The Role of TTP Phosphorylation in the Regulation of Inflammatory Cytokine Production by MK2/3
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Tristetraprolin Limits Inflammatory Cytokine Production in Tumor-Associated Macrophages in an Mrna Decay− Independent Manner
Published OnlineFirst July 16, 2015; DOI: 10.1158/0008-5472.CAN-15-0205 Cancer Microenvironment and Immunology Research Tristetraprolin Limits Inflammatory Cytokine Production in Tumor-Associated Macrophages in an mRNA Decay–Independent Manner Franz Kratochvill1,2, Nina Gratz1, Joseph E. Qualls1,2, Lee-Ann Van De Velde1,2, Hongbo Chi2, Pavel Kovarik3, and Peter J. Murray1,2 Abstract Tristetraprolin (TTP) is an inducible zinc finger AU-rich (TAM). However, TTP's effects on AU-rich mRNA stability RNA-binding protein essential for enforcing degradation of were negligible and limited by constitutive p38a MAPK activ- mRNAs encoding inflammatory chemokines and cytokines. ity, which was the main driver of proinflammatory cytokine Most studies on TTP center on the connection between mRNA production in TAMs at the posttranscriptional level. Instead, half-life and inflammatory output, because loss of TTP ampli- elimination of TTP caused excessive protein production of fies inflammation by increasing the stability of AU-rich inflammatory mediators, suggesting TTP-dependent transla- mRNAs. Here, we focused on how TTP controls cytokine and tional suppression of AU-rich mRNAs. Manipulation of the chemokine production in the nonresolving inflammation of p38a–TTP axis in macrophages has significant effects on the cancer using tissue-specific approaches. In contrast with mod- growth of tumors and therefore represents a means to mani- el in vitro macrophage systems, we found constitutive TTP pulate inflammation in the tumor microenvironment. Cancer expression in late-stage tumor-associated macrophages Res; 75(15); 1–11. Ó2015 AACR. Introduction TLR signaling phosphorylates TTP thereby blocking its function and sustaining TNF output (9, 10). -

Table S1 the Four Gene Sets Derived from Gene Expression Profiles of Escs and Differentiated Cells
Table S1 The four gene sets derived from gene expression profiles of ESCs and differentiated cells Uniform High Uniform Low ES Up ES Down EntrezID GeneSymbol EntrezID GeneSymbol EntrezID GeneSymbol EntrezID GeneSymbol 269261 Rpl12 11354 Abpa 68239 Krt42 15132 Hbb-bh1 67891 Rpl4 11537 Cfd 26380 Esrrb 15126 Hba-x 55949 Eef1b2 11698 Ambn 73703 Dppa2 15111 Hand2 18148 Npm1 11730 Ang3 67374 Jam2 65255 Asb4 67427 Rps20 11731 Ang2 22702 Zfp42 17292 Mesp1 15481 Hspa8 11807 Apoa2 58865 Tdh 19737 Rgs5 100041686 LOC100041686 11814 Apoc3 26388 Ifi202b 225518 Prdm6 11983 Atpif1 11945 Atp4b 11614 Nr0b1 20378 Frzb 19241 Tmsb4x 12007 Azgp1 76815 Calcoco2 12767 Cxcr4 20116 Rps8 12044 Bcl2a1a 219132 D14Ertd668e 103889 Hoxb2 20103 Rps5 12047 Bcl2a1d 381411 Gm1967 17701 Msx1 14694 Gnb2l1 12049 Bcl2l10 20899 Stra8 23796 Aplnr 19941 Rpl26 12096 Bglap1 78625 1700061G19Rik 12627 Cfc1 12070 Ngfrap1 12097 Bglap2 21816 Tgm1 12622 Cer1 19989 Rpl7 12267 C3ar1 67405 Nts 21385 Tbx2 19896 Rpl10a 12279 C9 435337 EG435337 56720 Tdo2 20044 Rps14 12391 Cav3 545913 Zscan4d 16869 Lhx1 19175 Psmb6 12409 Cbr2 244448 Triml1 22253 Unc5c 22627 Ywhae 12477 Ctla4 69134 2200001I15Rik 14174 Fgf3 19951 Rpl32 12523 Cd84 66065 Hsd17b14 16542 Kdr 66152 1110020P15Rik 12524 Cd86 81879 Tcfcp2l1 15122 Hba-a1 66489 Rpl35 12640 Cga 17907 Mylpf 15414 Hoxb6 15519 Hsp90aa1 12642 Ch25h 26424 Nr5a2 210530 Leprel1 66483 Rpl36al 12655 Chi3l3 83560 Tex14 12338 Capn6 27370 Rps26 12796 Camp 17450 Morc1 20671 Sox17 66576 Uqcrh 12869 Cox8b 79455 Pdcl2 20613 Snai1 22154 Tubb5 12959 Cryba4 231821 Centa1 17897 -

