Cryo-EM Structure of the Receptor-Activated TRPC5 Ion Channel
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

The Structural Basis for an On–Off Switch Controlling Gβγ-Mediated Inhibition of TRPM3 Channels
The structural basis for an on–off switch controlling Gβγ-mediated inhibition of TRPM3 channels Marc Behrendta,b,1,2, Fabian Grussc,1,3, Raissa Enzerotha,b, Sandeep Demblaa,b, Siyuan Zhaod, Pierre-Antoine Crassousd, Florian Mohra, Mieke Nysc, Nikolaos Lourose, Rodrigo Gallardoe,4, Valentina Zorzinif,5, Doris Wagnera, Anastassios Economou (Αναστάσιoς Οικoνόμoυ)f, Frederic Rousseaue, Joost Schymkowitze, Stephan E. Philippg, Tibor Rohacsd, Chris Ulensc,6, and Johannes Oberwinklera,b,6 aInstitut für Physiologie und Pathophysiologie, Philipps-Universität Marburg, 35037 Marburg, Germany;bCenter for Mind, Brain and Behavior (CMBB), Philipps-Universität Marburg and Justus-Liebig-Universität Giessen, 35032 Marburg, Germany; cLaboratory of Structural Neurobiology, Department of Cellular and Molecular Medicine, KU Leuven, 3000 Leuven, Belgium; dDepartment of Pharmacology, Physiology and Neuroscience, Rutgers New Jersey Medical School, Newark, NJ 07103; eSwitch Laboratory, VIB Center for Brain and Disease Research, Department of Cellular and Molecular Medicine, KU Leuven, 3000 Leuven, Belgium; fLaboratory of Molecular Bacteriology, Department of Microbiology and Immunology, Rega Institute for Medical Research, KU Leuven, 3000 Leuven, Belgium; and gExperimentelle und Klinische Pharmakologie und Toxikologie, Universität des Saarlandes, 66421 Homburg, Germany Edited by László Csanády, Semmelweis University, Budapest, Hungary, and accepted by Editorial Board Member David E. Clapham August 27, 2020 (received for review February 13, 2020) TRPM3 channels play important roles in the detection of noxious also subject to inhibition by activated μORsorotherGPCRs(6, heat and in inflammatory thermal hyperalgesia. The activity of 24–28), a process that has been shown to take place in peripheral these ion channels in somatosensory neurons is tightly regulated by endings of nociceptive neurons since locally applied μOR or μ βγ -opioid receptors through the signaling of G proteins, thereby GABA receptor agonists inhibit TRPM3-dependent pain (24–26). -

The TRPP2-Dependent Channel of Renal Primary Cilia Also Requires TRPM3
The TRPP2-dependent channel of renal primary cilia also requires TRPM3 Steven J. Kleene1, Brian J. Siroky2, Julio A. Landero-Figueroa3, Bradley P. Dixon4, Nolan W. Pachciarz2, Lu Lu2, and Nancy K. Kleene1 1Department of Pharmacology and Systems Physiology, University of Cincinnati, Cincinnati, Ohio; 2Division of Nephrology and Hypertension, Cincinnati Children's Hospital Medical Center, Cincinnati, Ohio; 3Department of Chemistry, University of Cincinnati, Cincinnati, Ohio; 4Renal Section, Department of Pediatrics, University of Colorado School of Medicine, Aurora, Colorado IntroductIon Primary cilia of renal epithelial cells express several members of the transient receptor potential (TRP) class of cation-conducting channel, including TRPC1, TRPM3, TRPM4, TRPP2, and TRPV4. Some cases of autosomal dominant polycystic kidney disease (ADPKD) are caused by defects in TRPP2 (also called polycystin-2, PC2, or PKD2). A large-conductance, TRPP2- dependent channel in renal cilia has been well described, but it is not known whether this channel includes any other protein subunits. Methods To study this question, we investigated the pharmacology of the TRPP2-dependent channel through electrical recordings from the cilia of mIMCD-3 cells, a murine cell line of renal epithelial origin, results The pharmacology was found to match that of TRPM3 channels. The ciliary TRPP2-dependent channel is known to be activated by depolarization and/or increasing cytoplasmic Ca2+. This activation was greatly enhanced by external pregnenolone sulfate, an agonist of TRPM3 channels. Pregnenolone sulfate did not change the current-voltage relation of the channel. CIM0216, another TRPM3 agonist, modestly increased the activity of the ciliary channels. The channels were effectively blocked by isosakuranetin, a specific inhibitor of TRPM3 channels. -

