Proteomic Profiling of Phosphoproteins and Secreted
Total Page:16
File Type:pdf, Size:1020Kb
Load more
Recommended publications
-

The Regulation of Carbamoyl Phosphate Synthetase-Aspartate Transcarbamoylase-Dihydroorotase (Cad) by Phosphorylation and Protein-Protein Interactions
THE REGULATION OF CARBAMOYL PHOSPHATE SYNTHETASE-ASPARTATE TRANSCARBAMOYLASE-DIHYDROOROTASE (CAD) BY PHOSPHORYLATION AND PROTEIN-PROTEIN INTERACTIONS Eric M. Wauson A dissertation submitted to the faculty of the University of North Carolina at Chapel Hill in partial fulfillment of the requirements for the degree of Doctor of Philosophy in the Department of Pharmacology. Chapel Hill 2007 Approved by: Lee M. Graves, Ph.D. T. Kendall Harden, Ph.D. Gary L. Johnson, Ph.D. Aziz Sancar M.D., Ph.D. Beverly S. Mitchell, M.D. 2007 Eric M. Wauson ALL RIGHTS RESERVED ii ABSTRACT Eric M. Wauson: The Regulation of Carbamoyl Phosphate Synthetase-Aspartate Transcarbamoylase-Dihydroorotase (CAD) by Phosphorylation and Protein-Protein Interactions (Under the direction of Lee M. Graves, Ph.D.) Pyrimidines have many important roles in cellular physiology, as they are used in the formation of DNA, RNA, phospholipids, and pyrimidine sugars. The first rate- limiting step in the de novo pyrimidine synthesis pathway is catalyzed by the carbamoyl phosphate synthetase II (CPSase II) part of the multienzymatic complex Carbamoyl phosphate synthetase, Aspartate transcarbamoylase, Dihydroorotase (CAD). CAD gene induction is highly correlated to cell proliferation. Additionally, CAD is allosterically inhibited or activated by uridine triphosphate (UTP) or phosphoribosyl pyrophosphate (PRPP), respectively. The phosphorylation of CAD by PKA and ERK has been reported to modulate the response of CAD to allosteric modulators. While there has been much speculation on the identity of CAD phosphorylation sites, no definitive identification of in vivo CAD phosphorylation sites has been performed. Therefore, we sought to determine the specific CAD residues phosphorylated by ERK and PKA in intact cells. -

Proteomic Analysis of the Rad18 Interaction Network in DT40 – a Chicken B Cell Line
Proteomic analysis of the Rad18 interaction network in DT40 – a chicken B cell line Thesis submitted for the degree of Doctor of Natural Sciences at the Faculty of Biology, Ludwig-Maximilians-University Munich 15th January, 2009 Submitted by Sushmita Gowri Sreekumar Chennai, India Completed at the Helmholtz Zentrum München German Research Center for Environmental Health Institute of Clinical Molecular Biology and Tumor Genetics, Munich Examiners: PD Dr. Berit Jungnickel Prof. Heinrich Leonhardt Prof. Friederike Eckardt-Schupp Prof. Harry MacWilliams Date of Examination: 16th June 2009 To my Parents, Sister, Brother & Rajesh Table of Contents 1. SUMMARY ........................................................................................................................ 1 2. INTRODUCTION ............................................................................................................. 2 2.1. MECHANISMS OF DNA REPAIR ......................................................................................... 3 2.2. ADAPTIVE GENETIC ALTERATIONS – AN ADVANTAGE ....................................................... 5 2.3. THE PRIMARY IG DIVERSIFICATION DURING EARLY B CELL DEVELOPMENT ...................... 6 2.4. THE SECONDARY IG DIVERSIFICATION PROCESSES IN THE GERMINAL CENTER .................. 7 2.4.1. Processing of AID induced DNA lesions during adaptive immunity .................. 9 2.5. TARGETING OF SOMATIC HYPERMUTATION TO THE IG LOCI ............................................ 10 2.6. ROLE OF THE RAD6 PATHWAY IN IG DIVERSIFICATION -

