KIAA0556 Is a Novel Ciliary Basal Body Component Mutated in Joubert Syndrome Anna A
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-
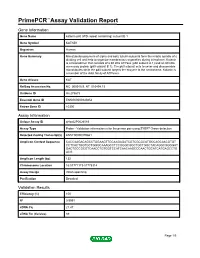
Primepcr™Assay Validation Report
PrimePCR™Assay Validation Report Gene Information Gene Name katanin p80 (WD repeat containing) subunit B 1 Gene Symbol KATNB1 Organism Human Gene Summary Microtubules polymers of alpha and beta tubulin subunits form the mitotic spindle of a dividing cell and help to organize membranous organelles during interphase. Katanin is a heterodimer that consists of a 60 kDa ATPase (p60 subunit A 1) and an 80 kDa accessory protein (p80 subunit B 1). The p60 subunit acts to sever and disassemble microtubules while the p80 subunit targets the enzyme to the centrosome. Katanin is a member of the AAA family of ATPases. Gene Aliases KAT RefSeq Accession No. NC_000016.9, NT_010498.15 UniGene ID Hs.275675 Ensembl Gene ID ENSG00000140854 Entrez Gene ID 10300 Assay Information Unique Assay ID qHsaCIP0026589 Assay Type Probe - Validation information is for the primer pair using SYBR® Green detection Detected Coding Transcript(s) ENST00000379661 Amplicon Context Sequence CACCAAGACAGCCTGGAAGTTGCAAGAGATCGTCGCGCATGCCAGCAACGTGT CCTCACTGGTGCTGGGCAAAGCCTCCGGGCGGCTGCTGGCTACAGGCGGGGAT GACTGCCGCGTCAACCTGTGGTCCATCAACAAGCCCAACTGCATCATGAGCCTG ACG Amplicon Length (bp) 132 Chromosome Location 16:57771173-57778314 Assay Design Intron-spanning Purification Desalted Validation Results Efficiency (%) 100 R2 0.9991 cDNA Cq 21.47 cDNA Tm (Celsius) 89 Page 1/5 PrimePCR™Assay Validation Report gDNA Cq 35.43 Specificity (%) 100 Information to assist with data interpretation is provided at the end of this report. Page 2/5 PrimePCR™Assay Validation Report KATNB1, Human Amplification -

Dual Proteome-Scale Networks Reveal Cell-Specific Remodeling of the Human Interactome
bioRxiv preprint doi: https://doi.org/10.1101/2020.01.19.905109; this version posted January 19, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. All rights reserved. No reuse allowed without permission. Dual Proteome-scale Networks Reveal Cell-specific Remodeling of the Human Interactome Edward L. Huttlin1*, Raphael J. Bruckner1,3, Jose Navarrete-Perea1, Joe R. Cannon1,4, Kurt Baltier1,5, Fana Gebreab1, Melanie P. Gygi1, Alexandra Thornock1, Gabriela Zarraga1,6, Stanley Tam1,7, John Szpyt1, Alexandra Panov1, Hannah Parzen1,8, Sipei Fu1, Arvene Golbazi1, Eila Maenpaa1, Keegan Stricker1, Sanjukta Guha Thakurta1, Ramin Rad1, Joshua Pan2, David P. Nusinow1, Joao A. Paulo1, Devin K. Schweppe1, Laura Pontano Vaites1, J. Wade Harper1*, Steven P. Gygi1*# 1Department of Cell Biology, Harvard Medical School, Boston, MA, 02115, USA. 2Broad Institute, Cambridge, MA, 02142, USA. 3Present address: ICCB-Longwood Screening Facility, Harvard Medical School, Boston, MA, 02115, USA. 4Present address: Merck, West Point, PA, 19486, USA. 5Present address: IQ Proteomics, Cambridge, MA, 02139, USA. 6Present address: Vor Biopharma, Cambridge, MA, 02142, USA. 7Present address: Rubius Therapeutics, Cambridge, MA, 02139, USA. 8Present address: RPS North America, South Kingstown, RI, 02879, USA. *Correspondence: [email protected] (E.L.H.), [email protected] (J.W.H.), [email protected] (S.P.G.) #Lead Contact: [email protected] bioRxiv preprint doi: https://doi.org/10.1101/2020.01.19.905109; this version posted January 19, 2020. The copyright holder for this preprint (which was not certified by peer review) is the author/funder. -