A Computational Approach for Defining a Signature of Β-Cell Golgi Stress in Diabetes Mellitus
Page 1 of 781 Diabetes A Computational Approach for Defining a Signature of β-Cell Golgi Stress in Diabetes Mellitus Robert N. Bone1,6,7, Olufunmilola Oyebamiji2, Sayali Talware2, Sharmila Selvaraj2, Preethi Krishnan3,6, Farooq Syed1,6,7, Huanmei Wu2, Carmella Evans-Molina 1,3,4,5,6,7,8* Departments of 1Pediatrics, 3Medicine, 4Anatomy, Cell Biology & Physiology, 5Biochemistry & Molecular Biology, the 6Center for Diabetes & Metabolic Diseases, and the 7Herman B. Wells Center for Pediatric Research, Indiana University School of Medicine, Indianapolis, IN 46202; 2Department of BioHealth Informatics, Indiana University-Purdue University Indianapolis, Indianapolis, IN, 46202; 8Roudebush VA Medical Center, Indianapolis, IN 46202. *Corresponding Author(s): Carmella Evans-Molina, MD, PhD ([email protected]) Indiana University School of Medicine, 635 Barnhill Drive, MS 2031A, Indianapolis, IN 46202, Telephone: (317) 274-4145, Fax (317) 274-4107 Running Title: Golgi Stress Response in Diabetes Word Count: 4358 Number of Figures: 6 Keywords: Golgi apparatus stress, Islets, β cell, Type 1 diabetes, Type 2 diabetes 1 Diabetes Publish Ahead of Print, published online August 20, 2020 Diabetes Page 2 of 781 ABSTRACT The Golgi apparatus (GA) is an important site of insulin processing and granule maturation, but whether GA organelle dysfunction and GA stress are present in the diabetic β-cell has not been tested. We utilized an informatics-based approach to develop a transcriptional signature of β-cell GA stress using existing RNA sequencing and microarray datasets generated using human islets from donors with diabetes and islets where type 1(T1D) and type 2 diabetes (T2D) had been modeled ex vivo. To narrow our results to GA-specific genes, we applied a filter set of 1,030 genes accepted as GA associated. -

Human RIPK1 Deficiency Causes Combined Immunodeficiency and Inflammatory Bowel Diseases
Human RIPK1 deficiency causes combined immunodeficiency and inflammatory bowel diseases Yue Lia,1, Marita Führerb,1, Ehsan Bahramia,1, Piotr Sochac, Maja Klaudel-Dreszlerc, Amira Bouzidia, Yanshan Liua, Anna S. Lehlea, Thomas Magga, Sebastian Hollizecka, Meino Rohlfsa, Raffaele Concaa, Michael Fieldd, Neil Warnere,f, Slae Mordechaig, Eyal Shteyerh, Dan Turnerh,i, Rachida Boukarij, Reda Belbouabj, Christoph Walzk, Moritz M. Gaidtl,m, Veit Hornungl,m, Bernd Baumannn, Ulrich Pannickeb, Eman Al Idrissio, Hamza Ali Alghamdio, Fernando E. Sepulvedap,q, Marine Gilp,q, Geneviève de Saint Basilep,q,r, Manfred Hönigs, Sibylle Koletzkoa,i, Aleixo M. Muisee,f,i,t,u, Scott B. Snapperd,i,v,w, Klaus Schwarzb,x,2, Christoph Kleina,i,2, and Daniel Kotlarza,i,2,3 aDr. von Hauner Children’s Hospital, Department of Pediatrics, University Hospital, Ludwig-Maximilians-Universität (LMU) Munich, 80337 Munich, Germany; bThe Institute for Transfusion Medicine, University of Ulm, 89081 Ulm, Germany; cDepartment of Gastroenterology, Hepatology, Nutritional Disorders and Pediatrics, The Children’s Memorial Health Institute, 04730 Warsaw, Poland; dDivision of Gastroenterology, Hepatology and Nutrition, Boston Children’s Hospital, Boston, MA 02115; eSickKids Inflammatory Bowel Disease Center, Research Institute, Hospital for Sick Children, Toronto, ON M5G1X8, Canada; fCell Biology Program, Research Institute, Hospital for Sick Children, Toronto, ON M5G1X8, Canada; gPediatric Gastroenterology, Hadassah University Hospital, Jerusalem 91120, Israel; hThe Juliet Keidan -