Protein Identities in Evs Isolated from U87-MG GBM Cells As Determined by NG LC-MS/MS
Protein identities in EVs isolated from U87-MG GBM cells as determined by NG LC-MS/MS. No. Accession Description Σ Coverage Σ# Proteins Σ# Unique Peptides Σ# Peptides Σ# PSMs # AAs MW [kDa] calc. pI 1 A8MS94 Putative golgin subfamily A member 2-like protein 5 OS=Homo sapiens PE=5 SV=2 - [GG2L5_HUMAN] 100 1 1 7 88 110 12,03704523 5,681152344 2 P60660 Myosin light polypeptide 6 OS=Homo sapiens GN=MYL6 PE=1 SV=2 - [MYL6_HUMAN] 100 3 5 17 173 151 16,91913397 4,652832031 3 Q6ZYL4 General transcription factor IIH subunit 5 OS=Homo sapiens GN=GTF2H5 PE=1 SV=1 - [TF2H5_HUMAN] 98,59 1 1 4 13 71 8,048185945 4,652832031 4 P60709 Actin, cytoplasmic 1 OS=Homo sapiens GN=ACTB PE=1 SV=1 - [ACTB_HUMAN] 97,6 5 5 35 917 375 41,70973209 5,478027344 5 P13489 Ribonuclease inhibitor OS=Homo sapiens GN=RNH1 PE=1 SV=2 - [RINI_HUMAN] 96,75 1 12 37 173 461 49,94108966 4,817871094 6 P09382 Galectin-1 OS=Homo sapiens GN=LGALS1 PE=1 SV=2 - [LEG1_HUMAN] 96,3 1 7 14 283 135 14,70620005 5,503417969 7 P60174 Triosephosphate isomerase OS=Homo sapiens GN=TPI1 PE=1 SV=3 - [TPIS_HUMAN] 95,1 3 16 25 375 286 30,77169764 5,922363281 8 P04406 Glyceraldehyde-3-phosphate dehydrogenase OS=Homo sapiens GN=GAPDH PE=1 SV=3 - [G3P_HUMAN] 94,63 2 13 31 509 335 36,03039959 8,455566406 9 Q15185 Prostaglandin E synthase 3 OS=Homo sapiens GN=PTGES3 PE=1 SV=1 - [TEBP_HUMAN] 93,13 1 5 12 74 160 18,68541938 4,538574219 10 P09417 Dihydropteridine reductase OS=Homo sapiens GN=QDPR PE=1 SV=2 - [DHPR_HUMAN] 93,03 1 1 17 69 244 25,77302971 7,371582031 11 P01911 HLA class II histocompatibility antigen, -
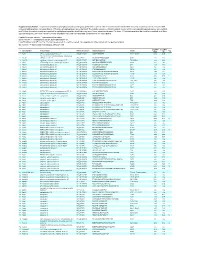
Supplementary Table 1
Supplementary Table 1. Large-scale quantitative phosphoproteomic profiling was performed on paired vehicle- and hormone-treated mTAL-enriched suspensions (n=3). A total of 654 unique phosphopeptides corresponding to 374 unique phosphoproteins were identified. The peptide sequence, phosphorylation site(s), and the corresponding protein name, gene symbol, and RefSeq Accession number are reported for each phosphopeptide identified in any one of three experimental pairs. For those 414 phosphopeptides that could be quantified in all three experimental pairs, the mean Hormone:Vehicle abundance ratio and corresponding standard error are also reported. Peptide Sequence column: * = phosphorylated residue Site(s) column: ^ = ambiguously assigned phosphorylation site Log2(H/V) Mean and SE columns: H = hormone-treated, V = vehicle-treated, n/a = peptide not observable in all 3 experimental pairs Sig. column: * = significantly changed Log 2(H/V), p<0.05 Log (H/V) Log (H/V) # Gene Symbol Protein Name Refseq Accession Peptide Sequence Site(s) 2 2 Sig. Mean SE 1 Aak1 AP2-associated protein kinase 1 NP_001166921 VGSLT*PPSS*PK T622^, S626^ 0.24 0.95 PREDICTED: ATP-binding cassette, sub-family A 2 Abca12 (ABC1), member 12 XP_237242 GLVQVLS*FFSQVQQQR S251^ 1.24 2.13 3 Abcc10 multidrug resistance-associated protein 7 NP_001101671 LMT*ELLS*GIRVLK T464, S468 -2.68 2.48 4 Abcf1 ATP-binding cassette sub-family F member 1 NP_001103353 QLSVPAS*DEEDEVPVPVPR S109 n/a n/a 5 Ablim1 actin-binding LIM protein 1 NP_001037859 PGSSIPGS*PGHTIYAK S51 -3.55 1.81 6 Ablim1 actin-binding -

2. Improvement of Itraq-SPROX Protocol