TRPC5 Regulates Axonal Outgrowth in Developing Retinal Ganglion Cells
Laboratory Investigation (2020) 100:297–310 https://doi.org/10.1038/s41374-019-0347-1 ARTICLE TRPC5 regulates axonal outgrowth in developing retinal ganglion cells 1 1 2 1 1 Mai Oda ● Hanako Yamamoto ● Hidetaka Matsumoto ● Yasuki Ishizaki ● Koji Shibasaki Received: 22 July 2019 / Revised: 15 November 2019 / Accepted: 15 November 2019 / Published online: 16 December 2019 © The Author(s), under exclusive licence to United States and Canadian Academy of Pathology 2019 Abstract The TRPC5 ion channel is activated upon depletion of intracellular calcium stores, as well as by various stimuli such as nitric oxide (NO), membrane stretch, and cold temperatures. TRPC5 is abundantly expressed in the central nervous system where it has important neuronal functions. In the chick retina, TRPC5 expression was shown to be restricted to amacrine cells (ACs) and Müller glial cells, although its expression was also observed in the ganglion cell layer (GCL) in displaced ACs, as determined by their characteristic cell morphology. However, it is possible that this expression analysis alone might be insufficient to fully understand the expression of TRPC5 in retinal ganglion cells (RGCs). Hence, we analyzed TRPC5 expression by in situ hybridization and immunostaining in the developing mouse retina, and for the first time identified that 1234567890();,: 1234567890();,: developing and mature RGCs strongly express TRPC5. The expression begins at E14.5, and is restricted to ACs and RGCs. It was reported that TRPC5 negatively regulates axonal outgrowth in hippocampal neurons. We thus hypothesized that TRPC5 might have similar functions in RGCs since they extend very long axons toward the brain, and this characteristic significantly differs from other retinal cell types. -

TRPC1 Regulates the Activity of a Voltage-Dependent Nonselective Cation Current in Hippocampal CA1 Neurons
cells Article TRPC1 Regulates the Activity of a Voltage-Dependent Nonselective Cation Current in Hippocampal CA1 Neurons 1, 1 1,2 1,3, Frauke Kepura y, Eva Braun , Alexander Dietrich and Tim D. Plant * 1 Pharmakologisches Institut, BPC-Marburg, Fachbereich Medizin, Philipps-Universität Marburg, Karl-von-Frisch-Straße 2, 35043 Marburg, Germany; [email protected] (F.K.); braune@staff.uni-marburg.de (E.B.); [email protected] (A.D.) 2 Walther-Straub-Institut für Pharmakologie und Toxikologie, Ludwig-Maximilians-Universität München, 80336 München, Germany 3 Center for Mind, Brain and Behavior, Philipps-Universität Marburg, 35032 Marburg, Germany * Correspondence: plant@staff.uni-marburg.de; Tel.: +49-6421-28-65038 Present address: Institut für Bodenkunde und Pflanzenernährung/Institut für angewandte Ökologie, y Hochschule Geisenheim University, Von-Lade-Str. 1, 65366 Geisenheim, Germany. Received: 27 November 2019; Accepted: 14 February 2020; Published: 18 February 2020 Abstract: The cation channel subunit TRPC1 is strongly expressed in central neurons including neurons in the CA1 region of the hippocampus where it forms complexes with TRPC4 and TRPC5. To investigate the functional role of TRPC1 in these neurons and in channel function, we compared current responses to group I metabotropic glutamate receptor (mGluR I) activation and looked +/+ / for major differences in dendritic morphology in neurons from TRPC1 and TRPC1− − mice. mGluR I stimulation resulted in the activation of a voltage-dependent nonselective cation current in both genotypes. Deletion of TRPC1 resulted in a modification of the shape of the current-voltage relationship, leading to an inward current increase. In current clamp recordings, the percentage of neurons that responded to depolarization in the presence of an mGluR I agonist with a plateau / potential was increased in TRPC1− − mice. -