Westminsterresearch ZFP36 Proteins and Mrna Targets in B Cell
WestminsterResearch http://www.westminster.ac.uk/westminsterresearch ZFP36 proteins and mRNA targets in B cell malignancies Alcaraz, A. This is an electronic version of a PhD thesis awarded by the University of Westminster. © Miss Amor Alcaraz, 2015. The WestminsterResearch online digital archive at the University of Westminster aims to make the research output of the University available to a wider audience. Copyright and Moral Rights remain with the authors and/or copyright owners. Whilst further distribution of specific materials from within this archive is forbidden, you may freely distribute the URL of WestminsterResearch: ((http://westminsterresearch.wmin.ac.uk/). In case of abuse or copyright appearing without permission e-mail [email protected] ZFP36 proteins and mRNA targets in B cell malignancies Maria del Amor Alcaraz-Serrano A Thesis submitted in partial fulfilment of the requirements of the University of Westminster for the degree of Doctor of Philosophy September 2015 Abstract The ZFP36 proteins are a family of post-transcriptional regulator proteins that bind to adenine uridine rich elements (AREs) in 3’ untranslated (3’UTR) regions of mRNAs. The members of the human family, ZFP36L1, ZFP36L2 and ZFP36 are able to degrade mRNAs of important cell regulators that include cytokines, cell signalling proteins and transcriptional factors. This project investigated two proposed targets for the protein family that have important roles in B cell biology, BCL2 and CD38 mRNAs. BCL2 is an anti-apoptotic protein with key roles in cell survival and carcinogenesis; CD38 is a membrane protein differentially expressed in B cells and with a prognostic value in B chronic lymphocytic leukaemia (B-CLL), patients positive for CD38 are considered to have a poor prognosis. -

Supplementary Table S4. FGA Co-Expressed Gene List in LUAD
Supplementary Table S4. FGA co-expressed gene list in LUAD tumors Symbol R Locus Description FGG 0.919 4q28 fibrinogen gamma chain FGL1 0.635 8p22 fibrinogen-like 1 SLC7A2 0.536 8p22 solute carrier family 7 (cationic amino acid transporter, y+ system), member 2 DUSP4 0.521 8p12-p11 dual specificity phosphatase 4 HAL 0.51 12q22-q24.1histidine ammonia-lyase PDE4D 0.499 5q12 phosphodiesterase 4D, cAMP-specific FURIN 0.497 15q26.1 furin (paired basic amino acid cleaving enzyme) CPS1 0.49 2q35 carbamoyl-phosphate synthase 1, mitochondrial TESC 0.478 12q24.22 tescalcin INHA 0.465 2q35 inhibin, alpha S100P 0.461 4p16 S100 calcium binding protein P VPS37A 0.447 8p22 vacuolar protein sorting 37 homolog A (S. cerevisiae) SLC16A14 0.447 2q36.3 solute carrier family 16, member 14 PPARGC1A 0.443 4p15.1 peroxisome proliferator-activated receptor gamma, coactivator 1 alpha SIK1 0.435 21q22.3 salt-inducible kinase 1 IRS2 0.434 13q34 insulin receptor substrate 2 RND1 0.433 12q12 Rho family GTPase 1 HGD 0.433 3q13.33 homogentisate 1,2-dioxygenase PTP4A1 0.432 6q12 protein tyrosine phosphatase type IVA, member 1 C8orf4 0.428 8p11.2 chromosome 8 open reading frame 4 DDC 0.427 7p12.2 dopa decarboxylase (aromatic L-amino acid decarboxylase) TACC2 0.427 10q26 transforming, acidic coiled-coil containing protein 2 MUC13 0.422 3q21.2 mucin 13, cell surface associated C5 0.412 9q33-q34 complement component 5 NR4A2 0.412 2q22-q23 nuclear receptor subfamily 4, group A, member 2 EYS 0.411 6q12 eyes shut homolog (Drosophila) GPX2 0.406 14q24.1 glutathione peroxidase -