Development and Application of Mass Spectrometry-Based Approaches for Thermodynamic Analysis of Protein-Ligand Binding Interactions by Xiaopu (Lorrain) Jin Department of Chemistry Duke University Date:_______________________ Approved: ___________________________ Michael C. Fitzgerald, Supervisor ___________________________ Jie Liu ___________________________ Terrence G. Oas ___________________________ Katherine J. Franz Dissertation submitted in partial fulfillment of the requirements for the degree of Doctor of Philosophy in the Department of Chemistry in the Graduate School of Duke University 2017 i v ABSTRACT Development and Application of Mass Spectrometry-Based Approaches for Thermodynamic Analysis of Protein-Ligand Binding Interactions by Xiaopu (Lorrain) Jin Department of Chemistry Duke University Date:_______________________ Approved: ___________________________ Michael C. Fitzgerald, Supervisor ___________________________ Jie Liu ___________________________ Terrence G. Oas ___________________________ Katherine J. Franz An abstract of a dissertation submitted in partial fulfillment of the requirements for the degree of Doctor of Philosophy in the Department of Chemistry in the Graduate School of Duke University 2017 Copyright by Xiaopu (Lorrain) Jin 2017 Abstract The characterization of protein stability changes and protein-ligand interactions on the proteomic scale is important for understanding the biology of cellular processes. The identification and quantification of protein-ligand binding affinities is critical for disease -

ABSTRACT Title of Dissertation: INTEGRATION OF
ABSTRACT Title of Dissertation: INTEGRATION OF 18O LABELING AND SOLUTION ISOELECTRIC FOCUSING IN A SHOTGUN ANALYSIS OF MITOCHONDRIAL PROTEINS Jinshan Wang, Doctor of Philosophy, 2007 Dissertation Directed by: Professor Catherine Fenselau Department of Chemistry and Biochemistry The coupling of efficient separations and mass spectrometry instrumentation is highly desirable to provide global proteomic analysis. When quantitative comparisons are part of the strategy, separation and analytical methods should be selected, which optimize the isotope labeling procedure. Enzyme-catalyzed 18O labeling is considered to be the labeling method most compatible with analysis of proteins from tissue and other limited samples. The introduction of label at the peptide stage mandates that protein manipulation be minimized in favor of peptide fractionation post-labeling. In the present study, forward and reverse 18O labeling are integrated with solution isoelectric focusing and capillary LC-tandem mass spectrometry to study changes in mitochondrial proteins associated with drug resistance in human cancer cells. A total of 637 peptides corresponding to 278 proteins were identified in this analysis. Of these, twelve proteins have been demonstrated from the forward and reverse labeling experiments to have abundances altered by greater than a factor of two between the drug susceptible MCF-7 cell line and the MCF-7 cell line selected for resistance to mitoxantrone. Galectin-3 binding protein precursor was detected in the resistant cell line, but was not detected in the drug susceptible line. Such proteins are challenging to 18O and other isotope strategies and a solution is offered, based on reverse labeling. These twelve proteins play a role in several pathways including apoptosis, oxidative phosphorylation, fatty acid metabolism and amino acid metabolism. -
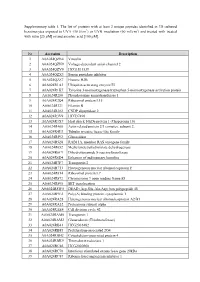
