Downloaded from Oncogenomics Database Pob01p.Ncifcrf.Gov/Cgi-Bin/JK Pediatric Sarcoma Expression Downloaded from St
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

SUPPLEMENTARY MATERIAL Bone Morphogenetic Protein 4 Promotes
www.intjdevbiol.com doi: 10.1387/ijdb.160040mk SUPPLEMENTARY MATERIAL corresponding to: Bone morphogenetic protein 4 promotes craniofacial neural crest induction from human pluripotent stem cells SUMIYO MIMURA, MIKA SUGA, KAORI OKADA, MASAKI KINEHARA, HIROKI NIKAWA and MIHO K. FURUE* *Address correspondence to: Miho Kusuda Furue. Laboratory of Stem Cell Cultures, National Institutes of Biomedical Innovation, Health and Nutrition, 7-6-8, Saito-Asagi, Ibaraki, Osaka 567-0085, Japan. Tel: 81-72-641-9819. Fax: 81-72-641-9812. E-mail: [email protected] Full text for this paper is available at: http://dx.doi.org/10.1387/ijdb.160040mk TABLE S1 PRIMER LIST FOR QRT-PCR Gene forward reverse AP2α AATTTCTCAACCGACAACATT ATCTGTTTTGTAGCCAGGAGC CDX2 CTGGAGCTGGAGAAGGAGTTTC ATTTTAACCTGCCTCTCAGAGAGC DLX1 AGTTTGCAGTTGCAGGCTTT CCCTGCTTCATCAGCTTCTT FOXD3 CAGCGGTTCGGCGGGAGG TGAGTGAGAGGTTGTGGCGGATG GAPDH CAAAGTTGTCATGGATGACC CCATGGAGAAGGCTGGGG MSX1 GGATCAGACTTCGGAGAGTGAACT GCCTTCCCTTTAACCCTCACA NANOG TGAACCTCAGCTACAAACAG TGGTGGTAGGAAGAGTAAAG OCT4 GACAGGGGGAGGGGAGGAGCTAGG CTTCCCTCCAACCAGTTGCCCCAAA PAX3 TTGCAATGGCCTCTCAC AGGGGAGAGCGCGTAATC PAX6 GTCCATCTTTGCTTGGGAAA TAGCCAGGTTGCGAAGAACT p75 TCATCCCTGTCTATTGCTCCA TGTTCTGCTTGCAGCTGTTC SOX9 AATGGAGCAGCGAAATCAAC CAGAGAGATTTAGCACACTGATC SOX10 GACCAGTACCCGCACCTG CGCTTGTCACTTTCGTTCAG Suppl. Fig. S1. Comparison of the gene expression profiles of the ES cells and the cells induced by NC and NC-B condition. Scatter plots compares the normalized expression of every gene on the array (refer to Table S3). The central line -

Transcriptional and Post-Transcriptional Regulation of ATP-Binding Cassette Transporter Expression
Transcriptional and Post-transcriptional Regulation of ATP-binding Cassette Transporter Expression by Aparna Chhibber DISSERTATION Submitted in partial satisfaction of the requirements for the degree of DOCTOR OF PHILOSOPHY in Pharmaceutical Sciences and Pbarmacogenomies in the Copyright 2014 by Aparna Chhibber ii Acknowledgements First and foremost, I would like to thank my advisor, Dr. Deanna Kroetz. More than just a research advisor, Deanna has clearly made it a priority to guide her students to become better scientists, and I am grateful for the countless hours she has spent editing papers, developing presentations, discussing research, and so much more. I would not have made it this far without her support and guidance. My thesis committee has provided valuable advice through the years. Dr. Nadav Ahituv in particular has been a source of support from my first year in the graduate program as my academic advisor, qualifying exam committee chair, and finally thesis committee member. Dr. Kathy Giacomini graciously stepped in as a member of my thesis committee in my 3rd year, and Dr. Steven Brenner provided valuable input as thesis committee member in my 2nd year. My labmates over the past five years have been incredible colleagues and friends. Dr. Svetlana Markova first welcomed me into the lab and taught me numerous laboratory techniques, and has always been willing to act as a sounding board. Michael Martin has been my partner-in-crime in the lab from the beginning, and has made my days in lab fly by. Dr. Yingmei Lui has made the lab run smoothly, and has always been willing to jump in to help me at a moment’s notice. -

