Chem Compute Quickstart
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

Parameterizing a Novel Residue
University of Illinois at Urbana-Champaign Luthey-Schulten Group, Department of Chemistry Theoretical and Computational Biophysics Group Computational Biophysics Workshop Parameterizing a Novel Residue Rommie Amaro Brijeet Dhaliwal Zaida Luthey-Schulten Current Editors: Christopher Mayne Po-Chao Wen February 2012 CONTENTS 2 Contents 1 Biological Background and Chemical Mechanism 4 2 HisH System Setup 7 3 Testing out your new residue 9 4 The CHARMM Force Field 12 5 Developing Topology and Parameter Files 13 5.1 An Introduction to a CHARMM Topology File . 13 5.2 An Introduction to a CHARMM Parameter File . 16 5.3 Assigning Initial Values for Unknown Parameters . 18 5.4 A Closer Look at Dihedral Parameters . 18 6 Parameter generation using SPARTAN (Optional) 20 7 Minimization with new parameters 32 CONTENTS 3 Introduction Molecular dynamics (MD) simulations are a powerful scientific tool used to study a wide variety of systems in atomic detail. From a standard protein simulation, to the use of steered molecular dynamics (SMD), to modelling DNA-protein interactions, there are many useful applications. With the advent of massively parallel simulation programs such as NAMD2, the limits of computational anal- ysis are being pushed even further. Inevitably there comes a time in any molecular modelling scientist’s career when the need to simulate an entirely new molecule or ligand arises. The tech- nique of determining new force field parameters to describe these novel system components therefore becomes an invaluable skill. Determining the correct sys- tem parameters to use in conjunction with the chosen force field is only one important aspect of the process. -
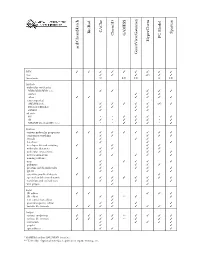
D:\Doc\Workshops\2005 Molecular Modeling\Notebook Pages\Software Comparison\Summary.Wpd
CAChe BioRad Spartan GAMESS Chem3D PC Model HyperChem acd/ChemSketch GaussView/Gaussian WIN TTTT T T T T T mac T T T (T) T T linux/unix U LU LU L LU Methods molecular mechanics MM2/MM3/MM+/etc. T T T T T Amber T T T other TT T T T T semi-empirical AM1/PM3/etc. T T T T T (T) T Extended Hückel T T T T ZINDO T T T ab initio HF * * T T T * T dft T * T T T * T MP2/MP4/G1/G2/CBS-?/etc. * * T T T * T Features various molecular properties T T T T T T T T T conformer searching T T T T T crystals T T T data base T T T developer kit and scripting T T T T molecular dynamics T T T T molecular interactions T T T T movies/animations T T T T T naming software T nmr T T T T T polymers T T T T proteins and biomolecules T T T T T QSAR T T T T scientific graphical objects T T spectral and thermodynamic T T T T T T T T transition and excited state T T T T T web plugin T T Input 2D editor T T T T T 3D editor T T ** T T text conversion editor T protein/sequence editor T T T T various file formats T T T T T T T T Output various renderings T T T T ** T T T T various file formats T T T T ** T T T animation T T T T T graphs T T spreadsheet T T T * GAMESS and/or GAUSSIAN interface ** Text only. -

Jaguar 5.5 User Manual Copyright © 2003 Schrödinger, L.L.C
Jaguar 5.5 User Manual Copyright © 2003 Schrödinger, L.L.C. All rights reserved. Schrödinger, FirstDiscovery, Glide, Impact, Jaguar, Liaison, LigPrep, Maestro, Prime, QSite, and QikProp are trademarks of Schrödinger, L.L.C. MacroModel is a registered trademark of Schrödinger, L.L.C. To the maximum extent permitted by applicable law, this publication is provided “as is” without warranty of any kind. This publication may contain trademarks of other companies. October 2003 Contents Chapter 1: Introduction.......................................................................................1 1.1 Conventions Used in This Manual.......................................................................2 1.2 Citing Jaguar in Publications ...............................................................................3 Chapter 2: The Maestro Graphical User Interface...........................................5 2.1 Starting Maestro...................................................................................................5 2.2 The Maestro Main Window .................................................................................7 2.3 Maestro Projects ..................................................................................................7 2.4 Building a Structure.............................................................................................9 2.5 Atom Selection ..................................................................................................10 2.6 Toolbar Controls ................................................................................................11 -