Supplementary Table 1. the List of Proteins with at Least 2 Unique
Supplementary table 1. The list of proteins with at least 2 unique peptides identified in 3D cultured keratinocytes exposed to UVA (30 J/cm2) or UVB irradiation (60 mJ/cm2) and treated with treated with rutin [25 µM] or/and ascorbic acid [100 µM]. Nr Accession Description 1 A0A024QZN4 Vinculin 2 A0A024QZN9 Voltage-dependent anion channel 2 3 A0A024QZV0 HCG1811539 4 A0A024QZX3 Serpin peptidase inhibitor 5 A0A024QZZ7 Histone H2B 6 A0A024R1A3 Ubiquitin-activating enzyme E1 7 A0A024R1K7 Tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein 8 A0A024R280 Phosphoserine aminotransferase 1 9 A0A024R2Q4 Ribosomal protein L15 10 A0A024R321 Filamin B 11 A0A024R382 CNDP dipeptidase 2 12 A0A024R3V9 HCG37498 13 A0A024R3X7 Heat shock 10kDa protein 1 (Chaperonin 10) 14 A0A024R408 Actin related protein 2/3 complex, subunit 2, 15 A0A024R4U3 Tubulin tyrosine ligase-like family 16 A0A024R592 Glucosidase 17 A0A024R5Z8 RAB11A, member RAS oncogene family 18 A0A024R652 Methylenetetrahydrofolate dehydrogenase 19 A0A024R6C9 Dihydrolipoamide S-succinyltransferase 20 A0A024R6D4 Enhancer of rudimentary homolog 21 A0A024R7F7 Transportin 2 22 A0A024R7T3 Heterogeneous nuclear ribonucleoprotein F 23 A0A024R814 Ribosomal protein L7 24 A0A024R872 Chromosome 9 open reading frame 88 25 A0A024R895 SET translocation 26 A0A024R8W0 DEAD (Asp-Glu-Ala-Asp) box polypeptide 48 27 A0A024R9E2 Poly(A) binding protein, cytoplasmic 1 28 A0A024RA28 Heterogeneous nuclear ribonucleoprotein A2/B1 29 A0A024RA52 Proteasome subunit alpha 30 A0A024RAE4 Cell division cycle 42 31 -

Global Proteomic Detection of Native, Stable, Soluble Human Protein Complexes
GLOBAL PROTEOMIC DETECTION OF NATIVE, STABLE, SOLUBLE HUMAN PROTEIN COMPLEXES by Pierre Claver Havugimana A thesis submitted in conformity with the requirements for the degree of Doctor of Philosophy Graduate Department of Molecular Genetics University of Toronto © Copyright by Pierre Claver Havugimana 2012 Global Proteomic Detection of Native, Stable, Soluble Human Protein Complexes Pierre Claver Havugimana Doctor of Philosophy Graduate Department of Molecular Genetics University of Toronto 2012 Abstract Protein complexes are critical to virtually every biological process performed by living organisms. The cellular “interactome”, or set of physical protein-protein interactions, is of particular interest, but no comprehensive study of human multi-protein complexes has yet been reported. In this Thesis, I describe the development of a novel high-throughput profiling method, which I term Fractionomic Profiling-Mass Spectrometry (or FP-MS), in which biochemical fractionation using non-denaturing high performance liquid chromatography (HPLC), as an alternative to affinity purification (e.g. TAP tagging) or immuno-precipitation, is coupled with tandem mass spectrometry-based protein identification for the global detection of stably- associated protein complexes in mammalian cells or tissues. Using a cell culture model system, I document proof-of-principle experiments confirming the suitability of this method for monitoring large numbers of soluble, stable protein complexes from either crude protein extracts or enriched sub-cellular compartments. Next, I document how, using orthogonal functional genomics information generated in collaboration with computational biology groups as filters, we applied FP-MS co-fractionation profiling to construct a high-quality map of 622 predicted unique soluble human protein complexes that could be biochemically enriched from HeLa and HEK293 nuclear and cytoplasmic extracts. -