Rapid Evolution of Mammalian X-Linked Testis-Expressed Homeobox Genes
Copyright 2004 by the Genetics Society of America DOI: 10.1534/genetics.103.025072 Rapid Evolution of Mammalian X-Linked Testis-Expressed Homeobox Genes Xiaoxia Wang and Jianzhi Zhang1 Department of Ecology and Evolutionary Biology, University of Michigan, Ann Arbor, Michigan 48109 Manuscript received November 26, 2003 Accepted for publication February 11, 2004 ABSTRACT Homeobox genes encode transcription factors that function in various developmental processes and are usually evolutionarily conserved in their sequences. However, two X-chromosome-linked testis-expressed homeobox genes, one from rodents and the other from fruit flies, are known to evolve rapidly under positive Darwinian selection. Here we report yet another case, from primates. TGIFLX is an X-linked homeobox gene that originated by retroposition of the autosomal gene TGIF2, most likely in a common ancestor of rodents and primates. While TGIF2 is ubiquitously expressed, TGIFLX is exclusively expressed in adult testis. A comparison of the TGIFLX sequences among 16 anthropoid primates revealed a signifi- cantly higher rate of nonsynonymous nucleotide substitution (dN) than synonymous substitution (dS), strongly suggesting the action of positive selection. Although the high dN/dS ratio is most evident outside ف the homeobox, the homeobox has a dN/dS of 0.89 and includes two codons that are likely under selection. Furthermore, the rate of radical amino acid substitutions that alter amino acid charge is significantly greater than that of conservative substitutions, suggesting that the selection promotes diversity of the protein charge profile. More interestingly, an analysis of 64 orthologous homeobox genes from humans and mice shows substantially higher rates of amino acid substitution in X-linked testis-expressed genes than in other genes. -

