provide feedback on it may be obtained at http://www.rfc-editor.org/info/rfc5892. Copyright Notice Copyright (c) 2010 IETF Trust and the persons identified as the document authors. All rights reserved. This document is subject to BCP 78 and the IETF Trust's Legal Provisions Relating to IETF Documents (http://trustee.ietf.org/license-info) in effect on the date of publication of this document. Please review these documents carefully, as they describe your rights and restrictions with respect to this document. Code Components extracted from this document must include Simplified BSD License text as described in Section 4.e of the Trust Legal Provisions and are provided without warranty as described in the Simplified BSD License. -

Kyrillische Schrift Für Den Computer
Hanna-Chris Gast Kyrillische Schrift für den Computer Benennung der Buchstaben, Vergleich der Transkriptionen in Bibliotheken und Standesämtern, Auflistung der Unicodes sowie Tastaturbelegung für Windows XP Inhalt Seite Vorwort ................................................................................................................................................ 2 1 Kyrillische Schriftzeichen mit Benennung................................................................................... 3 1.1 Die Buchstaben im Russischen mit Schreibschrift und Aussprache.................................. 3 1.2 Kyrillische Schriftzeichen anderer slawischer Sprachen.................................................... 9 1.3 Veraltete kyrillische Schriftzeichen .................................................................................... 10 1.4 Die gebräuchlichen Sonderzeichen ..................................................................................... 11 2 Transliterationen und Transkriptionen (Umschriften) .......................................................... 13 2.1 Begriffe zum Thema Transkription/Transliteration/Umschrift ...................................... 13 2.2 Normen und Vorschriften für Bibliotheken und Standesämter....................................... 15 2.3 Tabellarische Übersicht der Umschriften aus dem Russischen ....................................... 21 2.4 Transliterationen veralteter kyrillischer Buchstaben ....................................................... 25 2.5 Transliterationen bei anderen slawischen -

Differential Integrin Adhesome Expression Defines Human Natural
bioRxiv preprint doi: https://doi.org/10.1101/2020.12.01.404806; this version posted December 1, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC 4.0 International license. Differential integrin adhesome expression defines human natural killer cell residency and developmental stage Everardo Hegewisch Solloa1, Seungmae Seo1, Bethany L. Mundy-Bosse2, Anjali Mishra3,a, Erik Waldman4,b, Sarah Maurrasse4,b, Eli Grunstein4, Thomas J. Connors5, Aharon G. Freud2,6, and Emily M. Mace1 1Department of Pediatrics, Columbia University College of Physicians and Surgeons, New York NY 10032 2Division of Hematology, Department of Internal Medicine, The Ohio State University, Columbus, OH 43210, USA; Comprehensive Cancer Center and The James Cancer Hospital and Solove Research Institute, The Ohio State University, Columbus, OH 43210 33Division of Dermatology, Department of Internal Medicine, The Ohio State University, Columbus, OH 43210, USA; Comprehensive Cancer Center and The James Cancer Hospital and Solove Research Institute, The Ohio State University, Columbus, OH 43210 4Department of Otolaryngology - Head and Neck Surgery, Columbia University Medical Center, New York, New York 10032 5Department of Pediatrics, Division of Pediatric Critical Care and Hospital Medicine, Columbia University Irving Medical Center, New York, NY 10024 6Department of Pathology, The Ohio State University, Columbus, -

Snipa Snpcard
SNiPAcard Block annotations Block info genomic range chr19:55,117,749-55,168,602 block size 50,854 bp variant count 74 variants Basic features Conservation/deleteriousness Linked genes μ = -0.557 [-4.065 – AC009892.10 , AC009892.5 , AC009892.9 , AC245036.1 , AC245036.2 , phyloP 2.368] gene(s) hit or close-by AC245036.3 , AC245036.4 , AC245036.5 , AC245036.6 , LILRA1 , LILRB1 , LILRB4 , MIR8061 , VN1R105P phastCons μ = 0.059 [0 – 0.633] eQTL gene(s) CTB-83J4.2 , LILRA1 , LILRA2 , LILRB2 , LILRB5 , LILRP1 μ = -0.651 [-4.69 – 2.07] AC008984.5 , AC008984.5 , AC008984.6 , AC008984.6 , AC008984.7 , AC008984.7 , AC009892.10 , AC009892.10 , AC009892.2 , AC009892.2 , AC009892.5 , AC010518.3 , AC010518.3 , AC011515.2 , AC011515.2 , AC012314.19 , AC012314.19 , FCAR , FCAR , FCAR , FCAR , FCAR , FCAR , FCAR , FCAR , FCAR , FCAR , KIR2DL1 , KIR2DL1 , KIR2DL1 , KIR2DL1 , KIR2DL1 , KIR2DL1 , KIR2DL1 , KIR2DL1 , KIR2DL1 , KIR2DL1 , KIR2DL1 , KIR2DL1 , KIR2DL1 , KIR2DL1 , KIR2DL1 , KIR2DL1 , KIR2DL1 , KIR2DL1 , KIR2DL1 , KIR2DL1 , KIR2DL1 , KIR2DL1 , KIR2DL1 , KIR2DL1 , KIR2DL1 , KIR2DL3 , KIR2DL3 , KIR2DL3 , KIR2DL3 , KIR2DL3 , KIR2DL3 , KIR2DL3 , KIR2DL3 , KIR2DL3 , KIR2DL3 , KIR2DL3 , KIR2DL3 , KIR2DL3 , KIR2DL3 , KIR2DL3 , KIR2DL3 , KIR2DL3 , KIR2DL3 , KIR2DL3 , KIR2DL3 , KIR2DL4 , KIR2DL4 , KIR2DL4 , KIR2DL4 , KIR2DL4 , KIR2DL4 , KIR2DL4 , KIR2DL4 , KIR2DL4 , KIR2DL4 , KIR2DL4 , KIR2DL4 , KIR2DL4 , KIR2DL4 , KIR2DL4 , KIR2DL4 , KIR2DL4 , KIR2DL4 , KIR2DL4 , KIR2DL4 , KIR2DL4 , KIR2DL4 , KIR2DL4 , KIR2DL4 , KIR2DL4 , KIR2DL4 , KIR2DL4 , KIR2DL4 , -