Microtubule-Severing Enzymes at the Cutting Edge
Commentary 2561 Microtubule-severing enzymes at the cutting edge David J. Sharp1,* and Jennifer L. Ross2 1Department of Physiology and Biophysics, Albert Einstein College of Medicine, 1300 Morris Park Avenue, Bronx, NY 10461, USA 2Department of Physics, University of Massachusetts-Amherst, 302 Hasbrouck Laboratory, Amherst, MA 01003, USA *Author for correspondence ([email protected]) Journal of Cell Science 125, 2561–2569 ß 2012. Published by The Company of Biologists Ltd doi: 10.1242/jcs.101139 Summary ATP-dependent severing of microtubules was first reported in Xenopus laevis egg extracts in 1991. Two years later this observation led to the purification of the first known microtubule-severing enzyme, katanin. Katanin homologs have now been identified throughout the animal kingdom and in plants. Moreover, members of two closely related enzyme subfamilies, spastin and fidgetin, have been found to sever microtubules and might act alongside katanins in some contexts (Roll-Mecak and McNally, 2010; Yu et al., 2008; Zhang et al., 2007). Over the past few years, it has become clear that microtubule-severing enzymes contribute to a wide range of cellular activities including mitosis and meiosis, morphogenesis, cilia biogenesis and disassembly, and migration. Thus, this group of enzymes is revealing itself to be among the most important of the microtubule regulators. This Commentary focuses on our growing understanding of how microtubule-severing enzymes contribute to the organization and dynamics of diverse microtubule arrays, as well as the structural and biophysical characteristics that afford them the unique capacity to catalyze the removal of tubulin from the interior microtubule lattice. Our goal is to provide a broader perspective, focusing on a limited number of particularly informative, representative and/or timely findings. -

Katanin-P80 Gene Promoter Characterization and Regulation Via Elk1
Katanin-p80 Gene Promoter Characterization and Regulation via Elk1 Ece Selc¸uk., Koray Kırımtay., Derya Canbaz, Gu¨ her Is¸ık Cesur, Sirin Korulu, Arzu Karabay* Department of Molecular Biology and Genetics, Istanbul Technical University, Istanbul, Turkey Abstract Katanin is an ATPase family member protein that participates in microtubule severing. It has heterodimeric structure consisting of 60 kDa (katanin-p60) and 80 kDa (katanin-p80) subunits encoded by KATNA1 and KATNB1 genes, respectively. Katanin-p60 has the enzymatic activity for microtubule severing, whereas katanin-p80 consists of multiple domains with different functions such as targeting katanin-p60 to the centrosome, augmenting microtubule severing by katanin-p60, and even suppressing microtubule severing. Despite the various important functions of katanin-p80, its transcriptional regulation has not been studied yet. Elk1 transcription factor has been shown to interact with microtubules and regulate the transcription of another microtubule severing protein, spastin. In spite of katanin’s importance, and structural and functional similarities to spastin, there is no study on the transcriptional regulation of katanin yet. In this study, we aimed to characterize KATNB1 promoter and analyze the effects of Elk1 on katanin-p80 expression. We identified a 518- bp TATA-less promoter including a critical CpG island and GC boxes as an optimal promoter, and sequential deletion of CpG island and the GC elements gradually decreased the KATNB1 promoter activity. In addition, we showed Elk1 binding on the KATNB1 promoter by EMSA. We found that Elk1 activated KATNB1 promoter, and increased both mRNA and protein levels of katanin- p80 in SH-SY5Y cells. On the other hand, KCl treatment increasing SUMOylation decreased KATNB1 promoter activity. -

