Laboratory Guidelines for Enumerating CD4 T Lymphocytes in the Context of HIV/AIDS SEA-HLM-392 Distribution: Limited
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Human and Mouse CD Marker Handbook Human and Mouse CD Marker Key Markers - Human Key Markers - Mouse
Welcome to More Choice CD Marker Handbook For more information, please visit: Human bdbiosciences.com/eu/go/humancdmarkers Mouse bdbiosciences.com/eu/go/mousecdmarkers Human and Mouse CD Marker Handbook Human and Mouse CD Marker Key Markers - Human Key Markers - Mouse CD3 CD3 CD (cluster of differentiation) molecules are cell surface markers T Cell CD4 CD4 useful for the identification and characterization of leukocytes. The CD CD8 CD8 nomenclature was developed and is maintained through the HLDA (Human Leukocyte Differentiation Antigens) workshop started in 1982. CD45R/B220 CD19 CD19 The goal is to provide standardization of monoclonal antibodies to B Cell CD20 CD22 (B cell activation marker) human antigens across laboratories. To characterize or “workshop” the antibodies, multiple laboratories carry out blind analyses of antibodies. These results independently validate antibody specificity. CD11c CD11c Dendritic Cell CD123 CD123 While the CD nomenclature has been developed for use with human antigens, it is applied to corresponding mouse antigens as well as antigens from other species. However, the mouse and other species NK Cell CD56 CD335 (NKp46) antibodies are not tested by HLDA. Human CD markers were reviewed by the HLDA. New CD markers Stem Cell/ CD34 CD34 were established at the HLDA9 meeting held in Barcelona in 2010. For Precursor hematopoetic stem cell only hematopoetic stem cell only additional information and CD markers please visit www.hcdm.org. Macrophage/ CD14 CD11b/ Mac-1 Monocyte CD33 Ly-71 (F4/80) CD66b Granulocyte CD66b Gr-1/Ly6G Ly6C CD41 CD41 CD61 (Integrin b3) CD61 Platelet CD9 CD62 CD62P (activated platelets) CD235a CD235a Erythrocyte Ter-119 CD146 MECA-32 CD106 CD146 Endothelial Cell CD31 CD62E (activated endothelial cells) Epithelial Cell CD236 CD326 (EPCAM1) For Research Use Only. -

Role of IL-4 Receptor &Alpha
Role of IL-4 receptor a–positive CD41 T cells in chronic airway hyperresponsiveness Frank Kirstein, PhD, Natalie E. Nieuwenhuizen, PhD,* Jaisubash Jayakumar, PhD, William G. C. Horsnell, PhD, and Frank Brombacher, PhD Cape Town, South Africa, and Berlin, Germany Background: TH2 cells and their cytokines are associated with IL-17–producing T cells and, consequently, increased airway allergic asthma in human subjects and with mouse models of neutrophilia. allergic airway disease. IL-4 signaling through the IL-4 receptor Conclusion: IL-4–responsive T helper cells are dispensable for 1 a (IL-4Ra) chain on CD4 T cells leads to TH2 cell acute OVA-induced airway disease but crucial in maintaining differentiation in vitro, implying that IL-4Ra–responsive CD41 chronic asthmatic pathology. (J Allergy Clin Immunol T cells are critical for the induction of allergic asthma. However, 2016;137:1852-62.) mechanisms regulating acute and chronic allergen-specific T 2 H Key words: responses in vivo remain incompletely understood. TH2 cell, acute allergic airway disease, chronic asthma, Objective: This study defines the requirements for IL-4Ra– cytokine receptors, IL-4, IL-13, gene-deficient mice responsive CD41 T cells and the IL-4Ra ligands IL-4 and IL-13 in the development of allergen-specific TH2 responses during the Allergic asthma is a chronic inflammatory disease of the airways onset and chronic phase of experimental allergic airway disease. characterized by an inappropriate immune response to harmless Methods: Development of acute and chronic ovalbumin environmental antigens. T 2 cells regulate adaptive immune 1 H (OVA)–induced allergic asthma was assessed weekly in CD4 T responses to allergens, and their presence correlates with disease 2 cell–specific IL-4Ra–deficient BALB/c mice (LckcreIL-4Ra /lox) symptoms in human subjects and mice.1 IL-4 plays a crucial role and respective control mice in the presence or absence of IL-4 in the in vitro and in vivo differentiation of TH2 cells, suggesting or IL-13. -