Porting the DFT Code CASTEP to Gpgpus
Porting the DFT code CASTEP to GPGPUs Toni Collis [email protected] EPCC, University of Edinburgh CASTEP and GPGPUs Outline • Why are we interested in CASTEP and Density Functional Theory codes. • Brief introduction to CASTEP underlying computational problems. • The OpenACC implementation http://www.nu-fuse.com CASTEP: a DFT code • CASTEP is a commercial and academic software package • Capable of Density Functional Theory (DFT) and plane wave basis set calculations. • Calculates the structure and motions of materials by the use of electronic structure (atom positions are dictated by their electrons). • Modern CASTEP is a re-write of the original serial code, developed by Universities of York, Durham, St. Andrews, Cambridge and Rutherford Labs http://www.nu-fuse.com CASTEP: a DFT code • DFT/ab initio software packages are one of the largest users of HECToR (UK national supercomputing service, based at University of Edinburgh). • Codes such as CASTEP, VASP and CP2K. All involve solving a Hamiltonian to explain the electronic structure. • DFT codes are becoming more complex and with more functionality. http://www.nu-fuse.com HECToR • UK National HPC Service • Currently 30- cabinet Cray XE6 system – 90,112 cores • Each node has – 2×16-core AMD Opterons (2.3GHz Interlagos) – 32 GB memory • Peak of over 800 TF and 90 TB of memory http://www.nu-fuse.com HECToR usage statistics Phase 3 statistics (Nov 2011 - Apr 2013) Ab initio codes (VASP, CP2K, CASTEP, ONETEP, NWChem, Quantum Espresso, GAMESS-US, SIESTA, GAMESS-UK, MOLPRO) GS2NEMO ChemShell 2%2% SENGA2% 3% UM Others 4% 34% MITgcm 4% CASTEP 4% GROMACS 6% DL_POLY CP2K VASP 5% 8% 19% http://www.nu-fuse.com HECToR usage statistics Phase 3 statistics (Nov 2011 - Apr 2013) 35% of the Chemistry software on HECToR is using DFT methods. -
Massive-Parallel Implementation of the Resolution-Of-Identity Coupled
Article Cite This: J. Chem. Theory Comput. 2019, 15, 4721−4734 pubs.acs.org/JCTC Massive-Parallel Implementation of the Resolution-of-Identity Coupled-Cluster Approaches in the Numeric Atom-Centered Orbital Framework for Molecular Systems † § † † ‡ § § Tonghao Shen, , Zhenyu Zhu, Igor Ying Zhang,*, , , and Matthias Scheffler † Department of Chemistry, Fudan University, Shanghai 200433, China ‡ Shanghai Key Laboratory of Molecular Catalysis and Innovative Materials, MOE Key Laboratory of Computational Physical Science, Fudan University, Shanghai 200433, China § Fritz-Haber-Institut der Max-Planck-Gesellschaft, Faradayweg 4-6, 14195 Berlin, Germany *S Supporting Information ABSTRACT: We present a massive-parallel implementation of the resolution of identity (RI) coupled-cluster approach that includes single, double, and perturbatively triple excitations, namely, RI-CCSD(T), in the FHI-aims package for molecular systems. A domain-based distributed-memory algorithm in the MPI/OpenMP hybrid framework has been designed to effectively utilize the memory bandwidth and significantly minimize the interconnect communication, particularly for the tensor contraction in the evaluation of the particle−particle ladder term. Our implementation features a rigorous avoidance of the on- the-fly disk storage and excellent strong scaling of up to 10 000 and more cores. Taking a set of molecules with different sizes, we demonstrate that the parallel performance of our CCSD(T) code is competitive with the CC implementations in state-of- the-art high-performance-computing computational chemistry packages. We also demonstrate that the numerical error due to the use of RI approximation in our RI-CCSD(T) method is negligibly small. Together with the correlation-consistent numeric atom-centered orbital (NAO) basis sets, NAO-VCC-nZ, the method is applied to produce accurate theoretical reference data for 22 bio-oriented weak interactions (S22), 11 conformational energies of gaseous cysteine conformers (CYCONF), and 32 Downloaded via FRITZ HABER INST DER MPI on January 8, 2021 at 22:13:06 (UTC). -