Supplemental Information
Supplemental information Dissection of the genomic structure of the miR-183/96/182 gene. Previously, we showed that the miR-183/96/182 cluster is an intergenic miRNA cluster, located in a ~60-kb interval between the genes encoding nuclear respiratory factor-1 (Nrf1) and ubiquitin-conjugating enzyme E2H (Ube2h) on mouse chr6qA3.3 (1). To start to uncover the genomic structure of the miR- 183/96/182 gene, we first studied genomic features around miR-183/96/182 in the UCSC genome browser (http://genome.UCSC.edu/), and identified two CpG islands 3.4-6.5 kb 5’ of pre-miR-183, the most 5’ miRNA of the cluster (Fig. 1A; Fig. S1 and Seq. S1). A cDNA clone, AK044220, located at 3.2-4.6 kb 5’ to pre-miR-183, encompasses the second CpG island (Fig. 1A; Fig. S1). We hypothesized that this cDNA clone was derived from 5’ exon(s) of the primary transcript of the miR-183/96/182 gene, as CpG islands are often associated with promoters (2). Supporting this hypothesis, multiple expressed sequences detected by gene-trap clones, including clone D016D06 (3, 4), were co-localized with the cDNA clone AK044220 (Fig. 1A; Fig. S1). Clone D016D06, deposited by the German GeneTrap Consortium (GGTC) (http://tikus.gsf.de) (3, 4), was derived from insertion of a retroviral construct, rFlpROSAβgeo in 129S2 ES cells (Fig. 1A and C). The rFlpROSAβgeo construct carries a promoterless reporter gene, the β−geo cassette - an in-frame fusion of the β-galactosidase and neomycin resistance (Neor) gene (5), with a splicing acceptor (SA) immediately upstream, and a polyA signal downstream of the β−geo cassette (Fig. -

Chromosome Substitution Strain Assessment of a Huntington's
Mamm Genome DOI 10.1007/s00335-014-9552-9 Chromosome substitution strain assessment of a Huntington’s disease modifier locus Eliana Marisa Ramos • Marina Kovalenko • Jolene R. Guide • Jason St. Claire • Tammy Gillis • Jayalakshmi S. Mysore • Jorge Sequeiros • Vanessa C. Wheeler • Isabel Alonso • Marcy E. MacDonald Received: 26 August 2014 / Accepted: 3 December 2014 Ó The Author(s) 2015. This article is published with open access at Springerlink.com Abstract Huntington’s disease (HD) is a dominant neu- with the human 6q23–24 region, is derived from the A/J rodegenerative disorder that is due to expansion of an (AJ) strain. Crosses were performed to assess the possi- unstable HTT CAG repeat for which genome-wide genetic bility of dominantly acting chr10 AJ-B6J variants of strong scans are now revealing chromosome regions that contain effect that may modulate CAG-dependent HdhQ111/? phe- disease-modifying genes. We have explored a novel notypes. Testing of F1 progeny confirmed that a single AJ human–mouse cross-species functional prioritisation chromosome had a significant effect on the rate of body approach, by evaluating the HD modifier 6q23–24 linkage weight gain and in HdhQ111 mice the AJ chromosome was interval. This unbiased strategy employs C57BL/6J (B6J) associated subtle alterations in somatic CAG instability in HdhQ111 knock-in mice, replicates of the HD mutation, and the liver and the formation of intra-nuclear inclusions, as the C57BL/6J-chr10A/J/NaJ chromosome substitution strain well as DARPP-32 levels, in the striatum. These findings in (CSS10), in which only chromosome 10 (chr10), in synteny relatively small cohorts are suggestive of dominant chr10 AJ-B6 variants that may modify effects of the CAG expansion, and encourage a larger study with CSS10 and sub-strains. -