Table S2.Up Or Down Regulated Genes in Tcof1 Knockdown Neuroblastoma N1E-115 Cells Involved in Differentbiological Process Anal
Table S2.Up or down regulated genes in Tcof1 knockdown neuroblastoma N1E-115 cells involved in differentbiological process analysed by DAVID database Pop Pop Fold Term PValue Genes Bonferroni Benjamini FDR Hits Total Enrichment GO:0044257~cellular protein catabolic 2.77E-10 MKRN1, PPP2R5C, VPRBP, MYLIP, CDC16, ERLEC1, MKRN2, CUL3, 537 13588 1.944851 8.64E-07 8.64E-07 5.02E-07 process ISG15, ATG7, PSENEN, LOC100046898, CDCA3, ANAPC1, ANAPC2, ANAPC5, SOCS3, ENC1, SOCS4, ASB8, DCUN1D1, PSMA6, SIAH1A, TRIM32, RNF138, GM12396, RNF20, USP17L5, FBXO11, RAD23B, NEDD8, UBE2V2, RFFL, CDC GO:0051603~proteolysis involved in 4.52E-10 MKRN1, PPP2R5C, VPRBP, MYLIP, CDC16, ERLEC1, MKRN2, CUL3, 534 13588 1.93519 1.41E-06 7.04E-07 8.18E-07 cellular protein catabolic process ISG15, ATG7, PSENEN, LOC100046898, CDCA3, ANAPC1, ANAPC2, ANAPC5, SOCS3, ENC1, SOCS4, ASB8, DCUN1D1, PSMA6, SIAH1A, TRIM32, RNF138, GM12396, RNF20, USP17L5, FBXO11, RAD23B, NEDD8, UBE2V2, RFFL, CDC GO:0044265~cellular macromolecule 6.09E-10 MKRN1, PPP2R5C, VPRBP, MYLIP, CDC16, ERLEC1, MKRN2, CUL3, 609 13588 1.859332 1.90E-06 6.32E-07 1.10E-06 catabolic process ISG15, RBM8A, ATG7, LOC100046898, PSENEN, CDCA3, ANAPC1, ANAPC2, ANAPC5, SOCS3, ENC1, SOCS4, ASB8, DCUN1D1, PSMA6, SIAH1A, TRIM32, RNF138, GM12396, RNF20, XRN2, USP17L5, FBXO11, RAD23B, UBE2V2, NED GO:0030163~protein catabolic process 1.81E-09 MKRN1, PPP2R5C, VPRBP, MYLIP, CDC16, ERLEC1, MKRN2, CUL3, 556 13588 1.87839 5.64E-06 1.41E-06 3.27E-06 ISG15, ATG7, PSENEN, LOC100046898, CDCA3, ANAPC1, ANAPC2, ANAPC5, SOCS3, ENC1, SOCS4, -

Targeting RIP Kinases in Chronic Inflammatory Disease
biomolecules Review Targeting RIP Kinases in Chronic Inflammatory Disease Mary Speir 1,2, Tirta M. Djajawi 1,2 , Stephanie A. Conos 1,2, Hazel Tye 1 and Kate E. Lawlor 1,2,* 1 Centre for Innate Immunity and Infectious Diseases, Hudson Institute of Medical Research, Clayton, VIC 3168, Australia; [email protected] (M.S.); [email protected] (T.M.D.); [email protected] (S.A.C.); [email protected] (H.T.) 2 Department of Molecular and Translational Science, Monash University, Clayton, VIC 3168, Australia * Correspondence: [email protected]; Tel.: +61-85722700 Abstract: Chronic inflammatory disorders are characterised by aberrant and exaggerated inflam- matory immune cell responses. Modes of extrinsic cell death, apoptosis and necroptosis, have now been shown to be potent drivers of deleterious inflammation, and mutations in core repressors of these pathways underlie many autoinflammatory disorders. The receptor-interacting protein (RIP) kinases, RIPK1 and RIPK3, are integral players in extrinsic cell death signalling by regulating the production of pro-inflammatory cytokines, such as tumour necrosis factor (TNF), and coordinating the activation of the NOD-like receptor protein 3 (NLRP3) inflammasome, which underpin patholog- ical inflammation in numerous chronic inflammatory disorders. In this review, we firstly give an overview of the inflammatory cell death pathways regulated by RIPK1 and RIPK3. We then discuss how dysregulated signalling along these pathways can contribute to chronic inflammatory disorders of the joints, skin, and gastrointestinal tract, and discuss the emerging evidence for targeting these RIP kinases in the clinic. Keywords: apoptosis; necroptosis; RIP kinases; chronic inflammatory disease; tumour necrosis factor; Citation: Speir, M.; Djajawi, T.M.; interleukin-1 Conos, S.A.; Tye, H.; Lawlor, K.E. -