Lawrence Berkeley National Laboratory Recent Work
Lawrence Berkeley National Laboratory Recent Work Title From NWChem to NWChemEx: Evolving with the Computational Chemistry Landscape. Permalink https://escholarship.org/uc/item/4sm897jh Journal Chemical reviews, 121(8) ISSN 0009-2665 Authors Kowalski, Karol Bair, Raymond Bauman, Nicholas P et al. Publication Date 2021-04-01 DOI 10.1021/acs.chemrev.0c00998 Peer reviewed eScholarship.org Powered by the California Digital Library University of California From NWChem to NWChemEx: Evolving with the computational chemistry landscape Karol Kowalski,y Raymond Bair,z Nicholas P. Bauman,y Jeffery S. Boschen,{ Eric J. Bylaska,y Jeff Daily,y Wibe A. de Jong,x Thom Dunning, Jr,y Niranjan Govind,y Robert J. Harrison,k Murat Keçeli,z Kristopher Keipert,? Sriram Krishnamoorthy,y Suraj Kumar,y Erdal Mutlu,y Bruce Palmer,y Ajay Panyala,y Bo Peng,y Ryan M. Richard,{ T. P. Straatsma,# Peter Sushko,y Edward F. Valeev,@ Marat Valiev,y Hubertus J. J. van Dam,4 Jonathan M. Waldrop,{ David B. Williams-Young,x Chao Yang,x Marcin Zalewski,y and Theresa L. Windus*,r yPacific Northwest National Laboratory, Richland, WA 99352 zArgonne National Laboratory, Lemont, IL 60439 {Ames Laboratory, Ames, IA 50011 xLawrence Berkeley National Laboratory, Berkeley, 94720 kInstitute for Advanced Computational Science, Stony Brook University, Stony Brook, NY 11794 ?NVIDIA Inc, previously Argonne National Laboratory, Lemont, IL 60439 #National Center for Computational Sciences, Oak Ridge National Laboratory, Oak Ridge, TN 37831-6373 @Department of Chemistry, Virginia Tech, Blacksburg, VA 24061 4Brookhaven National Laboratory, Upton, NY 11973 rDepartment of Chemistry, Iowa State University and Ames Laboratory, Ames, IA 50011 E-mail: [email protected] 1 Abstract Since the advent of the first computers, chemists have been at the forefront of using computers to understand and solve complex chemical problems. -

Computational Chemistry
Computational Chemistry as a Teaching Aid Melissa Boule Aaron Harshman Rexford Hoadley An Interactive Qualifying Project Report Submitted to the Faculty of the Worcester Polytechnic Institute In partial fulfillment of the requirements for the Bachelor of Science Degree Approved by: Professor N. Aaron Deskins 1i Abstract Molecular modeling software has transformed the capabilities of researchers. Molecular modeling software has the potential to be a helpful teaching tool, though as of yet its efficacy as a teaching technique has yet to be proven. This project focused on determining the effectiveness of a particular molecular modeling software in high school classrooms. Our team researched what topics students struggled with, surveyed current high school chemistry teachers, chose a modeling software, and then developed a lesson plan around these topics. Lastly, we implemented the lesson in two separate high school classrooms. We concluded that these programs show promise for the future, but with the current limitations of high school technology, these tools may not be as impactful. 2ii Acknowledgements This project would not have been possible without the help of many great educators. Our advisor Professor N. Aaron Deskins provided driving force to ensure the research ran smoothly and efficiently. Mrs. Laurie Hanlan and Professor Drew Brodeur provided insight and guidance early in the development of our topic. Mr. Mark Taylor set-up our WebMO server and all our other technology needs. Lastly, Mrs. Pamela Graves and Mr. Eric Van Inwegen were instrumental by allowing us time in their classrooms to implement our lesson plans. Without the support of this diverse group of people this project would never have been possible. -