Variation in Protein Coding Genes Identifies Information
bioRxiv preprint doi: https://doi.org/10.1101/679456; this version posted June 21, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. Animal complexity and information flow 1 1 2 3 4 5 Variation in protein coding genes identifies information flow as a contributor to 6 animal complexity 7 8 Jack Dean, Daniela Lopes Cardoso and Colin Sharpe* 9 10 11 12 13 14 15 16 17 18 19 20 21 22 23 24 Institute of Biological and Biomedical Sciences 25 School of Biological Science 26 University of Portsmouth, 27 Portsmouth, UK 28 PO16 7YH 29 30 * Author for correspondence 31 [email protected] 32 33 Orcid numbers: 34 DLC: 0000-0003-2683-1745 35 CS: 0000-0002-5022-0840 36 37 38 39 40 41 42 43 44 45 46 47 48 49 Abstract bioRxiv preprint doi: https://doi.org/10.1101/679456; this version posted June 21, 2019. The copyright holder for this preprint (which was not certified by peer review) is the author/funder, who has granted bioRxiv a license to display the preprint in perpetuity. It is made available under aCC-BY-NC-ND 4.0 International license. Animal complexity and information flow 2 1 Across the metazoans there is a trend towards greater organismal complexity. How 2 complexity is generated, however, is uncertain. Since C.elegans and humans have 3 approximately the same number of genes, the explanation will depend on how genes are 4 used, rather than their absolute number. -

Aspm (NM 009791) Mouse Tagged ORF Clone Product Data
OriGene Technologies, Inc. 9620 Medical Center Drive, Ste 200 Rockville, MD 20850, US Phone: +1-888-267-4436 [email protected] EU: [email protected] CN: [email protected] Product datasheet for MR219160 Aspm (NM_009791) Mouse Tagged ORF Clone Product data: Product Type: Expression Plasmids Product Name: Aspm (NM_009791) Mouse Tagged ORF Clone Tag: Myc-DDK Symbol: Aspm Synonyms: Calmbp1; D330028K02Rik; MCPH5; Sha1 Vector: pCMV6-Entry (PS100001) E. coli Selection: Kanamycin (25 ug/mL) Cell Selection: Neomycin ORF Nucleotide >MR219160 representing NM_009791 Sequence: Red=Cloning site Blue=ORF Green=Tags(s) TTTTGTAATACGACTCACTATAGGGCGGCCGGGAATTCGTCGACTGGATCCGGTACCGAGGAGATCTGCC GCCGCGATCGCC ATGGCGACGATGCAGGCAGCCTCCTGCCCAGAGGAGAGGGGGCGGCGGGCGCGACCAGATCCTGAGGCCG GGGACCCGTCTCCGCCGGTGCTGTTGCTCAGCCACTTCTGCGGCGTTCCCTTCCTCTGCTTCGGGGATGT CCGCGTGGGCACGTCGCGGACGCGGTCTCTGGTCCTGCACAACCCGCACGAGGAACCTCTGCAGGTGGAG CTGTCGCTGCTGCGCGCCGCAGGCCAGGGCTTCAGCGTGGCGCCGAACCGCTGCGAGCTGAAGCCTAAAG AAAAACTTACCATTTCCGTTACCTGGACGCCACTGCGAGAAGGGGGAGTGAGGGAGATTGTCACATTTCT TGTAAATGATTTCCTGAAGCACCAGGCTATATTACTAGGAAATGCAGAAGAGCCTAAGAAGAAAAAGAGA AGTCTTTGGAATACCAGTAAGAAGATTCCAGCCTCTTCAAAACATACAAAAAGGACTTCCAAAAACCAAC ATTTTAATGAATCATTTACTATTTCACAAAAAGACAGAATTAGGAGCCCACTGCAGCCTTGTGAAAATCT GGCTATGAGTGAATGCTCTTCCCCAACAGAAAACAAAGTCCCCACCCCATCCATTAGTCCTATTAGAGAA TGCCAGAGTGAAACTTGCTTGCCACTGTTTTTACGCGAGTCCACTGCCTATTCATCTCTTCATGAATCTG AAAATACACAAAATTTAAAAGTACAAGATGCCAGCATTTCACAAACTTTTGATTTTAATGAGGAAGTCGC AAATGAAACTTTTATTAATCCCATTAGTGTCTGTCACCAGAGTGAAGGGGATAGGAAACTCACGCTTGCC CCAAACTGTTCTTCACCTTTGAATAGTACACAGACCCAAATACACTTTCTAAGTCCAGATTCTTTTGTAA -