The TRPM4 Channel Inhibitor 9-Phenanthrol
W&M ScholarWorks Arts & Sciences Articles Arts and Sciences 2014 The TRPM4 channel inhibitor 9-phenanthrol R. Guinamard T. Hof C. A. Del Negro College of William and Mary Follow this and additional works at: https://scholarworks.wm.edu/aspubs Recommended Citation Guinamard, R., Hof, T., & Del Negro, C. A. (2014). The TRPM4 channel inhibitor 9‐phenanthrol. British Journal of Pharmacology, 171(7), 1600-1613. This Article is brought to you for free and open access by the Arts and Sciences at W&M ScholarWorks. It has been accepted for inclusion in Arts & Sciences Articles by an authorized administrator of W&M ScholarWorks. For more information, please contact [email protected]. British Journal of DOI:10.1111/bph.12582 www.brjpharmacol.org BJP Pharmacology REVIEW Correspondence Romain Guinamard, Groupe Signalisation, Electrophysiologie et Imagerie des Lésions The TRPM4 channel d’Ischémie/Reperfusion Myocardique, EA4650, Université de Caen Basse Normandie, inhibitor 9-phenanthrol Sciences D, Esplande de la Paix, CS 14032, 14032 Caen Cedex 5, R Guinamard1,2, T Hof1 and C A Del Negro2 France. E-mail: [email protected] ---------------------------------------------------------------- 1EA 4650, Groupe Signalisation, Electrophysiologie et Imagerie des Lésions d’Ischémie-Reperfusion Myocardique, UCBN, Normandie Université, Caen, France, 2Department Keywords 9-phenanthrol; TRPM4; of Applied Science, The College of William and Mary, Williamsburg, VA, USA calcium-activated non-selective cation channel; cardioprotection; NSCCa ---------------------------------------------------------------- Received 1 November 2013 Revised 17 December 2013 Accepted 8 January 2014 The phenanthrene-derivative 9-phenanthrol is a recently identified inhibitor of the transient receptor potential melastatin (TRPM) 4 channel, a Ca2+-activated non-selective cation channel whose mechanism of action remains to be determined. -
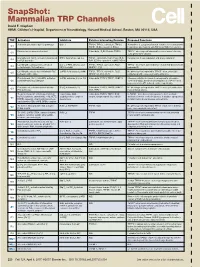
Snapshot: Mammalian TRP Channels David E
SnapShot: Mammalian TRP Channels David E. Clapham HHMI, Children’s Hospital, Department of Neurobiology, Harvard Medical School, Boston, MA 02115, USA TRP Activators Inhibitors Putative Interacting Proteins Proposed Functions Activation potentiated by PLC pathways Gd, La TRPC4, TRPC5, calmodulin, TRPC3, Homodimer is a purported stretch-sensitive ion channel; form C1 TRPP1, IP3Rs, caveolin-1, PMCA heteromeric ion channels with TRPC4 or TRPC5 in neurons -/- Pheromone receptor mechanism? Calmodulin, IP3R3, Enkurin, TRPC6 TRPC2 mice respond abnormally to urine-based olfactory C2 cues; pheromone sensing 2+ Diacylglycerol, [Ca ]I, activation potentiated BTP2, flufenamate, Gd, La TRPC1, calmodulin, PLCβ, PLCγ, IP3R, Potential role in vasoregulation and airway regulation C3 by PLC pathways RyR, SERCA, caveolin-1, αSNAP, NCX1 La (100 µM), calmidazolium, activation [Ca2+] , 2-APB, niflumic acid, TRPC1, TRPC5, calmodulin, PLCβ, TRPC4-/- mice have abnormalities in endothelial-based vessel C4 i potentiated by PLC pathways DIDS, La (mM) NHERF1, IP3R permeability La (100 µM), activation potentiated by PLC 2-APB, flufenamate, La (mM) TRPC1, TRPC4, calmodulin, PLCβ, No phenotype yet reported in TRPC5-/- mice; potentially C5 pathways, nitric oxide NHERF1/2, ZO-1, IP3R regulates growth cones and neurite extension 2+ Diacylglycerol, [Ca ]I, 20-HETE, activation 2-APB, amiloride, Cd, La, Gd Calmodulin, TRPC3, TRPC7, FKBP12 Missense mutation in human focal segmental glomerulo- C6 potentiated by PLC pathways sclerosis (FSGS); abnormal vasoregulation in TRPC6-/- -