Efficient Two-Component Relativistic Method to Obtain Electronic
View metadata, citation and similar papers at core.ac.uk brought to you by CORE provided by DSpace at Waseda University Efficient Two-Component Relativistic Method to Obtain Electronic and Molecular Structures of Heavy-Element Systems ͷిࢠঢ়ଶ͓ΑͼࢠߏͷͨΊͷܥॏݩૉ ޮతͳ 2 ૬ର๏ February 2017 Waseda University Graduate School of Advanced Science and Engineering Department of Chemistry and Biochemistry, Research on Electronic State Theory Yuya NAKAJIMA தౢɹ༟ Contents Chapter 1 General introduction 1 References . ................................ 6 Chapter 2 Theoretical background 7 2.1 Dirac Hamiltonian . ....................... 7 2.2 The IODKH method ....................... 9 2.3 The LUT scheme . ........................ 12 2.4 Spin-free and spin-dependent formalisms . ......... 14 2.5 The FCP method . ........................ 16 2.6 The GHF method . ........................ 19 2.7 The Analytical energy derivative for the GHF method . 21 References . ................................ 24 Chapter 3 Analytical energy gradient for spin-free infinite-order Douglas–Kroll–Hess method with local unitary transfor- mation 25 3.1 Introduction . .......................... 25 3.2 Theory and implementation . ................. 27 3.2.1 Energy gradient for IODKH . ........... 27 3.2.2 Analytical derivative for the space transformation matrices . ......................... 30 3.2.3 Energy gradient for LUT-IODKH . .......... 32 3.2.4 Implementation . ................... 33 i 3.3 Numerical assessments . .................. 34 3.3.1 Computational details . ................ 34 3.3.2 Numerical gradient values ............... 37 3.3.3 Accuracies of IODKH/C and LUT-IODKH/C methods 38 3.3.4 Computational cost of the LUT scheme ........ 45 3.3.5 Metal complexes . ................... 47 3.3.6 Heavier analogues of ethylene . ........... 48 3.3.7 Harmonic frequencies of diatomic molecules . 49 3.4 Conclusion . -

1 Advances in Electronic Structure Theory: Gamess a Decade Later Mark S. Gordon and Michael W. Schmidt Department of Chemistry A
ADVANCES IN ELECTRONIC STRUCTURE THEORY: GAMESS A DECADE LATER MARK S. GORDON AND MICHAEL W. SCHMIDT DEPARTMENT OF CHEMISTRY AND AMES LABORATORY, IOWA STATE UNIVERSITY, AMES, IA 50011 Abstract. Recent developments in advanced quantum chemistry and quantum chemistry interfaced with model potentials are discussed, with the primary focus on new implementations in the GAMESS electronic structure suite of programs. Applications to solvent effects and surface science are discussed. 1. Introduction The past decade has seen an extraordinary growth in novel new electronic structure methods and creative implementations of these methods. Concurrently, there have been important advances in “middleware”, software that enables the implementation of efficient electronic structure algorithms. Combined with continuing improvements in computer and interconnect hardware, these advances have extended both the accuracy of computations and the sizes of molecular systems to which such methods may be applied. The great majority of the new computational chemistry algorithms have found their way into one or more broadly distributed electronic structure packages. Since this work focuses on new features of the GAMESS (General Atomic and Molecular Electronic Structure System1) suite of codes, it is important at the outset to recognize the many other packages that offer both similar and complementary features. These include ACES2 CADPAC3, DALTON4, GAMESS-UK5, HYPERCHEM6, JAGUAR7, MOLCAS8 MOLPRO9, NWChem10, PQS11, PSI312, Q-Chem13, SPARTAN14, TURBOMOLE15, and UT-Chem16. Some1,3,4,10 of these packages are distributed at no cost to all, or to academic users, while others are commercial packages, but all are generally available to users without restriction or constraint. Indeed, the developers of these codes frequently collaborate to share features, a practice that clearly benefits all of their users. -