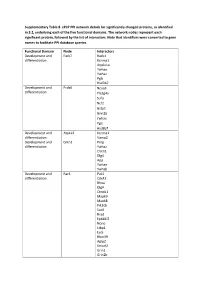
Supplementary Table 8. Cpcp PPI Network Details for Significantly Changed Proteins, As Identified in 3.2, Underlying Each of the Five Functional Domains
Supplementary Table 8. cPCP PPI network details for significantly changed proteins, as identified in 3.2, underlying each of the five functional domains. The network nodes represent each significant protein, followed by the list of interactors. Note that identifiers were converted to gene names to facilitate PPI database queries. Functional Domain Node Interactors Development and Park7 Rack1 differentiation Kcnma1 Atp6v1a Ywhae Ywhaz Pgls Hsd3b7 Development and Prdx6 Ncoa3 differentiation Pla2g4a Sufu Ncf2 Gstp1 Grin2b Ywhae Pgls Hsd3b7 Development and Atp1a2 Kcnma1 differentiation Vamp2 Development and Cntn1 Prnp differentiation Ywhaz Clstn1 Dlg4 App Ywhae Ywhab Development and Rac1 Pak1 differentiation Cdc42 Rhoa Dlg4 Ctnnb1 Mapk9 Mapk8 Pik3cb Sod1 Rrad Epb41l2 Nono Ltbp1 Evi5 Rbm39 Aplp2 Smurf2 Grin1 Grin2b Xiap Chn2 Cav1 Cybb Pgls Ywhae Development and Hbb-b1 Atp5b differentiation Hba Kcnma1 Got1 Aldoa Ywhaz Pgls Hsd3b4 Hsd3b7 Ywhae Development and Myh6 Mybpc3 differentiation Prkce Ywhae Development and Amph Capn2 differentiation Ap2a2 Dnm1 Dnm3 Dnm2 Atp6v1a Ywhab Development and Dnm3 Bin1 differentiation Amph Pacsin1 Grb2 Ywhae Bsn Development and Eef2 Ywhaz differentiation Rpgrip1l Atp6v1a Nphp1 Iqcb1 Ezh2 Ywhae Ywhab Pgls Hsd3b7 Hsd3b4 Development and Gnai1 Dlg4 differentiation Development and Gnao1 Dlg4 differentiation Vamp2 App Ywhae Ywhab Development and Psmd3 Rpgrip1l differentiation Psmd4 Hmga2 Development and Thy1 Syp differentiation Atp6v1a App Ywhae Ywhaz Ywhab Hsd3b7 Hsd3b4 Development and Tubb2a Ywhaz differentiation Nphp4 -

The Combination of TPL2 Knockdown and Tnfα Causes Synthetic Lethality Via Caspase-8 Activation in Human Carcinoma Cell Lines
The combination of TPL2 knockdown and TNFα causes synthetic lethality via caspase-8 activation in human carcinoma cell lines Oksana B. Serebrennikovaa, Maria D. Paraskevopouloua, Elia Aguado-Frailea,1, Vasiliki Tarasliaa, Wenying Rena,2, Geeta Thapaa, Jatin Ropera,3, Keyong Dua,2, Carlo M. Croceb,c,4, and Philip N. Tsichlisa,b,c,4 aMolecular Oncology Research Institute, Tufts Medical Center, Boston, MA 02111; bDepartment of Cancer Biology and Genetics, The Ohio State University, Columbus, OH 43210; and cThe Ohio State University Comprehensive Cancer Center, Columbus, OH 43210 Contributed by Carlo M. Croce, May 21, 2019 (sent for review January 29, 2019; reviewed by Emad S. Alnemri and Wafik El-Deiry) Most normal and tumor cells are protected from tumor necrosis subset of tumor cells and we delineate the relevant TPL2-regulated factor α (TNFα)-induced apoptosis. Here, we identify the MAP3 pathway(s). kinase tumor progression locus-2 (TPL2) as a player contributing TPL2 is an oncoprotein that is activated by provirus insertion to the protection of a subset of tumor cell lines. The combination in Moloney murine leukemia virus-induced rodent lymphomas of TPL2 knockdown and TNFα gives rise to a synthetic lethality and mammary tumor virus-induced mammary adenocarcinomas phenotype via receptor-interacting serine/threonine-protein ki- (13, 14). Expression of constitutively active TPL2 from a thymus- nase 1 (RIPK1)-dependent and -independent mechanisms. Whereas targeted transgene confirmed its oncogenic potential (15). Sub- wild-type TPL2 rescues -