Retroviral Vector Integration Deregulates Gene Expression but Has No Consequence on the Biology and Function of Transplanted T Cells
Retroviral vector integration deregulates gene expression but has no consequence on the biology and function of transplanted T cells Alessandra Recchia*†, Chiara Bonini*‡, Zulma Magnani*, Fabrizia Urbinati†, Daniela Sartori*§, Sara Muraro*, Enrico Tagliafico†, Attilio Bondanza*¶, Maria Teresa Lupo Stanghellini‡, Massimo Bernardi‡, Alessandra Pescarollo‡, Fabio Ciceri‡, Claudio Bordignon§¶, and Fulvio Mavilio†ʈ *Cancer Immunotherapy and Gene Therapy Program, and ‡Bone Marrow Transplantation Unit, Istituto Scientifico H. San Raffaele, Via Olgettina 58, 20132 Milan, Italy; †Department of Biomedical Sciences, University of Modena and Reggio Emilia, Via Campi 287, 41100 Modena, Italy; §MolMed S.p.A., Via Olgettina 58, 20132 Milan, Italy; and ¶San Raffaele ‘‘Vita e Salute’’ University, Via Olgettina 58, 20132 Milan, Italy Edited by Malcolm A. Martin, National Institutes of Health, Bethesda, MD, and approved December 6, 2005 (received for review September 1, 2005) The use of retroviral vectors in gene therapy has raised safety is currently profiled to provide a graft-versus-leukemia effect to concerns for the genotoxic risk associated with their uncontrolled patients in relapse after HLA-identical HSC transplantation (14), insertion into the human genome. We have analyzed the conse- or to promote immune reconstitution and prevent relapse in quences of retroviral transduction in T cells from leukemic patients patients undergoing HLA-haploidentical HSC transplantation. In treated with allogeneic stem cell transplantation and donor lym- 46 patients treated since 1994 in both contexts, we were able to phocytes genetically modified with a suicide gene (HSV-TK). Ret- control GvHD in 100% of the cases, while preserving antiviral and roviral vectors integrate preferentially within or near transcribed antitumor activity (14–16). -

Genetic and Chemical Modulation of Spastin-Dependent Axon Outgrowth
Disease Models & Mechanisms 3, 743-751 (2010) doi:10.1242/dmm.004002 © 2010. Published by The Company of Biologists Ltd RESEARCH ARTICLE Genetic and chemical modulation of spastin-dependent axon outgrowth in zebrafish embryos indicates a role for impaired microtubule dynamics in hereditary spastic paraplegia Richard Butler1,*,‡, Jonathan D. Wood1,2,*,§, Jennifer A. Landers1 and Vincent T. Cunliffe1,§ SUMMARY Mutations in the SPAST (SPG4) gene, which encodes the microtubule-severing protein spastin, are the most common cause of autosomal dominant hereditary spastic paraplegia (HSP). Following on from previous work in our laboratory showing that spastin is required for axon outgrowth, we report here that the related microtubule-severing protein katanin is also required for axon outgrowth in vivo. Using confocal time-lapse imaging, we have identified requirements for spastin and katanin in maintaining normal axonal microtubule dynamics and growth cone motility in vivo, supporting a model in which microtubule severing is required for concerted growth of neuronal microtubules. Simultaneous knockdown of spastin and katanin caused a more severe phenotype than did individual knockdown of either gene, suggesting that they have different but related functions in supporting axon outgrowth. In addition, the microtubule-destabilising drug nocodazole abolished microtubule dynamics and growth cone motility, and enhanced phenotypic severity in spast-knockdown zebrafish embryos. Thus, disruption of microtubule dynamics might underlie neuronal DMM dysfunction -