Introduction to Flow Cytometry Principles Data Analysis Protocols Troubleshooting
Flow Cytometry ipl.qxd 11/12/06 11:14 Page i Introduction to Flow Cytometry Principles Data analysis Protocols Troubleshooting By Misha Rahman, Ph.D. Technical advisors Andy Lane, Ph.D. Angie Swindell, M.Sc. Sarah Bartram, B.Sc. Your first choice for antibodies! Flow Cytometry ipl.qxd 11/12/06 11:14 Page ii Flow Cytometry ipl.qxd 11/12/06 11:14 Page iii Introduction to Flow Cytometry Principles Data analysis Protocols Troubleshooting By Misha Rahman, Ph.D. Technical advisors Andy Lane, Ph.D. Angie Swindell, M.Sc. Sarah Bartram, B.Sc. Flow Cytometry ipl.qxd 11/12/06 11:14 Page 2 Preface How can I explain what flow cytometry is to someone that knows nothing about it? Well, imagine it to be a lot like visiting a supermarket. You choose the goods you want and take them to the cashier. Usually you have to pile them onto a conveyor. The clerk picks up one item at a time and interrogates it with a laser to read the barcode. Once identified and if sense prevails, similar goods are collected together e.g. fruit and vegetables go into one shopping bag and household goods into another. Now picture in your mind the whole process automated; replace shopping with biological cells; and substitute the barcode with cellular markers – welcome to the world of flow cytometry and cell sorting! We aim to give you a basic overview of all the important facets of flow cytometry without delving too deeply into the complex mathematics and physics behind it all. For that there are other books (some recommended at the back). -
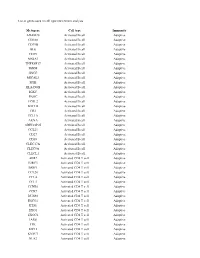
List of Genes Used in Cell Type Enrichment Analysis
List of genes used in cell type enrichment analysis Metagene Cell type Immunity ADAM28 Activated B cell Adaptive CD180 Activated B cell Adaptive CD79B Activated B cell Adaptive BLK Activated B cell Adaptive CD19 Activated B cell Adaptive MS4A1 Activated B cell Adaptive TNFRSF17 Activated B cell Adaptive IGHM Activated B cell Adaptive GNG7 Activated B cell Adaptive MICAL3 Activated B cell Adaptive SPIB Activated B cell Adaptive HLA-DOB Activated B cell Adaptive IGKC Activated B cell Adaptive PNOC Activated B cell Adaptive FCRL2 Activated B cell Adaptive BACH2 Activated B cell Adaptive CR2 Activated B cell Adaptive TCL1A Activated B cell Adaptive AKNA Activated B cell Adaptive ARHGAP25 Activated B cell Adaptive CCL21 Activated B cell Adaptive CD27 Activated B cell Adaptive CD38 Activated B cell Adaptive CLEC17A Activated B cell Adaptive CLEC9A Activated B cell Adaptive CLECL1 Activated B cell Adaptive AIM2 Activated CD4 T cell Adaptive BIRC3 Activated CD4 T cell Adaptive BRIP1 Activated CD4 T cell Adaptive CCL20 Activated CD4 T cell Adaptive CCL4 Activated CD4 T cell Adaptive CCL5 Activated CD4 T cell Adaptive CCNB1 Activated CD4 T cell Adaptive CCR7 Activated CD4 T cell Adaptive DUSP2 Activated CD4 T cell Adaptive ESCO2 Activated CD4 T cell Adaptive ETS1 Activated CD4 T cell Adaptive EXO1 Activated CD4 T cell Adaptive EXOC6 Activated CD4 T cell Adaptive IARS Activated CD4 T cell Adaptive ITK Activated CD4 T cell Adaptive KIF11 Activated CD4 T cell Adaptive KNTC1 Activated CD4 T cell Adaptive NUF2 Activated CD4 T cell Adaptive PRC1 Activated -

T Cells the Usual Subsets
T cells: the usual subsets Chen Dong and Gustavo J. Martinez T cells have important roles in immune responses and function by directly secreting soluble mediators or important for adaptation of immune responses in different microenvironments and might be particularly through cell contact-dependent mechanisms. Many T cell subsets have been characterized. Although relevant for host defence against pathogens that colonize different tissues. Distinct T cell subsets, or effector T cells were originally considered to be terminally differentiated, a growing body of evidence has differentiation states, can be identified based on the cell surface markers expressed and/or the effector challenged this view and suggested that the phenotype of effector T cells is not completely fixed but is molecules produced by a particular T cell population. This Poster summarizes our current understanding of more flexible or plastic. T cells can have ‘mixed’ phenotypes (that is, have characteristics usually the surface markers, transcriptional regulators, effector molecules and functions of the different T cell associated with more than one T cell subset) and can interconvert from one subset phenotype to another, subsets that participate in immune responses. Further knowledge of how these T cell subsets are regulated IMMUNOLOGY although instructive signalling can lead to long-term fixation of cytokine memory. T cell plasticity can be and cooperate with each other will provide us with better tools to treat immune-related diseases. Cytotoxic T cell Exhausted T cell -