Computational Chemistry: a Practical Guide for Applying Techniques to Real-World Problems
Computational Chemistry: A Practical Guide for Applying Techniques to Real-World Problems. David C. Young Copyright ( 2001 John Wiley & Sons, Inc. ISBNs: 0-471-33368-9 (Hardback); 0-471-22065-5 (Electronic) COMPUTATIONAL CHEMISTRY COMPUTATIONAL CHEMISTRY A Practical Guide for Applying Techniques to Real-World Problems David C. Young Cytoclonal Pharmaceutics Inc. A JOHN WILEY & SONS, INC., PUBLICATION New York . Chichester . Weinheim . Brisbane . Singapore . Toronto Designations used by companies to distinguish their products are often claimed as trademarks. In all instances where John Wiley & Sons, Inc., is aware of a claim, the product names appear in initial capital or all capital letters. Readers, however, should contact the appropriate companies for more complete information regarding trademarks and registration. Copyright ( 2001 by John Wiley & Sons, Inc. All rights reserved. No part of this publication may be reproduced, stored in a retrieval system or transmitted in any form or by any means, electronic or mechanical, including uploading, downloading, printing, decompiling, recording or otherwise, except as permitted under Sections 107 or 108 of the 1976 United States Copyright Act, without the prior written permission of the Publisher. Requests to the Publisher for permission should be addressed to the Permissions Department, John Wiley & Sons, Inc., 605 Third Avenue, New York, NY 10158-0012, (212) 850-6011, fax (212) 850-6008, E-Mail: PERMREQ @ WILEY.COM. This publication is designed to provide accurate and authoritative information in regard to the subject matter covered. It is sold with the understanding that the publisher is not engaged in rendering professional services. If professional advice or other expert assistance is required, the services of a competent professional person should be sought. -

CGPLUS a Package Supporting the CHARMM/GAMESSPLUS Combination Package for Incorporating the Generalized Hybrid Orbital QM/MM Methods of GAMESSPLUS Into CHARMM
CGPLUS A Package Supporting the CHARMM/GAMESSPLUS Combination Package for Incorporating the Generalized Hybrid Orbital QM/MM Methods of GAMESSPLUS Into CHARMM Users Manual Version 2008 date of finalization of the latest version of software: April 18, 2008 date of most recent change in this document: April 18, 2008 Jingzhi Pu, Masahiro Higashi, Jiali Gao, and Donald G. Truhlar Department of Chemistry and Supercomputer Institute, University of Minnesota, Minneapolis, MN 55455-0431, U. S. A. Distribution site: http://comp.chem.umn.edu/cgplus The code and manual are copyrighted, 2004-2008. Contents CGPLUS Abstract ............................................................................................................... 1 Introduction ......................................................................................................................... 1 Referencing CGPLUS ......................................................................................................... 2 Utility for Modifying CHARMM and GAMESSPLUS ........................................................ 4 Program Distribution .......................................................................................................... 5 Description of the CGPLUS Installation Script "install_cgplus.com" ................................ 6 Description of the Modifications of CHARMM Files Made by CGPLUS .......................... 7 Description of the Make Files and Include File Provided by CGPLUS ............................. 8 Compiling CHARMM with GAMESSPLUS....................................................................... -

The Aomix Manual and the FAQ Webpage, You Cannot Resolve Your Problem, Contact the Aomix Developer with the Detailed Description of Your Problem
AOMix version 6.94 Software Manual Dr. S. I. Gorelsky, AOMix manual (www.sg-chem.net) Updated on January 21, 2021 AOMix is a user-friendly, comprehensive package for the molecular orbital analysis. AOMix calculates percentage contributions of different molecular fragments (atoms, ligands, groups of atomic orbitals / basis functions, groups of fragment molecular orbitals, etc.) to molecular orbitals from output files generated by different computational chemistry packages and produces data tables (in the ASCII text format) with relevant MO information, condensed Fukui functions, etc. In addition, it generates total, partial and overlap population density-of-states (DOS) plots and can be used for MO composition analysis in systems with many fragments. It also calculates the MO compositions in the basis of fragment molecular orbitals (FOs), occupation numbers for FOs and atomic orbitals (AOs), and, if the number of fragments is greater than 1, the amounts of electron donation / back-donation between molecular fragments (charge decomposition analysis, CDA), electronic polarizations of fragments, and generates plot data for MO interaction diagrams. In addition, it can be used for Morokuma’s energy decomposition analysis (EDA) and to generate a guess wave function of multi-fragment molecular systems from the wave functions of fragments. The software calculates total and free valence indices of fragments, 2-center (Wiberg, Löwdin, and Mayer) and 3-, 4-, 5- and 6-center bond orders between molecular fragments (which can be defined as atoms, groups of atoms, or groups of atomic orbitals) and performs the Löwdin population analysis. The software can be also used for recovery of the initial guess (as the converged wave function) and the analysis of spin-unrestricted MO calculations: the program projects β-spin molecular orbitals on to -spin molecular orbitals and prints the overlap matrix i j .