A Post-Transcriptional Mechanism Pacing Expression of Neural Genes with Precursor Cell Differentiation Status
ARTICLE Received 24 Sep 2014 | Accepted 21 May 2015 | Published 6 Jul 2015 DOI: 10.1038/ncomms8576 OPEN A post-transcriptional mechanism pacing expression of neural genes with precursor cell differentiation status Weijun Dai1, Wencheng Li2, Mainul Hoque2, Zhuyun Li1, Bin Tian2 & Eugene V. Makeyev1,3 Nervous system (NS) development relies on coherent upregulation of extensive sets of genes in a precise spatiotemporal manner. How such transcriptome-wide effects are orchestrated at the molecular level remains an open question. Here we show that 30-untranslated regions (30 UTRs) of multiple neural transcripts contain AU-rich cis-elements (AREs) recognized by tristetraprolin (TTP/Zfp36), an RNA-binding protein previously implicated in regulation of mRNA stability. We further demonstrate that the efficiency of ARE-dependent mRNA degradation declines in the neural lineage because of a decrease in the TTP protein expression mediated by the NS-enriched microRNA miR-9. Importantly, TTP downregulation in this context is essential for proper neuronal differentiation. On the other hand, inactivation of TTP in non-neuronal cells leads to dramatic upregulation of multiple NS-specific genes. We conclude that the newly identified miR-9/TTP circuitry limits unscheduled accumulation of neuronal mRNAs in non-neuronal cells and ensures coordinated upregulation of these transcripts in neurons. 1 School of Biological Sciences, Nanyang Technological University, Singapore 637551, Singapore. 2 Department of Microbiology, Biochemistry, and Molecular Genetics, Rutgers New Jersey Medical School, Newark, New Jersey 07103, USA. 3 MRC Centre for Developmental Neurobiology, King’s College London, London SE1 1UL, UK. Correspondence and requests for materials should be addressed to E.V.M. -

RIPK1, Active Recombinant Mouse Protein Expressed in Sf9 Cells
Catalog # Aliquot Size R07M-11G -05 5 µg R07M-11G -10 10 µg RIPK1, Active Recombinant mouse protein expressed in Sf9 cells Catalog # R07M-11G Lot # V2253-5 Product Description Specific Activity Recombinant mouse RIPK1 (1-327) was expressed by baculovirus in Sf9 insect cells using an N-terminal GST tag. 20,000 The RIPK1 gene accession number is NM_009068. 15,000 Gene Aliases 10,000 D330015H01Rik; Rinp; RIP; Rip1 Activity (cpm) 5,000 0 Formulation 0 120 240 360 480 Recombinant protein stored in 50mM Tris-HCl, pH 7.5, Protein (ng) 150mM NaCl, 10mM glutathione, 0.1mM EDTA, 0.25mM The specific activity of RIPK1 was determined to be 2.6 nmol DTT, 0.1mM PMSF, 25% glycerol. /min/mg as per activity assay protocol. Storage and Stability Purity Store product at –70oC. For optimal storage, aliquot target into smaller quantities after centrifugation and store at recommended temperature. For most favorable performance, avoid repeated handling and multiple The purity of RIPK1 was determined freeze/thaw cycles. to be >70% by densitometry, approx. MW 62 kDa. Scientific Background RIPK1 or Receptor Interacting Protein Kinase 1 is a serine/threonine kinase that was originally identified as interacting with the cytoplasmic domain of FAS. RIPK1 has been deemed as an important element in the signal transduction machinery that mediates programmed cell death. RIPK1 has been shown to interact with TRADD, TRAF1, TRAF2 and TRAF3 and TRADD can act as an RIPK1, Active adaptor protein to recruit RIPK1 to the TNFR1 complex in a Recombinant mouse protein expressed in Sf9 cells TNF-dependent process (1).