Bioanalytical and Proteomic Approaches in the Study of Pathologic Ecs Dysfunctionality, Oxidative Stress and the Effects of PFKFB3 Modulators
Bioanalytical and proteomic approaches in the study of pathologic ECs dysfunctionality, oxidative stress and the effects of PFKFB3 modulators Sarath Babu Nukala ADVERTIMENT. La consulta d’aquesta tesi queda condicionada a l’acceptació de les següents condicions d'ús: La difusió d’aquesta tesi per mitjà del servei TDX (www.tdx.cat) i a través del Dipòsit Digital de la UB (diposit.ub.edu) ha estat autoritzada pels titulars dels drets de propietat intel·lectual únicament per a usos privats emmarcats en activitats d’investigació i docència. No s’autoritza la seva reproducció amb finalitats de lucre ni la seva difusió i posada a disposició des d’un lloc aliè al servei TDX ni al Dipòsit Digital de la UB. No s’autoritza la presentació del seu contingut en una finestra o marc aliè a TDX o al Dipòsit Digital de la UB (framing). Aquesta reserva de drets afecta tant al resum de presentació de la tesi com als seus continguts. En la utilització o cita de parts de la tesi és obligat indicar el nom de la persona autora. ADVERTENCIA. La consulta de esta tesis queda condicionada a la aceptación de las siguientes condiciones de uso: La difusión de esta tesis por medio del servicio TDR (www.tdx.cat) y a través del Repositorio Digital de la UB (diposit.ub.edu) ha sido autorizada por los titulares de los derechos de propiedad intelectual únicamente para usos privados enmarcados en actividades de investigación y docencia. No se autoriza su reproducción con finalidades de lucro ni su difusión y puesta a disposición desde un sitio ajeno al servicio TDR o al Repositorio Digital de la UB. -

Cell-Based Analysis of CAD Variants Identifies Individuals Likely to Benefit from Uridine Therapy
ARTICLE © American College of Medical Genetics and Genomics Cell-based analysis of CAD variants identifies individuals likely to benefit from uridine therapy Francisco del Caño-Ochoa, PhD1,2,25, Bobby G. Ng, BS3, Malak Abedalthagafi, MD4, Mohammed Almannai, MD5, Ronald D. Cohn, MD6,7,8,9, Gregory Costain, MD6,10, Orly Elpeleg, MD11, Henry Houlden, MD, PhD12, Ehsan Ghayoor Karimiani, MD, PhD13, Pengfei Liu, PhD14,15, M. Chiara Manzini, PhD16, Reza Maroofian, PhD12, Michael Muriello, MD17,18, Ali Al-Otaibi, MD19, Hema Patel, MD20, Edvardson Shimon, MD21, V. Reid Sutton, MD22, Mehran Beiraghi Toosi, MD23, Lynne A. Wolfe, MS24, Jill A. Rosenfeld, MS14,15, Hudson H. Freeze, PhD3 and Santiago Ramón-Maiques, PhD 1,2,25 Purpose: Pathogenic autosomal recessive variants in CAD,encoding Results: We identified 25 individuals with biallelic variants in CAD the multienzymatic protein initiating pyrimidine de novo biosynth- and a phenotype consistent with a CAD deficit. We used the CAD- esis, cause a severe inborn metabolic disorder treatable with a dietary knockout complementation assay to test a total of 34 variants, supplement of uridine. This condition is difficult to diagnose given identifying 16 as deleterious for CAD activity. Combination of these the large size of CAD with over 1000 missense variants and pathogenic variants confirmed 11 subjects with a CAD deficit, for the nonspecific clinical presentation. We aimed to develop a whom we describe the clinical phenotype. reliable and discerning assay to assess the pathogenicity of CAD Conclusions: We designed a cell-based assay to test the variants and to select affected individuals that might benefit from pathogenicity of CAD variants, identifying 11 CAD-deficient uridine therapy. -