Involvement of TRPC4 and 5 Channels in Persistent Firing in Hippocampal CA1 Pyramidal Cells
cells Article Involvement of TRPC4 and 5 Channels in Persistent Firing in Hippocampal CA1 Pyramidal Cells Alberto Arboit 1,2,3, Antonio Reboreda 1,4 and Motoharu Yoshida 1,3,4,5,* 1 German Center for Neurodegenerative Diseases (DZNE), 39120 Magdeburg, Germany; [email protected] (A.A.); [email protected] (A.R.) 2 Otto-von-Guericke University, 39120 Magdeburg, Germany 3 Faculty of Psychology, Ruhr University Bochum (RUB), Universitätsstraße 150, 44801 Bochum, Germany 4 Leibniz Institute for Neurobiology (LIN), 39118 Magdeburg, Germany 5 Center for Behavioral Brain Sciences (CBBS), 39106 Magdeburg, Germany * Correspondence: [email protected] Received: 1 December 2019; Accepted: 1 February 2020; Published: 5 February 2020 Abstract: Persistent neural activity has been observed in vivo during working memory tasks, and supports short-term (up to tens of seconds) retention of information. While synaptic and intrinsic cellular mechanisms of persistent firing have been proposed, underlying cellular mechanisms are not yet fully understood. In vitro experiments have shown that individual neurons in the hippocampus and other working memory related areas support persistent firing through intrinsic cellular mechanisms that involve the transient receptor potential canonical (TRPC) channels. Recent behavioral studies demonstrating the involvement of TRPC channels on working memory make the hypothesis that TRPC driven persistent firing supports working memory a very attractive one. However, this view has been challenged by recent findings that persistent firing in vitro is unchanged in TRPC knock out (KO) mice. To assess the involvement of TRPC channels further, we tested novel and highly specific TRPC channel blockers in cholinergically induced persistent firing in mice CA1 pyramidal cells for the first time. -

FKBP52 Regulates TRPC3-Dependent Ca<Sup>
© 2019. Published by The Company of Biologists Ltd | Journal of Cell Science (2019) 132, jcs231506. doi:10.1242/jcs.231506 RESEARCH ARTICLE FKBP52 regulates TRPC3-dependent Ca2+ signals and the hypertrophic growth of cardiomyocyte cultures Sandra Bandleon1, Patrick P. Strunz1, Simone Pickel2, Oleksandra Tiapko3, Antonella Cellini1, Erick Miranda-Laferte2 and Petra Eder-Negrin1,* ABSTRACT A single TRPC subunit is composed of six transmembrane The transient receptor potential (TRP; C-classical, TRPC) channel domains with a pore-forming loop connecting the transmembrane ‘ ’ TRPC3 allows a cation (Na+/Ca2+) influx that is favored by the domains 5 and 6, a preserved 25 amino acid sequence called a TRP domain and two cytosolic domains, an N-terminal ankyrin repeat stimulation of Gq protein-coupled receptors (GPCRs). An enhanced TRPC3 activity is related to adverse effects, including pathological domain and a C-terminal coiled-coil domain (Eder et al., 2007; Fan hypertrophy in chronic cardiac disease states. In the present study, et al., 2018). The cytosolic domains mediate ion channel formation we identified FK506-binding protein 52 (FKBP52, also known as and are implicated in ion channel regulation and plasma membrane FKBP4) as a novel interaction partner of TRPC3 in the heart. FKBP52 targeting (Eder et al., 2007). Among several protein interaction sites, was recovered from a cardiac cDNA library by a C-terminal TRPC3 the C-terminus of all TRPC subunits harbors a highly conserved fragment (amino acids 742–848) in a yeast two-hybrid screen. proline-rich sequence that corresponds to the binding domain in the Drosophila Downregulation of FKBP52 promoted a TRPC3-dependent photoreceptor channel TRPL for the FK506-binding hypertrophic response in neonatal rat cardiomyocytes (NRCs). -