Galectin-4 Interaction with CD14 Triggers the Differentiation of Monocytes Into Macrophage-Like Cells Via the MAPK Signaling Pathway
Immune Netw. 2019 Jun;19(3):e17 https://doi.org/10.4110/in.2019.19.e17 pISSN 1598-2629·eISSN 2092-6685 Original Article Galectin-4 Interaction with CD14 Triggers the Differentiation of Monocytes into Macrophage-like Cells via the MAPK Signaling Pathway So-Hee Hong 1,2,3,4,5, Jun-Seop Shin 1,2,3,5, Hyunwoo Chung 1,2,4,5, Chung-Gyu Park 1,2,3,4,5,* 1Xenotransplantation Research Center, Seoul National University College of Medicine, Seoul 03080, Korea 2Institute of Endemic Diseases, Seoul National University College of Medicine, Seoul 03080, Korea 3Cancer Research Institute, Seoul National University College of Medicine, Seoul 03080, Korea 4Department of Biomedical Sciences, Seoul National University College of Medicine, Seoul 03080, Korea 5Department of Microbiology and Immunology, Seoul National University College of Medicine, Seoul 03080, Korea Received: Jan 28, 2019 ABSTRACT Revised: May 13, 2019 Accepted: May 19, 2019 Galectin-4 (Gal-4) is a β-galactoside-binding protein mostly expressed in the gastrointestinal *Correspondence to tract of animals. Although intensive functional studies have been done for other galectin Chung-Gyu Park isoforms, the immunoregulatory function of Gal-4 still remains ambiguous. Here, we Department of Microbiology and Immunology, Seoul National University College of Medicine, demonstrated that Gal-4 could bind to CD14 on monocytes and induce their differentiation 103 Daehak-ro, Jongno-gu, Seoul 03080, into macrophage-like cells through the MAPK signaling pathway. Gal-4 induced the phenotypic Korea. changes on monocytes by altering the expression of various surface molecules, and induced E-mail: [email protected] functional changes such as increased cytokine production and matrix metalloproteinase expression and reduced phagocytic capacity. -

Comparison of Count Reproducibility, Accuracy, and Time to Results
Comparison of Count Reproducibility, Accuracy, and Time to Results between a Hemocytometer and the TC20™ Automated Cell Counter Tech Frank Hsiung, Tom McCollum, Eli Hefner, and Teresa Rubio Note Bio-Rad Laboratories, Inc., Hercules, CA 94547 USA Cell Counting Bulletin 6003 Introduction Bead counts were performed by sequentially loading For over 100 years the hemocytometer has been used by and counting the same chamber of a Bright-Line glass cell biologists to quantitate cells. It was first developed for the hemocytometer (Hausser Scientific). This was repeated ten quantitation of blood cells but became a popular and effective times. The number of beads was recorded for all nine tool for counting a variety of cell types, particles, and even 1 x 1 mm grids. small organisms. Currently, hemocytometers, armed with Flow Cytometry improved Neubauer grids, are a mainstay of cell biology labs. Flow cytometry was performed using a BD FACSCalibur flow Despite its longevity and versatility, hemocytometer counting cytometer (BD Biosciences) and CountBright counting beads suffers from a variety of shortcomings. These shortcomings (Life Technologies Corporation). Medium containing 50,000 include, but are not limited to, a lack of statistical robustness at CountBright beads was combined one-to-one with 250 µl low sample concentration, poor counts due to device misuse, of cells in suspension, yielding a final solution containing and subjectivity of counts among users, in addition to a time- 100 beads/µl. This solution was run through the flow consuming and tedious operation. In recent years automated cytometer until 10,000 events were collected in the gate cell counting has become an attractive alternative to manual previously defined as appropriate for non-doublet beads in hemocytometer–based cell counting, offering more reliable the FSC x SSC channel. -