DNMT Inhibitors Increase Methylation at Subset of Cpgs in Colon
bioRxiv preprint doi: https://doi.org/10.1101/395467; this version posted August 25, 2018. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. 1 Title: DNMT inhibitors increase methylation at subset of CpGs in colon, bladder, lymphoma, 2 breast, and ovarian, cancer genome 3 Running title: Decitabine/azacytidine increases DNA methylation 4 Anil K Giri1, Tero Aittokallio1,2 5 1Institute for Molecular Medicine Finland, FIMM, University of Helsinki, Helsinki, Finland. 6 2Department of Mathematics and Statistics, University of Turku, Turku, Finland. 7 Correspondence to 8 Dr. Anil K Giri 9 Institute for Molecular Medicine Finland FIMM, University of Helsinki, Helsinki, Finland. 10 Email: [email protected] 11 Financial disclosure: This work was funded by the Academy of Finland (grants 269862, 292611, 12 310507 and 313267), Cancer Society of Finland, and the Sigrid Juselius Foundation. 13 Ethical disclosure: This study is an independent analysis of existing data available in the public 14 domain and does not involve any animal or human samples that have been collected by the authors 15 themselves. 16 Author contribution: AKG conceptualized, analyzed the data and wrote the manuscript. TA 17 critically revised and edited the manuscript. The authors report no conflict of interest. 18 19 Word count: 20 Figure number: 5 21 Table number: 1 22 23 Abstract 24 Background: DNA methyltransferase inhibitors (DNMTi) decitabine and azacytidine are approved 25 therapies for acute myeloid leukemia and myelodysplastic syndrome. -
Conserved Immunomodulatory Transcriptional Networks Underlie Antipsychotic-Induced Weight Gain
Translational Psychiatry www.nature.com/tp ARTICLE OPEN Conserved immunomodulatory transcriptional networks underlie antipsychotic-induced weight gain 1,3 1,3 1 1 2 2 Rizaldy C. Zapata✉ , Besma S. Chaudry , Mariela Lopez Valencia , Dinghong Zhang , Scott A. Ochsner , Neil J. McKenna and Olivia Osborn 1 © The Author(s) 2021 Although antipsychotics, such as olanzapine, are effective in the management of psychiatric conditions, some patients experience excessive antipsychotic-induced weight gain (AIWG). To illuminate pathways underlying AIWG, we compared baseline blood gene expression profiles in two cohorts of mice that were either prone (AIWG-P) or resistant (AIWG-R) to weight gain in response to olanzapine treatment for two weeks. We found that transcripts elevated in AIWG-P mice relative to AIWG-R are enriched for high- confidence transcriptional targets of numerous inflammatory and immunomodulatory signaling nodes. Moreover, these nodes are themselves enriched for genes whose disruption in mice is associated with reduced body fat mass and slow postnatal weight gain. In addition, we identified gene expression profiles in common between our mouse AIWG-P gene set and an existing human AIWG-P gene set whose regulation by immunomodulatory transcription factors is highly conserved between species. Finally, we identified striking convergence between mouse AIWG-P transcriptional regulatory networks and those associated with body weight and body mass index in humans. We propose that immunomodulatory transcriptional networks drive AIWG, and that these networks have broader conserved roles in whole body-metabolism. Translational Psychiatry (2021) 11:405 ; https://doi.org/10.1038/s41398-021-01528-y INTRODUCTION incompletely understood. Previous studies have shown that the Antipsychotic drugs are effective medications for the treatment of effect of olanzapine on AIWG can be effectively modeled in mice psychiatric disease but have significant side effects, including [15, 16]. -
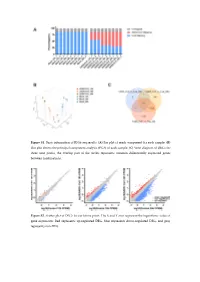
Figure S1. Basic Information of RNA-Seq Results. (A) Bar Plot of Reads Component for Each Sample
Figure S1. Basic information of RNA-seq results. (A) Bar plot of reads component for each sample. (B) Dot plot shows the principal component analysis (PCA) of each sample. (C) Venn diagram of DEGs for three time points, the overlap part of the circles represents common differentially expressed genes between combinations. Figure S2. Scatter plot of DEGs for each time point. The X and Y axes represent the logarithmic value of gene expression. Red represents up-regulated DEG, blue represents down-regulated DEG, and gray represents non-DEG. Table S1. Primers used for quantitative real-time PCR analysis of DEGs. Gene Primer Sequence Forward 5’-CTACGAGTGGATGGTCAAGAGC-3’ FOXO1 Reverse 5’-CCAGTTCCTTCATTCTGCACACG-3’ Forward 5’-GACGTCCGGCATCAGAGAAA-3’ IRS2 Reverse 5’-TCCACGGCTAATCGTCACAG-3’ Forward 5’-CACAACCAGGACCTCACACC-3’ IRS1 Reverse 5’-CTTGGCACGATAGAGAGCGT-3’ Forward 5’-AGGATACCACTCCCAACAGACCT-3’ IL6 Reverse 5’-CAAGTGCATCATCGTTGTTCATAC-3’ Forward 5’-TCACGTTGTACGCAGCTACC-3’ CCL5 Reverse 5’-CAGTCCTCTTACAGCCTTTGG-3’ Forward 5’-CTGTGCAGCCGCAGTGCCTACC-3’ BMP7 Reverse 5’-ATCCCTCCCCACCCCACCATCT-3’ Forward 5’-CTCTCCCCCTCGACTTCTGA-3’ BCL2 Reverse 5’-AGTCACGCGGAACACTTGAT-3’ Forward 5’-CTGTCGAACACAGTGGTACCTG-3’ FGF7 Reverse 5’-CCAACTGCCACTGTCCTGATTTC-3’ Forward 5’-GGGAGCCAAAAGGGTCATCA-3’ GAPDH Reverse 5’-CGTGGACTGTGGTCATGAGT-3’ Supplementary material: Differentially expressed genes log2(SADS-CoV_12h/ Qvalue (SADS-CoV _12h/ Gene Symbol Control_12h) Control_12h) PTGER4 -1.03693 6.79E-04 TMEM72 -3.08132 3.66E-04 IFIT2 -1.02918 2.11E-07 FRAT2 -1.09282 4.66E-05 -