TRPC5 Does Not Cause Or Aggravate Glomerular Disease
BRIEF COMMUNICATION www.jasn.org TRPC5 Does Not Cause or Aggravate Glomerular Disease Xuexiang Wang , Ranadheer R. Dande, Hao Yu, Beata Samelko, Rachel E. Miller, Mehmet M. Altintas, and Jochen Reiser Department of Medicine, Rush University Medical Center, Chicago, Illinois ABSTRACT Transient receptor potential channel 5 (TRPC5) is highly expressed in brain and or pharmacologic inhibition of TRPC5 pro- kidney and mediates calcium influx and promotes cell migration. In the kidney, loss tected mice from albuminuria. Yet, the di- of TRPC5 function has been reported to benefit kidney filter dynamics by balancing rect effect of TRPC5 overexpression in mice podocyte cytoskeletal remodeling. However, in vivo gain-in-function studies of was not shown and a direct pathogenic TRPC5 with respect to kidney function have not been reported. To address this role of TRPC5 for proteinuria remained gap, we developed two transgenic mouse models on the C57BL/6 background by obscure. overexpressing either wild-type TRPC5 or a TRPC5 ion-pore mutant. Compared with We generated two novel transgenic nontransgenic controls, neither transgenic model exhibited an increase in protein- mouse models by overexpression of ei- uria at 8 months of age or a difference in LPS-induced albuminuria. Moreover, acti- ther wild-type TRPC5 (TG) or pore mu- vation of TRPC5 by Englerin A did not stimulate proteinuria, and inhibition of TRPC5 tant dominant negative TRPC5 (DN) in by ML204 did not significantly lower the level of LPS-induced proteinuria in any mice on a C57BL/6 background (B/6). group. Collectively, these data suggest that the overexpression or activation of With these unique models, we have the the TRPC5 ion channel does not cause kidney barrier injury or aggravate such injury opportunity to investigate the functional under pathologic conditions. -

Structure of Full-Length Human TRPM4
Structure of full-length human TRPM4 Jingjing Duana,1, Zongli Lib,c,1, Jian Lid,e,1, Ana Santa-Cruza, Silvia Sanchez-Martineza, Jin Zhangd,2, and David E. Claphama,f,g,2 aHoward Hughes Medical Institute, Ashburn, VA 20147; bHoward Hughes Medical Institute, Harvard Medical School, Boston, MA 02115; cDepartment of Biological Chemistry and Molecular Pharmacology, Harvard Medical School, Boston, MA 02115; dSchool of Basic Medical Sciences, Nanchang University, Nanchang, 330031 Jiangxi, China; eDepartment of Molecular and Cellular Biochemistry, University of Kentucky, Lexington, KY 40536; fDepartment of Neurobiology, Harvard Medical School, Boston, MA 02115; and gDepartment of Cardiology, Boston Children’s Hospital, Boston, MA 02115 Contributed by David E. Clapham, January 25, 2018 (sent for review December 19, 2017; reviewed by Mark T. Nelson, Dejian Ren, and Thomas Voets) Transient receptor potential melastatin subfamily member 4 Structure of Human TRPM4 (TRPM4) is a widely distributed, calcium-activated, monovalent- Full-length human TRPM4 (1,214 residues) was expressed using selective cation channel. Mutations in human TRPM4 (hTRPM4) the BacMam expression system (Materials and Methods). The result in progressive familial heart block. Here, we report the + TRPM4 construct used for expression retained the key func- electron cryomicroscopy structure of hTRPM4 in a closed, Na - tional properties of the wild-type channel, such as permeability + + bound, apo state at pH 7.5 to an overall resolution of 3.7 Å. Five to Na and K and activation by intracellular calcium (Fig. S1A). partially hydrated sodium ions are proposed to occupy the center The protein, obtained in the absence of exogenous calcium ions, of the conduction pore and the entrance to the coiled-coil domain. -