Single-Cell RNA Sequencing Demonstrates the Molecular and Cellular Reprogramming of Metastatic Lung Adenocarcinoma
ARTICLE https://doi.org/10.1038/s41467-020-16164-1 OPEN Single-cell RNA sequencing demonstrates the molecular and cellular reprogramming of metastatic lung adenocarcinoma Nayoung Kim 1,2,3,13, Hong Kwan Kim4,13, Kyungjong Lee 5,13, Yourae Hong 1,6, Jong Ho Cho4, Jung Won Choi7, Jung-Il Lee7, Yeon-Lim Suh8,BoMiKu9, Hye Hyeon Eum 1,2,3, Soyean Choi 1, Yoon-La Choi6,10,11, Je-Gun Joung1, Woong-Yang Park 1,2,6, Hyun Ae Jung12, Jong-Mu Sun12, Se-Hoon Lee12, ✉ ✉ Jin Seok Ahn12, Keunchil Park12, Myung-Ju Ahn 12 & Hae-Ock Lee 1,2,3,6 1234567890():,; Advanced metastatic cancer poses utmost clinical challenges and may present molecular and cellular features distinct from an early-stage cancer. Herein, we present single-cell tran- scriptome profiling of metastatic lung adenocarcinoma, the most prevalent histological lung cancer type diagnosed at stage IV in over 40% of all cases. From 208,506 cells populating the normal tissues or early to metastatic stage cancer in 44 patients, we identify a cancer cell subtype deviating from the normal differentiation trajectory and dominating the metastatic stage. In all stages, the stromal and immune cell dynamics reveal ontological and functional changes that create a pro-tumoral and immunosuppressive microenvironment. Normal resident myeloid cell populations are gradually replaced with monocyte-derived macrophages and dendritic cells, along with T-cell exhaustion. This extensive single-cell analysis enhances our understanding of molecular and cellular dynamics in metastatic lung cancer and reveals potential diagnostic and therapeutic targets in cancer-microenvironment interactions. 1 Samsung Genome Institute, Samsung Medical Center, Seoul 06351, Korea. -

The Past, Present and Future of Flow Cytometry in Central Nervous System Malignancies
Review The Past, Present and Future of Flow Cytometry in Central Nervous System Malignancies Evrysthenis Vartholomatos 1, George Vartholomatos 2, George A. Alexiou 1,3 and Georgios S. Markopoulos 1,2,* 1 Faculty of Medicine, Neurosurgical Institute, School of Health Sciences, University of Ioannina, 45110 Ioannina, Greece; [email protected] (E.V.); [email protected] (G.A.A.) 2 Haematology Laboratory-Unit of Molecular Biology, University Hospital of Ioannina, 45110 Ioannina, Greece; [email protected] 3 Department of Neurosurgery, University of Ioannina, 45110 Ioannina, Greece * Correspondence: [email protected] Abstract: Central nervous system malignancies (CNSMs) are categorized among the most aggressive and deadly types of cancer. The low median survival in patients with CNSMs is partly explained by the objective difficulties of brain surgeries as well as by the acquired chemoresistance of CNSM cells. Flow Cytometry is an analytical technique with the ability to quantify cell phenotype and to categorize cell populations on the basis of their characteristics. In the current review, we summarize the Flow Cytometry methodologies that have been used to study different phenotypic aspects of CNSMs. These include DNA content analysis for the determination of malignancy status and phenotypic characterization, as well as the methodologies used during the development of novel therapeutic agents. We conclude with the historical and current utility of Flow Cytometry in the field, and we propose how we can exploit current and possible future methodologies in the battle against this dreadful type of malignancy. Citation: Vartholomatos, E.; Keywords: central nervous system malignancies; flow cytometry; glioblastoma; intraoperative flow Vartholomatos, G.; Alexiou, G.A.; cytometry; phenotypic analysis; DNA content analysis Markopoulos, G.S. -