Overview Gene List Target Scan Vs DIANA Group a Group B Group A
Overview Gene list Target scan vs DIANA Group A Group B Group A hsa-miR-181a hsa-miR-323 hsa-miR-326 Target scan Diana microT Overlap Target scan Diana microT Overlap Target scan SEPT3 SEPT3 SEPT3 SEPT7 ADARB1 HPCAL4 ABHD2 ABL2 ABHD13 ACVR2A ADCYAP1R1 AKAP13 PDPK1 ACRBP ACAN ABI1 ADAMTS1 ALAD APOBR ACVRL1 ACCN2 ABLIM1 ADAMTSL1 ANKRD52 ATXN1 ADAM19 ACER3 ACSL1 AKAP7 ARID2 C18orf23,RNF165 ADAM33 ACVR2A ACTN2 ANKRD43 ARL3 C20orf29 ADAMTS2 ADAMTS1 ACVR2A AP1S3 ARRB1 CACNG4 AHCYL2 ADAMTS18 ACVR2B ARID2 BBC3 CCNJL ALOX15B ADAMTS5 ADAM11 ATP11A BTG1 CYP2E1 ANK1 ADAMTSL1 ADAM22 ATXN1 C18orf62 GNB1L ANKS6 ADARB1 ADAMTS1 B4GALT1 C1orf21 GPR61 APBA1 AFAP1 ADAMTS6 BAG4 CADM4 GTSE1 ARCN1 AFTPH ADAMTSL1 BAI3 CALML4 HPCAL4 ARHGEF37 AK3 ADCY9 BNC2 CAPN6 KIAA0152 ARID3B AKAP7 ADRBK1 BRD1 CBFA2T2 KIF1A ARL8A ANAPC16 AFF2 BRWD1 CEBPA MACF1 ATP2B2 ANK1 AHCTF1,AHCTF1PBTBD3 CHD1 MYO1D ATP6V1G2 ANKRD12 AKAP2,PALM2 C13orf23 CIT PCNT AUP1 ANKRD33B AKAP6 C14orf43 CLASP2 PDPK1 BCL2L2 ANKRD43 AKAP7 CAPRIN1 CLCN5 PLEKHG4B BHLHE40 ANKRD44 AKAP9 CARM1 CLIP3 PPARA BTBD3 ANKRD52 AKT3 CBX4 COL5A2 PRB1,PRB2,PRB4 BTRC AP1S3 ALG9 CCDC117 CTNS PTPRT C10orf26 APBA1 ANKRD13C CCNJ DCTN4 PYCR1 C14orf1 APLP2 ANKRD20B CDH13 DCUN1D4 RAPGEF1 C16orf45 APOO ANKRD43 CDON DDB1 SRCAP C16orf54 ARID2 ANKRD50 CDYL DDX39B TMEM63C C1orf106 ARL3 AP1G1 CEP350 DIP2C C1orf27 ARRDC3 AP1S3 CHD7 DNAJB3 C22orf29 ATF7 API5 CHIC1 EEPD1 C9orf3 ATG2B ARFGEF2 CLIP1 EIF2C1 CACNA1E ATG7 ARHGAP12 CNOT6L ELFN2 CAPN12 ATP11A ARHGAP26 CNR1 ELK1 CASKIN1 ATP2B3 ARHGAP29 CNTN4 FAM172A CBFA2T3 ATP8B2 ARHGEF3 CNTNAP2 -

Six1 and Six4 Homeoproteins Are Required for Pax3 and Mrf
Research article Development and disease 2235 Six1 and Six4 homeoproteins are required for Pax3 and Mrf expression during myogenesis in the mouse embryo Raphaelle Grifone1, Josiane Demignon1,*, Christophe Houbron2,*, Evelyne Souil3, Claire Niro1, Mary J. Seller4, Ghislaine Hamard2 and Pascal Maire1,† 1Département Génétique, Développement et Pathologie Moléculaire, Institut Cochin – INSERM 567, CNRS UMR 8104, Université Paris V, 24 Rue du Faubourg Saint Jacques 75014 Paris, France 2Plateforme de recombinaison homologue, Institut Cochin – INSERM 567, CNRS UMR 8104, Université Paris V, 24 Rue du Faubourg Saint Jacques 75014 Paris, France 3Plateforme d’histologie, Institut Cochin – INSERM 567, CNRS UMR 8104, Université Paris V, 24 Rue du Faubourg Saint Jacques 75014 Paris, France 4Division of Medical and Molecular Genetics, Guy’s Hospital, London SE1 9RT, UK *These authors contributed equally to this work †Author for correspondence (e-mail: [email protected]) Accepted 9 February 2005 Development 132, 2235-2249 Published by The Company of Biologists 2005 doi:10.1242/dev.01773 Summary In mammals, Six5, Six4 and Six1 genes are co-expressed abolished, while it is preserved in Pax3-deficient embryos. during mouse myogenesis. Six4 and Six5 single knockout Within the myotome, absence of Six1 and Six4 impairs the (KO) mice have no developmental defects, while Six1 KO expression of the myogenic regulatory factors myogenin mice die at birth and show multiple organ developmental and Myod1, and Mrf4 expression becomes undetectable. defects. We have generated Six1Six4 double KO mice and Myf5 expression is correctly initiated but becomes show an aggravation of the phenotype previously reported restricted to the caudal region of each somite. -