Host Cell Factors Necessary for Influenza a Infection: Meta-Analysis of Genome Wide Studies
Host Cell Factors Necessary for Influenza A Infection: Meta-Analysis of Genome Wide Studies Juliana S. Capitanio and Richard W. Wozniak Department of Cell Biology, Faculty of Medicine and Dentistry, University of Alberta Abstract: The Influenza A virus belongs to the Orthomyxoviridae family. Influenza virus infection occurs yearly in all countries of the world. It usually kills between 250,000 and 500,000 people and causes severe illness in millions more. Over the last century alone we have seen 3 global influenza pandemics. The great human and financial cost of this disease has made it the second most studied virus today, behind HIV. Recently, several genome-wide RNA interference studies have focused on identifying host molecules that participate in Influen- za infection. We used nine of these studies for this meta-analysis. Even though the overlap among genes identified in multiple screens was small, network analysis indicates that similar protein complexes and biological functions of the host were present. As a result, several host gene complexes important for the Influenza virus life cycle were identified. The biological function and the relevance of each identified protein complex in the Influenza virus life cycle is further detailed in this paper. Background and PA bound to the viral genome via nucleoprotein (NP). The viral core is enveloped by a lipid membrane derived from Influenza virus the host cell. The viral protein M1 underlies the membrane and anchors NEP/NS2. Hemagglutinin (HA), neuraminidase Viruses are the simplest life form on earth. They parasite host (NA), and M2 proteins are inserted into the envelope, facing organisms and subvert the host cellular machinery for differ- the viral exterior. -

A High-Throughput Approach to Uncover Novel Roles of APOBEC2, a Functional Orphan of the AID/APOBEC Family
Rockefeller University Digital Commons @ RU Student Theses and Dissertations 2018 A High-Throughput Approach to Uncover Novel Roles of APOBEC2, a Functional Orphan of the AID/APOBEC Family Linda Molla Follow this and additional works at: https://digitalcommons.rockefeller.edu/ student_theses_and_dissertations Part of the Life Sciences Commons A HIGH-THROUGHPUT APPROACH TO UNCOVER NOVEL ROLES OF APOBEC2, A FUNCTIONAL ORPHAN OF THE AID/APOBEC FAMILY A Thesis Presented to the Faculty of The Rockefeller University in Partial Fulfillment of the Requirements for the degree of Doctor of Philosophy by Linda Molla June 2018 © Copyright by Linda Molla 2018 A HIGH-THROUGHPUT APPROACH TO UNCOVER NOVEL ROLES OF APOBEC2, A FUNCTIONAL ORPHAN OF THE AID/APOBEC FAMILY Linda Molla, Ph.D. The Rockefeller University 2018 APOBEC2 is a member of the AID/APOBEC cytidine deaminase family of proteins. Unlike most of AID/APOBEC, however, APOBEC2’s function remains elusive. Previous research has implicated APOBEC2 in diverse organisms and cellular processes such as muscle biology (in Mus musculus), regeneration (in Danio rerio), and development (in Xenopus laevis). APOBEC2 has also been implicated in cancer. However the enzymatic activity, substrate or physiological target(s) of APOBEC2 are unknown. For this thesis, I have combined Next Generation Sequencing (NGS) techniques with state-of-the-art molecular biology to determine the physiological targets of APOBEC2. Using a cell culture muscle differentiation system, and RNA sequencing (RNA-Seq) by polyA capture, I demonstrated that unlike the AID/APOBEC family member APOBEC1, APOBEC2 is not an RNA editor. Using the same system combined with enhanced Reduced Representation Bisulfite Sequencing (eRRBS) analyses I showed that, unlike the AID/APOBEC family member AID, APOBEC2 does not act as a 5-methyl-C deaminase.