IL21R Expressing CD14+CD16+ Monocytes Expand in Multiple
Plasma Cell Disorders SUPPLEMENTARY APPENDIX IL21R expressing CD14 +CD16 + monocytes expand in multiple myeloma patients leading to increased osteoclasts Marina Bolzoni, 1 Domenica Ronchetti, 2,3 Paola Storti, 1,4 Gaetano Donofrio, 5 Valentina Marchica, 1,4 Federica Costa, 1 Luca Agnelli, 2,3 Denise Toscani, 1 Rosanna Vescovini, 1 Katia Todoerti, 6 Sabrina Bonomini, 7 Gabriella Sammarelli, 1,7 Andrea Vecchi, 8 Daniela Guasco, 1 Fabrizio Accardi, 1,7 Benedetta Dalla Palma, 1,7 Barbara Gamberi, 9 Carlo Ferrari, 8 Antonino Neri, 2,3 Franco Aversa 1,4,7 and Nicola Giuliani 1,4,7 1Myeloma Unit, Dept. of Medicine and Surgery, University of Parma; 2Dept. of Oncology and Hemato-Oncology, University of Milan; 3Hematology Unit, “Fondazione IRCCS Ca’ Granda”, Ospedale Maggiore Policlinico, Milan; 4CoreLab, University Hospital of Parma; 5Dept. of Medical-Veterinary Science, University of Parma; 6Laboratory of Pre-clinical and Translational Research, IRCCS-CROB, Referral Cancer Center of Basilicata, Rionero in Vulture; 7Hematology and BMT Center, University Hospital of Parma; 8Infectious Disease Unit, University Hospital of Parma and 9“Dip. Oncologico e Tecnologie Avanzate”, IRCCS Arcispedale Santa Maria Nuova, Reggio Emilia, Italy ©2017 Ferrata Storti Foundation. This is an open-access paper. doi:10.3324/haematol. 2016.153841 Received: August 5, 2016. Accepted: December 23, 2016. Pre-published: January 5, 2017. Correspondence: [email protected] SUPPLEMENTAL METHODS Immunophenotype of BM CD14+ in patients with monoclonal gammopathies. Briefly, 100 μl of total BM aspirate was incubated in the dark with anti-human HLA-DR-PE (clone L243; BD), anti-human CD14-PerCP-Cy 5.5, anti-human CD16-PE-Cy7 (clone B73.1; BD) and anti-human CD45-APC-H 7 (clone 2D1; BD) for 20 min. -

B Cell Checkpoints in Autoimmune Rheumatic Diseases
REVIEWS B cell checkpoints in autoimmune rheumatic diseases Samuel J. S. Rubin1,2,3, Michelle S. Bloom1,2,3 and William H. Robinson1,2,3* Abstract | B cells have important functions in the pathogenesis of autoimmune diseases, including autoimmune rheumatic diseases. In addition to producing autoantibodies, B cells contribute to autoimmunity by serving as professional antigen- presenting cells (APCs), producing cytokines, and through additional mechanisms. B cell activation and effector functions are regulated by immune checkpoints, including both activating and inhibitory checkpoint receptors that contribute to the regulation of B cell tolerance, activation, antigen presentation, T cell help, class switching, antibody production and cytokine production. The various activating checkpoint receptors include B cell activating receptors that engage with cognate receptors on T cells or other cells, as well as Toll-like receptors that can provide dual stimulation to B cells via co- engagement with the B cell receptor. Furthermore, various inhibitory checkpoint receptors, including B cell inhibitory receptors, have important functions in regulating B cell development, activation and effector functions. Therapeutically targeting B cell checkpoints represents a promising strategy for the treatment of a variety of autoimmune rheumatic diseases. Antibody- dependent B cells are multifunctional lymphocytes that contribute that serve as precursors to and thereby give rise to acti- cell- mediated cytotoxicity to the pathogenesis of autoimmune diseases -

Identification of 18 Immune Cell Subsets Using 13-Color Panel
Immunophenotyping Identification of 18 immune cell subsets in human blood using a 13-color panel Background Cell type Function Phenotype Flow cytometry has become the method of choice for Eosinophils Parasitic immunity CD45+, SSCmid/hi, CD14 –, CD16 –, CD19– immunophenotyping and identifying specific cellular + mid/hi subsets. Within seconds, it provides a thorough overview of Neutrophils Innate Immunity CD45 , SSC , CD14 –, CD16+, CD19– the major cell types that constitute a sample. Using multiple + mid markers simultaneously increases the number of parameters Classical Phagocytosis of CD45 , SSC , monocytes pathogens and CD14+, CD16– that can be analyzed per run and decreases the amount of antigen presentation starting material required to perform an assay. This can be Intermediate Phagocytosis of CD45+, SSCmid, critical for precious sample material and long-term immune- monocytes pathogens and CD14+, CD16mid monitoring studies. In this application note, we demonstrate antigen presentation 13-color immunophenotyping of human peripheral blood Non-classical Phagocytosis of CD45+, SSCmid, + + mononuclear cells (PBMCs) using the MACSQuant® Analyzer 16, monocytes pathogens and CD14 , CD16 antigen presentation a compact and reliable benchtop flow cytometer equipped + low + with three lasers. The markers selected allow for the Class-switched Adaptive immunity CD45 , SSC , CD19 , memory B cells CD27+, IgD–, CD14– simultaneous identification and analysis of 18 different cell Non-switched Adaptive immunity CD45+, SSClow, CD19+, populations, thus maximizing the amount of information that memory B cells CD27+, IgD+, CD14– can be retrieved from the sample material analyzed. This is Naive B cells Adaptive immunity – CD45+, SSClow, CD19+, critical when input material is limited, as is often the case for non-antigen CD27–, IgD+, CD14– pediatric or disease studies.