Functionally Diverse Human T Cells Recognize Non-Microbial Antigens Presented By
RESEARCH ARTICLE Functionally diverse human T cells recognize non-microbial antigens presented by MR1 Marco Lepore1, Artem Kalinichenko1, Salvatore Calogero1, Pavanish Kumar2, Bhairav Paleja2, Mathias Schmaler1, Vipin Narang2, Francesca Zolezzi2, Michael Poidinger2,3, Lucia Mori1,2, Gennaro De Libero1,2* 1Department of Biomedicine, University Hospital and University of Basel, Basel, Switzerland; 2Singapore Immunology Network, A*STAR, Singapore, Singapore; 3Singapore Institute for Clinical Sciences, A*STAR, Singapore, Singapore Abstract MHC class I-related molecule MR1 presents riboflavin- and folate-related metabolites to mucosal-associated invariant T cells, but it is unknown whether MR1 can present alternative antigens to other T cell lineages. In healthy individuals we identified MR1-restricted T cells (named MR1T cells) displaying diverse TCRs and reacting to MR1-expressing cells in the absence of microbial ligands. Analysis of MR1T cell clones revealed specificity for distinct cell-derived antigens and alternative transcriptional strategies for metabolic programming, cell cycle control and functional polarization following antigen stimulation. Phenotypic and functional characterization of MR1T cell clones showed multiple chemokine receptor expression profiles and secretion of diverse effector molecules, suggesting functional heterogeneity. Accordingly, MR1T cells exhibited distinct T helper-like capacities upon MR1-dependent recognition of target cells expressing physiological levels of surface MR1. These data extend the role of -

BMC Biology Biomed Central
BMC Biology BioMed Central Research article Open Access Classification and nomenclature of all human homeobox genes PeterWHHolland*†1, H Anne F Booth†1 and Elspeth A Bruford2 Address: 1Department of Zoology, University of Oxford, South Parks Road, Oxford, OX1 3PS, UK and 2HUGO Gene Nomenclature Committee, European Bioinformatics Institute (EMBL-EBI), Wellcome Trust Genome Campus, Hinxton, Cambridgeshire, CB10 1SA, UK Email: Peter WH Holland* - [email protected]; H Anne F Booth - [email protected]; Elspeth A Bruford - [email protected] * Corresponding author †Equal contributors Published: 26 October 2007 Received: 30 March 2007 Accepted: 26 October 2007 BMC Biology 2007, 5:47 doi:10.1186/1741-7007-5-47 This article is available from: http://www.biomedcentral.com/1741-7007/5/47 © 2007 Holland et al; licensee BioMed Central Ltd. This is an Open Access article distributed under the terms of the Creative Commons Attribution License (http://creativecommons.org/licenses/by/2.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited. Abstract Background: The homeobox genes are a large and diverse group of genes, many of which play important roles in the embryonic development of animals. Increasingly, homeobox genes are being compared between genomes in an attempt to understand the evolution of animal development. Despite their importance, the full diversity of human homeobox genes has not previously been described. Results: We have identified all homeobox genes and pseudogenes in the euchromatic regions of the human genome, finding many unannotated, incorrectly annotated, unnamed, misnamed or misclassified genes and pseudogenes. -

Setd1 Histone 3 Lysine 4 Methyltransferase Complex Components in Epigenetic Regulation
SETD1 HISTONE 3 LYSINE 4 METHYLTRANSFERASE COMPLEX COMPONENTS IN EPIGENETIC REGULATION Patricia A. Pick-Franke Submitted to the faculty of the University Graduate School in partial fulfillment of the requirements for the degree Master of Science in the Department of Biochemistry and Molecular Biology Indiana University December 2010 Accepted by the Faculty of Indiana University, in partial fulfillment of the requirements for the degree of Master of Science. _____________________________________ David Skalnik, Ph.D., Chair _____________________________________ Kristin Chun, Ph.D. Master’s Thesis Committee _____________________________________ Simon Rhodes, Ph.D. ii DEDICATION This thesis is dedicated to my sons, Zachary and Zephaniah who give me great joy, hope and continuous inspiration. I can only hope that I successfully set a good example demonstrating that one can truly accomplish anything, if you never give up and reach for your dreams. iii ACKNOWLEDGEMENTS I would like to thank my committee members Dr. Skalnik, Dr. Chun and Dr. Rhodes for allowing me to complete this dissertation. They have been incredibly generous with their flexibility. I must make a special thank you to Jeanette McClintock, who willingly gave her expertise in statistical analysis with the Cfp1 microarray data along with encouragement, support and guidance to complete this work. I would like to thank Courtney Tate for her ceaseless willingness to share ideas, and her methods and materials, and Erika Dolbrota for her generous instruction as well as the name of a good doctor. I would also like to acknowledge the superb mentorship of Dr. Jeon Heong Lee, PhD and the contagious passion and excitement for the life of science of Dr.