Distribution Profiles of Transient Receptor Potential Melastatin-Related and Vanilloid-Related Channels in Prostatic Tissue in Rat
TRPM and TRPV in rat prostate DOI: 10.1111/j.1745-7262.2007.00291.x www.asiaandro.com .Original Article . Distribution profiles of transient receptor potential melastatin-related and vanilloid-related channels in prostatic tissue in rat Huai-Peng Wang*, Xiao-Yong Pu*, Xing-Huan Wang Department of Urology, Guangdong Provnicial People’s Hospital, Guangzhou 510080, China Abstract Aim: To investigate the expression and distribution of the members of the transient receptor potential (TRP) channel members of TRP melastatin (TRPM) and TRP vanilloid (TRPV) subfamilies in rat prostatic tissue. Methods: Pros- tate tissue was obtained from male Sprague-Dawley rats. Reverse transcription polymerase chain reaction (RT-PCR) and quantitative real-time polymerase chain reaction (PCR) were used to check the expression of all TRPM and TRPV channel members with specific primers. Immunohistochemistry staining for TRPM8 and TRPV1 were also per- formed in rat tissues. Results: TRPM2, TRPM3, TRPM4, TRPM6, TRPM7, TRPM8, TRPV2 and TRPV4 mRNA were detected in all rat prostatic tissues. Very weak signals for TRPM1, TRPV1 and TRPV3 were also detected. The mRNA of TRPM5, TRPV5 and TRPV6 were not detected in all RT-PCR experiments. Quantitative real-time RT-PCR showed that TRPM2, TRPM3, TRPM4, TRPM8, TRPV2 and TRPV4 were the most abundantly expressed TRPM and TRPV subtypes, respectively. Fluorescence immunohistochemistry indicated that TRPM8 and TRPV1 are highly expressed in both epithelial and smooth muscle cells. Conclusion: Our results demonstrate that mRNA or protein for TRPM1, TRPM2, TRPM3, TRPM4, TRPM6, TRPM7, TRPM8, TRPV1, TRPV2, TRPV3 and TRPV4 exist in rat prostatic tissue. The data presented here assists in elucidating the physiological function of TRPM and TRPV channels. -

Transient Receptor Potential Cation Channel, Subfamily C, Member 5 (TRPC5) Is a Cold-Transducer in the Peripheral Nervous System
Transient receptor potential cation channel, subfamily C, member 5 (TRPC5) is a cold-transducer in the peripheral nervous system Katharina Zimmermanna,b,1,2, Jochen K. Lennerza,c,d,1,3, Alexander Heina,1,4, Andrea S. Linke, J. Stefan Kaczmareka,b, Markus Dellinga, Serdar Uysala, John D. Pfeiferc, Antonio Riccioa, and David E. Claphama,b,f,5 Departments of aCardiology, Manton Center for Orphan Disease, and bNeurobiology, Harvard Medical School, Boston, MA 02115; cDepartment of Pathology and Immunology, Washington University, St. Louis, MO, 63110; dDepartment of Pathology, Massachusetts General Hospital/Harvard Medical School, Boston, MA, 02116; eDepartment of Physiology and Pathophysiology, University Erlangen, 91054 Erlangen, Germany; and fHoward Hughes Medical Institute, Harvard Medical School, Children’s Hospital Boston, Boston, MA 02115 Contributed by David E. Clapham, September 20, 2011 (sent for review August 9, 2011) Detection and adaptation to cold temperature is crucial to survival. affected perforce by temperature, even if driven solely by con- Cold sensing in the innocuous range of cold (>10–15 °C) in the ventional NaV and KV channels—but high Q10 channels are likely mammalian peripheral nervous system is thought to rely primarily suspects in encoding temperature changes. Although TRPM8 on transient receptor potential (TRP) ion channels, most notably (a menthol receptor) is generally considered the primary cold – the menthol receptor, TRPM8. Here we report that TRP cation sensor for innocuous cold (11 13), and TRPA1 participates in channel, subfamily C member 5 (TRPC5), but not TRPC1/TRPC5 het- noxious cold sensing (9, 10) (14), many cold-sensitive neurons eromeric channels, are highly cold sensitive in the temperature lack TRPM8 as well as TRPA1 (15).