An NK Cell-Activating Receptor with Inhibitory Potential Mathias Faure and Eric O
KIR2DL4 (CD158d), an NK Cell-Activating Receptor with Inhibitory Potential Mathias Faure and Eric O. Long This information is current as J Immunol 2002; 168:6208-6214; ; of September 25, 2021. doi: 10.4049/jimmunol.168.12.6208 http://www.jimmunol.org/content/168/12/6208 Downloaded from References This article cites 43 articles, 21 of which you can access for free at: http://www.jimmunol.org/content/168/12/6208.full#ref-list-1 Why The JI? Submit online. http://www.jimmunol.org/ • Rapid Reviews! 30 days* from submission to initial decision • No Triage! Every submission reviewed by practicing scientists • Fast Publication! 4 weeks from acceptance to publication *average by guest on September 25, 2021 Subscription Information about subscribing to The Journal of Immunology is online at: http://jimmunol.org/subscription Permissions Submit copyright permission requests at: http://www.aai.org/About/Publications/JI/copyright.html Email Alerts Receive free email-alerts when new articles cite this article. Sign up at: http://jimmunol.org/alerts The Journal of Immunology is published twice each month by The American Association of Immunologists, Inc., 1451 Rockville Pike, Suite 650, Rockville, MD 20852 Copyright © 2002 by The American Association of Immunologists All rights reserved. Print ISSN: 0022-1767 Online ISSN: 1550-6606. The Journal of Immunology KIR2DL4 (CD158d), an NK Cell-Activating Receptor with Inhibitory Potential Mathias Faure and Eric O. Long1 KIR2DL4 (CD158d) is an unusual member of the killer cell Ig-like receptor family expressed in all NK cells and some T cells. KIR2DL4 activates the cytotoxicity of NK cells, despite the presence of an immunoreceptor tyrosine-based inhibition motif (ITIM) in its cytoplasmic tail. -

Crystal Structure of the HLA-Cw3 Allotype-Specific Killer Cell Inhibitory Receptor KIR2DL2
Proc. Natl. Acad. Sci. USA Vol. 96, pp. 3864–3869, March 1999 Immunology Crystal structure of the HLA-Cw3 allotype-specific killer cell inhibitory receptor KIR2DL2 GREG A. SNYDER*, ANDREW G. BROOKS†, AND PETER D. SUN‡§ *Department of Biochemistry, Molecular Biology, and Cell Biology, 2153 Sheridan, O. T. Hogan Hall, Room 2-100, Northwestern University, Evanston, IL 60208; †Department of Microbiology and Immunology, University of Melbourne, Royal Parade, Parkville, 3052 Victoria, Australia; and ‡Structural Biology Section, Office of the Scientific Director, National Institute of Allergy and Infectious Diseases, National Institutes of Health, 12441 Parklawn Drive, Rockville, MD 20852 Edited by David R. Davies, National Institute of Diabetes and Digestive and Kidney Diseases, Bethesda, MD, and approved January 19, 1999 (received for review November 11, 1998) ABSTRACT Killer cell inhibitory receptors (KIR) protect differ primarily in the transmembrane and the intracellular class I HLAs expressing target cells from natural killer (NK) regions. KIR2DL molecules are inhibitory receptors possess- cell-mediated lysis. To understand the molecular basis of this ing intracellular immunoreceptor tyrosine-based inhibitory receptor-ligand recognition, we have crystallized the extra- motifs (ITIM), whereas KIR2DS molecules appear to be cellular ligand-binding domains of KIR2DL2, a member of the noninhibitory receptors with shorter intracellular tails that Ig superfamily receptors that recognize HLA-Cw1, 3, 7, and 8 interact with an immunoreceptor tyrosine-based activation allotypes. The structure was determined in two different motif (ITAM) containing signaling component DAP12 (11). crystal forms, an orthorhombic P212121 and a trigonal P3221 KIR3D members have three Ig domains (known as D0, D1, space group, to resolutions of 3.0 and 2.9 Å, respectively.