C6cc08797c1.Pdf
Electronic Supplementary Material (ESI) for Chemical Communications. This journal is © The Royal Society of Chemistry 2017 Supplementary Information for Competition-based, quantitative chemical proteomics in breast cancer cells identifies new target profiles for sulforaphane James A. Clulow,a,‡ Elisabeth M. Storck,a,‡ Thomas Lanyon-Hogg,a,‡ Karunakaran A. Kalesh,a Lyn Jonesb and Edward W. Tatea,* a Department of Chemistry, Imperial College London, London, UK, SW7 2AZ b Worldwide Medicinal Chemistry, Pfizer, 200 Cambridge Park Drive, MA 02140, USA ‡ Authors share equal contribution * corresponding author, email: [email protected] Table of Contents 1 Supporting Figures..........................................................................................................................3 2 Supporting Tables.........................................................................................................................17 2.1 Supporting Table S1. Incorporation validation for the R10K8 label in the 'spike-in' SILAC proteome of the MCF7 cell line........................................................................................................17 2.2 Supporting Table S2. Incorporation validation for the R10K8 label in the 'spike-in' SILAC proteome of the MDA-MB-231 cell line ...........................................................................................17 2.3 Supporting Table S3. High- and medium- confidence targets of sulforaphane in the MCF7 cell line..............................................................................................................................................17 -

Supplementary Data
Supplemental Material Materials and Methods Immunohistochemistry Primary antibodies used for validation studies include: mouse anti-desmoglein-3 (Cat. # 32-6300, Invitrogen, CA, USA; 1:25), rabbit anti-cytokeratin 4 (Cat. # ab11215, Abcam, Cambridge, MA, USA; 1:100), mouse anti-cytokeratin 16 (Cat. # ab8741, Abcam; 1:25), rabbit anti-desmoplakin antibody (Cat. # ab14418, Abcam; 1:200), mouse anti-vimentin (Cat. # M7020, Dako, Carpinteria, CA, USA; 1:100). Secondary antibodies conjugated with biotin (Vector, Burlingame, CA, USA) were used, diluted to 1:400. Tissues slides containing archival FFPE sections, or tissue micro arrays (TMA) consisting of 508 HNSCC and controls, were dewaxed in SafeClear II (Fisher Scientific, Pittsburgh, PA, USA) hydrated through graded alcohols, immersed in 3% hydrogen peroxide in PBS for 30 min to quench the endogenous peroxidase, and processed for antigen retrieval and immunostaining with the appropriate primary antibodies and biotinylated secondary antibodies as described (1), followed by the avidin-biotin complex method (Vector Stain Elite, ABC kit; Vector). Slides were washed and developed in 3,3'- diaminobenzidine (Sigma FASTDAB tablet; Sigma Chemical) under microscopic control, and counterstained with Mayer's hematoxylin. For each stained TMA the number of positive cells in each core was visually evaluated and the results expressed as a percentage of stained cells/ total number of cells. According to their immunoreactivity the tissues array cores were divided according to tumor differentiation, where the percentage of stained cells in the three tumor classes were scored as more than 5% and less than 25% of the cells stained, 26 to 50%, 51 to 75% or, 75 to 100%.